Abstract
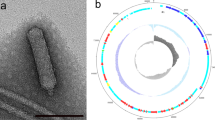
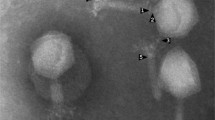
Multidrug-resistant Salmonella causing Salmonellosis is a food-borne pathogen and hence a public health hazard. Alternatives to antibiotics, such as phages, are possible solutions to this increasing drug resistance. In this context, several Salmonella phages were isolated and characterized. This paper describes the physiochemical and whole genome characterization of one such bacteriophage, ΦStp1, which efficiently infects serovars Salmonella Enteritidis and Salmonella Typhimurium. Morphological observations by transmission electron microscopy and phylogenetic analysis using terminase gene classified ΦStp1 to family Siphoviridae, closely resembling ‘T5 like phage’ morpho-types. With a maximum adsorption time of 50 min, ΦStp1 latent period was 30 min with 37 phages/cell burst size. ΦStp1 draft genome sequenced by shotgun method comprised 112,149 bp in 3 contigs with 37.99% GC content, 168 predicted ORFs, and 15 tRNAs. Genes involved in host shut down, DNA replication, regulation, nucleotide metabolism, lysis, and morphogenesis were also noted. The study not only provided an insight into the characteristics of phage genome, but also information about proteins encoded by bacteriophages, therefore contributing to understanding phage diversity. Sequence analysis also proved the absence of virulence and lysogeny-related genes, which only went to confirm ΦStp1 as a promising therapeutic agent against Salmonella infections.




Similar content being viewed by others
References
S.L. Foley, R. Nayak, I.B. Hanning, T.J. Johnson, J. Han, S.C. Ricke, Appl. Environ. Microbiol. 77, 4273–4279 (2011)
L.H. Su, C.H. Chiu, C. Chu, J.T Ou. Clin. Infect. Dis. 39, 546–551 (2004)
M. Kutateladze, R. Adamia, Med. Mal. Infect. 38, 426–430 (2008)
P. Garcia, B. Martinez, J.M. Obeso, A. Rodriguez, Lett. Appl. Microbiol. 47, 479–485 (2008)
S. Hagens, M.J. Loessner, Curr. Pharm. Biotechnol. 11, 58–68 (2010)
L.D. Goodridge, B. Bisha, Bacteriophage. 1, 130–137 (2011)
B. Leverentz, W.S. Conway, Z. Alavidze, W.J. Janisiewicz, Y. Fuchs, M.J. Camp, A. Sulakvelidze, J. Food Prot. 64, 1116–1121 (2001)
D. Goode, V.M. Allen, P.A. Barrow, Appl. Environ. Microbiol. 69, 5032–5036 (2003)
L. Fiorentin, N.D. Vieira, W. Barioni Júnior, Rev. Bras. Cienc. Avic. 7, 255–260 (2005)
M.H. Adams, Bacteriophages (Interscience Publisher Inc, London, 1959)
J. Sambrook, E. Fritsch, I. Maniatis, Molecular cloning – A laboratory manual, vol. 1 (Cold Spring Harbor Laboratory, Cold Spring Harbor, 2000)
Z. Lu, F. Breidt, H.P. Fleming, E. Altermann, T.R. Klaenhammer, Int. J. Food Microbiol. 84, 225–235 (2003)
E.A. Durmaz, MS Thesis. NC State Univ. Raleigh, NC, 1992
R. Ronen, C. Boucher, H. Chitsaz, P. Pevzner, Bioinformatics 28, 88–96 (2012)
J. Besemer, A. Lomsadze, M. Borodovsky, Nucl. Acids Res. 29, 2607–2618 (2001)
A. Marchler-Bauer, J.B. Anderson, M.K. Derbyshire, C. DeWeese-Scott, N.R. Gonzales, M. Gwadz, L. Hao, S. He, D.I. Hurwitz, J.D. Jackson, Z. Ke, D. Krylov, C.J. Lanczycki, C.A. Liebert, C. Liu, F. Lu, S. Lu, G.H. Marchler, M. Mullokandov, J.S. Song, N. Thanki, R.A. Yamashita, J.J. Yin, D. Zhang, S.H. Bryant, Nucl. Acids Res. 35, D237–D240 (2007)
E.M. Zdobnov, R. Apweiler, Bioinformatics 17, 847–848 (2001)
J. Söding, A. Biegert, A.N. Lupas, Nucl. Acids Res. 33, W244–W248 (2005)
T. Carver, N. Thomson, A. Bleasby, M. Berriman, J. Parkhill, Bioinformatics 25, 119–120 (2009)
T.M. Lowe, S.R. Eddy, Nucl. Acids Res. 25, 955–964 (1997)
K. Hofmann, W. Stoffel, Biol. Chem. Hoppe-Seyler 374, 166 (1993)
N. Saitou, M. Nei, Mol. Biol. Evol. 4, 406–425 (1987)
S. Kumar, G. Stecher, K. Tamura, Mol. Biol. Evol. 33, 1870–1874 (2016)
J.J. Gill, P. Hyman, Curr. Pharm. Biotechnol. 11, 2–14 (2010)
H.W. Ackermann, Methods Mol. Biol. 501, 127–140 (2009)
M. Krupovic, B.E. Dutilh, E.M. Adriaenssens, J. Wittmann, F.K. Vogensen, M.B. Sullivan, J. Rumnieks, D. Prangishvili, R. Lavigne, A.M. Kropinski, J. Klumpp, A. Gillis, F. Enault, R.A. Edwards, S. Duffy, M.R. Clokie, J. Barylski, H.W. Ackermann, J.H. Kuhn, Arch. Virol. 161, 1095–1099 (2016)
A.I.M. Switt, A. Sulakvelidze, M. Wiedmann, A.M. Kropinski, D.S. Wishart, C. Poppe, Y. Liang, In Salmonella ed. H. Schatten, A. Eisenstark, (Humana Press, New York, 2014) 237-287
S.T. Abedon, Bacteriophage. 1, 46–49 (2011)
J. Augustine, S.G. Bhat, J. Microbiol, Biotechnol. Food Sci. 4, 102 (2014)
J. Augustine, L. Louis, S.M. Varghese, S.G. Bhat, A. Kishore, J. Basic Microbiol. 53, 111–120 (2013)
Y. Wang, W. Wang, Y. Lv, W. Zheng, Z. Mi, G. Pei, X. An, X. Xu, C. Han, J. Liu, C. Zhou, J. Gen. Virol. 95, 2565–2575 (2014)
D. Turner, M. Hezwani, S. Nelson, V. Salisbury, D. Reynolds, J. Gen. Virol. 93, 2046–2056 (2012)
Y.A. Karpe, G.D. Kanade, K.D. Pingale, V.A. Arankalle, K. Banerjee, Virus Genes 52, 117–126 (2016)
D. Piya, Y. Xie, A.C.H. Morales, G.F.K. Everett, Genome Announc. 3, 1443–1444 (2015)
J. Wang, Y. Jiang, M. Vincent, Y. Sun, H. Yu, J. Wang, Q. Bao, H. Kong, S. Hu, Virology 332, 45–65 (2005)
S. Kala, N. Cumby, P.D. Sadowski, B.Z. Hyder, V. Kanelis, A.R. Davidson, K.L. Maxwell, Proc. Natl. Acad. Sci. U.S.A. 111, 6022–6027 (2014)
F. Rohwer, E. Edwards, J. Bacteriol. 184, 4529–4535 (2002)
S.R. Casjens, Curr. Opin. Microbiol. 8, 451–458 (2005)
F. Tétart, C. Desplats, M. Kutateladze, C. Monod, H.W. Ackermann, H.M. Krisch, J. Bacteriol. 183, 358–366 (2001)
G.F. Hatfull, R.W. Hendrix, Curr. Opin. Virol. 1, 98–303 (2011)
D. Lundin, E. Torrents, A.M. Poole, B.M. Sjöberg, BMC genomics 10, 1 (2009)
Y. Wang, X. Zhang, Virus Genes 37, 218–224 (2008)
C.K. Mathews, J. Biol. Chem. 242, 4083–4086 (1967)
L. Oliveira, P. Tavares, J.C. Alonso, Virus Res. 173, 247–259 (2013)
K.R. Kondabagil, V.B. Rao, J. Mol. Biol. 358, 67–82 (2006)
V.B. Rao, M. Feiss, Annu. Rev. Genet. 42, 647–681 (2008)
H. Oliveira, L.D. Melo, S.B. Santos, F.L. Nóbrega, E.C. Ferreira, N. Cerca, J. Azeredo, L.D. Kluskens, J. Virol. 87, 4558–4570 (2013)
K.M. Payne, G.F. Hatfull, PLoS ONE 7, e34052 (2012)
R.Y. Young, J. Mol, Microbiol. Biotechnol. 4, 21–36 (2002)
I.N. Wang, D.L. Smith, R. Young, Annu. Rev. Microbiol. 54, 799–825 (2000)
E.J. Summer, J. Berry, T.A.T. Tran, T.L. Niu, D.K. Struck, R. Young, Mol. Biol. 373, 1098–1112 (2007)
J. Berry, E.J. Summer, D.K. Struck, R. Young, Mol. Microbiol. 70, 341–351 (2008)
S.B. Santos, A.M. Kropinski, P.J. Ceyssens, H.W. Ackermann, A. Villegas, R. Lavigne, V.N. Krylov, C.M. Carvalho, E.C. Ferreira, J. Azeredo, J. Virol. 85, 11265–11273 (2011)
S.K. Kim, K. Makino, M. Amemura, H. Shinagawa, A. Nakata, J. Bacteriol. 175, 1316–1324 (2011)
J.M. Whichard, L.A. Weigt, D.J. Borris, L.L. Li, Q. Zhang, V. Kapur, F.W. Pierson, E.J. Lingohr, Y.M. She, A.M. Kropinski, N. Sriranganathan, Viruses. 2, 710–730 (2010)
M. Bailly-Bechet, M. Vergassola, E. Rocha, Genome Res. 17, 1486–1495 (2007)
Acknowledgements
The work was supported by KSCSTE project grant F.No. 009/SRSHS/2012/CSTE awarded to Dr. Sarita G. Bhat. The authors acknowledge Department of Biotechnology, Cochin University of Science and Technology for providing all facilities for research. The authors are grateful to Tina K.J for critical analysis of paper and Harisree P.Nair for suggestions in the analysis of data.
Author information
Authors and Affiliations
Contributions
Author SKS was responsible for bacteriophage isolation, preparation of DNA, genome annotation, analysis, and manuscript preparation. SGB conceived the study, aided the design of the experiments, and prepared manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Human and animals rights
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed consent
This article does not contain any studies with human participants.
Additional information
Edited by Joachim Jakob Bugert.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sritha, K.S., Bhat, S.G. Genomics of Salmonella phage ΦStp1: candidate bacteriophage for biocontrol. Virus Genes 54, 311–318 (2018). https://doi.org/10.1007/s11262-018-1538-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-018-1538-3