Abstract
Experimental methods of investigations of nanoparticle (NP)–protein interactions are limited, because they require a high amount of samples and the NPs tend to interfere with spectral results. Therefore, molecular modeling is a commonly accepted tool in such kind of investigations. Examining the molecule toxicity on the molecular level, we usually want to know, mainly, the location of the ligand on the protein surface and what is an influence of such a contact on the biological functions of the protein. In the presented work, we demonstrate that multiple-docking of the ligand from a random start and with large grid volume, to let the ligand search the whole protein surface, allows to find the best binding sites and gives reliable results considering ligand–protein interactions. In the present work, we have constructed six models of bronchoalveolar lavage fluids proteins: α1-antitripsin, albumin, ceruloplasmin, lactoferrin, lysozyme, and transferrin with fullerene, C60 utilizing molecular docking methods. The most probable results were examined with steered molecular dynamics (SMD) to see, if the simple docking method is able to predict the fullerene binding affinity. Albumin and lysozyme were already widely investigated and literature data is available for their complexes with fullerene C60 and/or its derivatives. Thus, we used these two models as a reference set to validate the used molecular modeling methods. With our best knowledge, interactions of the remaining four proteins with NPs have never been investigated in detail before. Our results indicate that fullerene C60 readily interacts with all studied proteins and may have a large impact on their biological functions.
Similar content being viewed by others
Introduction
Unique features of nanoparticles (NPs), in this number fullerenes, are the reason why these substances can be exploited in various fields of life, e.g., medicine, cosmetics, and electronics. On the other hand, there are still many questions related to unknown influence of nanomaterials on the environment and human health, and mainly related to their toxicity for alive organisms [1, 2]. NPs can be dangerous due to their high reactivity, easiness of dispersion, and migration through cellular membranes, or capability of transportation of other hazardous substances adsorbed on nanoparticle surface. Tiny size of NPs is related to a large surface area per unit mass, what rises chemical reactivity of these molecules [3].
The most dangerous seem to be NPs that can easily be transported through biological barriers (for example, the blood–brain barrier). It has been proven that NPs may modify biological responses and interactions of cells, for example, help bigger molecules to get inside organs; they can be transported along neurons [4], trigger oxidative stress and inflammations [3, 5]; and can be also responsible for genotoxicity and carcinogenesis [6]. Inhalation is one of the most probable and important exposure routes into the human body for various substances, in this number NPs [7–11]. Once a particle is deposited in the lungs, it starts to interact with epithelial or alveolar lining fluid proteins, before it is phagocytosed by macrophages or taken up by pneumocytes [7]. Inhaled NPs may also be transported through the whole body along various paths and utilizing various mechanisms. They can be distributed via the circulatory or lymphatic system and be deposited in various organs. As a result of accumulation of NPs in the liver, spleen, bone medulla, heart, and other organs, functions of an entire organism are disrupted [9, 10, 12, 13]. NP toxicity related to their routes of exposure are exhaustively described in [3, 14]. The smaller the particles and the more spherical the shape, the deeper they can get into the lungs. NPs smaller than 100 nm are deposited mainly in the alveolar area [8]. The observed negative influence of NPs on living organisms at the molecular level is directly related to their interactions with proteins and nucleic acids [15–18]. The lung epithelial lining fluid proteins are the first with which NPs can have a contact after being inhaled into human body. Therefore, interaction of NPs with these proteins is worth to study. The research at this point is more complicated, since NPs may be classified as substances usually tolerated by the organisms to some certain concentration, while when it is exceeded, they can reveal some toxic impact.
The most common and persistent are fullerenes built of 60 atoms (C60) [19], which possess 60 dislocated electrons. Experimental data indicates that the influence of fullerenes and their derivatives on biological structures depends on the size of a “carbon ball” and the structure of attached functional groups. Fullerenes can easily accept and donor electrons, so they have antioxidative and peroxidative capabilities, and have negative surface charge. Toxicity of fullerene C60 is mainly caused by generation of super oxide radical anion (O2−) [20] or by negative surface charge [21, 22]. Oxidative stress is especially intense in the lungs, where exists a large accumulation of reactive oxygen species (ROS) like the inflammatory phagocytes, neutrophils and macrophages [7]. The potential toxicity of the fullerenes may be caused by their shape. It was proven that C60 can block K+ channels, causing a biological effect [23]. Therefore, characteristic features of fullerenes as hydrophobicity, electrophilicity, and high reductive potential should not only be taken into consideration.
Fullerene C60 is insoluble in water, but creates stable aggregates with nanoscale dimensions, nC60 (25–500 nm) [24–27]. Aggregates remain in suspension for a long time [28] and their reactivity differs from separate C60 molecules. There are experimental data indicating that these aggregates are cytotoxic and cause damage triggered by electrostatic interactions with cell structures [21, 22] and by reactive forms of oxygen [20, 29]. It was previously reported that C60 nanoparticles are stabilized by adsorption of HSA and remain dispersed in physiological fluids [30]. Binding sites on HSA for nC60 occurred similar to these for separate molecules. The same type of binding sites as was reported for interactions C60–albumin [31] were experimentally detected in nC60–albumin complexes [32]. These observations indicate that even though investigations of separate C60 molecules are not complete as they should be for nC60, they bring reliable results, which can be extended on protein–nC60 complexes.
In the presented work, we indicate on the direct influence of C60 particles on proteins’ biological functions, e.g., blocking the ligand binding sites or interrupting conformational changes. We also describe C60–protein interactions and tried to find common features of such contacts for various proteins. The methods presented here, in contrary to computationally expensive molecular dynamics—were much less CPU time consuming. Our results indicate that in most of the cases, with the similar level of approximation, the predicted binding sites were reliable, as what was confirmed by a comparison with available experimental and computational data.
Materials and methods
In the presented study, we examined the interactions of fullerene C60 with selected epithelial lining fluid proteins in lungs. The other strategy is the database screening with reverse docking procedure [33]. All presented results were performed in silico, utilizing molecular modeling and docking methods. In-house tools and programs to models of visualization such as SwissPDBViewer 4.1 [34] and Chimera [35] were used for analysis of the identified binding sites. The protein structures examined here were taken from Protein Data Bank [36] and used for docking procedure: (1) albumin PDB ID: 1ao6 [37], (2) α1-antitripsin PDB ID: 3ne4 [38], (3) ceruloplasmin PDB ID: 4enz [39], (4) lactoferrin PDB ID: 1n76 [40], (5) lysozyme PDB ID: 3fe0 [41], and (6) transferrin PDB ID: 4h0w [42]. All computed models were prepared in SwissPDBViewer 4.1 program [34]. Missing amino acid residues were added where it was necessary; water, ligands, ions and other molecules unimportant for constructed models were removed. The 3D structure of nonsubstituted fullerene C60 was received by using Avogadro 1.1.1 program library [43]. Docking procedure was performed by AUTODOCK 1.5.6 package [44] with AMBER force field [45] as default for AUTODOCK program [46], since it is commonly used and a very potent tool in structure-based virtual screening [47], and it was already previously used to study fullerene derivatives with proteins [48–51].
Six complexes were constructed for the following proteins–C60 couples: (1) albumin–C60, (2) α1-antitripsin–C60, (3) ceruloplasmin–C60, (4) lactoferrin–C60, (5) lysozyme–C60, (6) transferrin–C60. No a priori settings were used to define searching space for a ligand, and all examined protein docking was carried with a large grid volume covering the entire surface of each protein. Initial conformation of the fullerene molecule in the docking space was generated randomly. Thus, fullerene was able to search the entire receptor surface and find the most preferable binding pockets. Docking parameters were as follows: 1000 runs from random start were performed for every protein (6000 result complexes in total), maximal number of retries was 1000; there were 150 individuals in the population and 1 individual was let to survive to next generation; there were maximally 27,000 generations; maximum number of energy evaluations was equal to 2,500,000; rate of gene mutation was set to 0.02 and rate of crossover to 0.8. During the docking procedure, the protein maintained rigid.
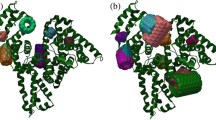
In-house tool was used to cluster ligand positions according to their distance from selected atoms in a protein structure. All clusters for each pair protein–ligand were ranked according to their free binding energies. Newly obtained complexes were visualized and analyzed in SPDBV program [34]. For the fullerene binding sites, three criteria were applied to select the most plausible ones for further investigations: (1) frequency of finding the site in repeated 1000 docking trials (free binding energy), (2) binding energy of the complex after docking, and (3) importance of location taken by the ligand for the protein functioning according to literature data. For each selected complex, one representative ligand with the lowest potential energy of binding was chosen and energy minimization was performed using YASARA package [52]. The amino acid residues within 7 Å radius from the ligand were acknowledged as the binding sites on proteins for C60. An example image for α1-antitripsin–C60 docking model is shown in Fig. 1. Collected data is presented in Table 1 and images of main binding sites for all investigated proteins are shown in Fig. 2.
Main binding sites detected on protein surface for fullerene C60 (red). a α1-antitripsin. b Albumin. c Ceruloplasmin. d Lactoferrin. e Lysozyme. f Transferrin. The protein hydrophobicity surface was drawn in the Kyte-Doolittle scale with colors ranging from dodger blue for the most hydrophilic to white at 0.0 to orange red for the most hydrophobic [35]
For a detailed investigation of the work required to extract a ligand from the identified binding sites (work done by pulling forces), we used steered molecular dynamics (SMD) as implemented in AMBER force field. The application of SMD was possible due to the discovery of the Jarzynski’s equation [53]. SMD applies an external force into a molecule and causes a change in coordinates with a specified time. Assuming that a single coordinate describes the investigated process, the external force can be described as F = 0.5k(x 0 + v t− x), where k is a force constant, x 0 and x are the initial and observed positions of the restraint point moving with a constant velovity v at time t. This force corresponds to the ligand being pulled by a harmonic spring stiffness k with its end moving with velocity v [54]. Jarzynski’s equation describes a relation between equilibrium of free energy differences between two states (bound and unbound) and work needed for transformation between those states exp.[−βΔF] = exp.[−βW] [55], where ΔF is a free energy difference between final (unbound) and initial (bounded) state, W is a work required to remove a ligand from a binding pocket and β = 1/RT.
Twenty-seven molecular systems were built (all investigated proteins and binding sites are listed in Table 1) and simulated using SMD as implemented in AMBER force field [56]. For every simulated system, 12 water layers of TIP3P water model and the necessary number of counter ions were added to neutralize a protein charge. Subsequently, the newly built systems were optimized with positional constraints applied to both ligand and protein in order to achieve a constant density. Finally, SMD simulation was performed with the following conditions: k = 40 [kcal/mol∙Å], T = 300 [K], v = 15 [Å/ns], MD step = 2 [fs], MD time = 1 [ns] and isothermal-isobaric (NTP) ensamble. For better and more reliable estimation of free energy, we used multisander procedure, running every simulation in 16 replicas as recommended in the AMBER manual [56]. Individual system parameters were as follows:
(1) albumin (HSA)—15 Na+ ions, 26,000 TIP3P water molecules, simulation box approximate size 97×84×103 Å; (2) α1-antitripsin—11 Na+ ions, 16,500 TIP3P water molecules, simulation box 95×68×84 Å; (3) ceruloplasmin—44 Na+ ions, 37,000 TIP3P water molecules, simulation box 105×112×108 Å; (4) lactoferrin—10 Cl− ions, 29,000 TIP3P water molecules, simulation box 92×90×115 Å; (5) lysozyme—8 Cl− ions, 7500 TIP3P water molecules, simulation box 63×60×66 Å; (6) transferrin—3 Na+ ions, 28,000 TIP3P water molecules, simulation box 110×93×90 Å. SMD results are collected in Table 1 and presented in Fig. 7.
Results and discussion
After docking, four to five main ligand binding sites were selected for each computed protein–C60 complex (see Table 1). For two of the examined proteins, previously published data exist regarding fullerene or fullerenol binding sites: human serum albumin (HSA) [31, 57, 58] and hen-egg white lysozyme with fullerenol C60(OH)x (x≈24) [50] and pristine fullerene C60 [59]. Thus, albumin–C60 and lysozyme–C60 complexes were used as a validation set.
Binding sites analysis
α1-antitripsin
Four binding sites were selected for α1-antitripsin according to the binding energy and ligand population criteria as described above (see Fig. 1 and Table 1). Two highly occupied (~10% ligands in each case) positions of fullerene were located in the reactive center loop (RCL) area (see Fig. 3), created by residues Lys343-Glu363 [60]. Additionally, Glu342 is involved in interactions with C60, and this residue creates a structurally important salt bridge Glu342-Lys290 [61]. Both, RCL and salt bridge play essential roles in α1-antitripsin biological functioning. All these residues are strongly conserved along serpin evolution and any changes or damages in this region lead to protein malfunction [74, 75]. However, as described, fullerene positions disrupting important regions in α1-antitripsin molecule are not “first choice” places taken by ligands during 1000 docking runs. There was a location detected with a much larger number of representatives and lower free binding energy of the complex (see Figs. 1 and 2 and Table 1). SMD mostly confirmed the results obtained by AutoDock. The largest work needed for extraction of C60 from site 1 is 29.24 kcal/mol (see Table 1). Site 4 contains lots of residues which were identified as important from the biological point of view for α1-antitripsin. In this case, the result obtained by SMD is 21.41 kcal/mol. The difference between site 2 and 3 is small, about 2.5 kcal/mol. These observations suggest that fullerene molecules may be hazardous for α1-antitripsin at higher concentration.
Albumin
From four main (highly populated) binding pockets detected for albumin–C60 model, the lowest energy location stays in an agreement with literature data [31] (see Table 1 and Fig. 4). This ligand position (ranked 3rd according to location occupancy, ~11% population) is equal to the type 1 binding site described by Benyamini et al. [31]. The location of the C60 close to the type 2 binding site introduced by those authors was also found with our calculation but its rank was too low (about 5% of the population). Those authors also reported that the type 1 binding site is favored by fullerene molecules interacting with HSA. Additionally, the type 1 binding site seems to be the more universal, because it can bind ligands, which are chemically very different, with a high affinity [76]. SMD results show the largest work was done for extraction of fullerene from site 1 (43.23 kcal/mol) (see Table 1 and Fig. 7); however, the most numerous ligand population was observed for site 3, as this site is located on the protein surface, and as a consequence, it is more accessible to the ligands. Site 2, similarly to site 1, possesses the nest shape, and the extraction work in this case is relatively high (30.45 kcal/mol).
Ceruloplasmin
Ceruloplasmin possesses strong oxidase activity towards numerous aromatic amines and phenols, e.g., norepinephrine, epinephrine, serotonin, dopa or (+)-lysergic acid diethylamine (LSD) [65]. Binding sites for metal cations—natural ligands of ceruloplasmin—are located in the inner parts of the protein molecule [62], so they cannot be reached by big C60 ligands.
Ceruloplasmin is involved in many protein–protein complexes being functional biological systems crucial for many cellular processes. It creates complexes with, e.g., myeloperoxidase [39] or lactoferrin [68, 77]. There are many amino acid residues involved in interactions with fullerene C60 on ceruloplasmin surface, which are considered as crucial for ceruloplasmin–lactoferrin or ceruloplasmin–myeloperoxidaze complexes (see Table 1). Around fragments Arg883-Arg892, His667-Trp669, and Cys699-Leu710 which are probably involved in interactions of cerruloplasmin with myeloperoxidaze [39, 64, 65] we found about 3, 10, and 8% populations of docked C60. Also, His1028-Val1037 fragment, previously recognized as important for binding various proteins to ceruloplasmin surface [66], was involved in interactions with fullerene molecules. Two binding places, with the population of about 8% in both cases, of the docked ligands, are located in these area (see Table 1). Additionally, Asn119 which carries a structurally important carbohydrate chain was involved in interactions with C60. These results are the hit to let us conclude that fullerene molecules attached to a protein surface may distinctly interrupt creating protein–protein complexes, mainly in higher concentrations. As a result, biological functions of all partners participating in such a complex are disturbed. Numerous fullerene molecules (or nC60) connected with protein may totally prevent to create a functional protein quaternary structures. SMD performed for the highly populated ceruloplasmin–C60 complexes showed very similar ligand detachment work for 1–4 pockets (22.99, 25.24, 20.27, and 28.77 kcal/mol, respectively). The highest work was observed during the pulling of C60 from site 5 (35.43 kcal/mol). Site 5 was confirmed as important for the biological function of ceruloplasmin [66].
Lactoferrin
Fullerene docking to lactoferrin gave the most fuzzy results, and binding energies were relatively high among all examined protein–fullerene complexes (see Table 1). Five binding pockets with the lowest energy have been selected for further analysis. They have about 7–8% of ligand occupancy. Significant dispersion of C60 molecules on lactoferrin surface suggests that this protein is receptive for C60 ligands. Although, relatively big fullerene molecules attached to lactoferrin surface cannot directly affect metal binding sites located inside protein structure, but they may interrupt conformational changes of lactoferrin which are required for iron release [78].
It is known that, similarly to ceruloplasmin, lactoferrin creates functional complexes with various proteins, in this number of heparin or ceruloplasmin. The N-terminal fragment Arg2-Arg5 was reported as crucial in this type of protein–protein interactions [66, 68] and we have found it in the fifth binding pocket, which reveals relatively low free energy. However, the interaction energy was the highest among all identified lactoferin binding sites (see Table 1). Similarly to ceruloplasmin, C60 molecules attached to the protein structure may seriously disturb forming functional protein–protein complexes. In the contrary to the previous protein–C60 complexes, in this case, there is no correlation between the interaction energy and ligand population as calculated by the AutoDock program and the results from the SMD simulation. It is mostly visible in comparison between site 3 and 4. In both cases, we have almost identical results as obtained by AutoDock, however SMD gave about 18 kcal/mol difference. It is caused by the location of the ligand. In the case of site 4 fullerene is buried inside of the protein, in the case of site 3, located on the protein surface. During the pulling simulation, the protein–ligand contact was preserved for much longer time and it caused such a difference in the result.
Lysozyme
Lysozyme is the smallest from all proteins presented here, and after docking, it gave the most strict results. Only seven ligand locations were obtained after the docking simulation. Five of them were taken into consideration as possible binding pockets for fullerene C60 (see Table 1). The lowest protein–ligand interactions energy (2nd in an occupancy rank ~26% of ligands) is located at the end of the active cleft of lysozyme and contains most of amino acid residues recognized previously as important for hen-egg white lysozyme original ligand binding [50, 69–71, 79] (see Table 1 and Fig. 5). Five other binding sites contain amino acid residues recognized as important for other ligand binding [72]; therefore, we considered them in the presented results. It was also experimentally proven that increasing concentration of fullerenol adsorbed to lysozyme decreases its enzymatic activity because of blocking a binding cleft [50]. Our computational results remain in an agreement with previous experimental and computational study data (see Table 1 and Fig. 5). SMD results for all examined binding sites were quite similar (from about 10 to 15 kcal/mol). The only exception was site 3 with extraction work of about 19 kcal/mol. This result is caused by the fact that fullerene molecule is relatively large in comparison to lysozyme protein. All identified sites were situated on the protein surface, with site 3 forming a kind of shallow nest.
Transferrin
The most plausible binding pocket observed for transferrin is placed in a functionally important area of this protein including Asn413 and Asn611 [73] and it was identified as site 1. The second binding site was the target of 20% of the ligand population. The third important binding pocket contains only one residue from transferrin binding site, Tyr95. The fourth ranked binding pocket (~6% population) consists of Asp63 and His249, which create the original protein binding site and Lys296 which, with other amino acid residues, stabilizes the metal binding site and play a crucial role in iron release (see Table 1). Our observations suggest that fullerene molecules attached to transferrin surfaces may partly inhibit the binding pocket and interrupt other biological functions of the protein but probably only at high concentrations of C60. This conclusion stays in agreement with experimental data related to transferrin–highly hydrophobic compound interactions, which shows that such complexes lead to irreparable damage of biological functions of transferrin [80]. SMD calculations pointed site 2 as the most attractive to ligands (42.79 kcal/mol), while the work of pulling for site 1 was the lowest observed (25.58 kcal/mol) and close to the values for the remaining binding sites. Site 1 contained twice as more polar than nonpolar residues, while for sites 2 and 3, this ratio was almost 1:1. This can explain the differences in the values obtained by SMD simulation. Moreover, site 2 has the deepest nest shape, since all remaining binding sites were located on the protein surface.
Characterization of the observed fullerene C60–protein interactions
One of the goals of our study was to characterize specific interactions present between fullerene C60 and protein to finding common features of such interactions. We found that there are almost two times more nonpolar (~260) than polar (~145) residues interacting with C60 in examined complexes. In the case of polar residues, glutamic acid and lysine are the most often appearing residues in all described binding sites. In the case of nonpolar residues, these are leucine and alanine (see Fig.6). With aromatic amino acid residues as tryptophan, phenylalanine, or tyrosine, fullerene C60 used to create π–π stacking interactions [50] and we detected many of them in the computed models. Aromatic residues make about 12% of all residues found in binding sites (see Fig. 6). Thus, we suggest these interactions are important for stabilization of fullerene–protein complex, but surely, they have to be supported by other kinds of interactions. Another type of interaction which should be taken into consideration in C60–protein complexes are cation–π interactions. They were found in many systems and should be acknowledged as important non-covalent binding forces [81]. Cation–π interactions are highly competitive with other kinds of non-covalent bonding forces which are known as strong ones [82]. Since they are not only “cation–aromatic” relations, for fullerene molecules with 60 dislocated electrons, they should be a very important kind of contact with proteins. Some computational studies confirmed this fact for semibuckminsterfullerene [83–86]. In our models there is plenty of arginine, lysine, asparagine, glutamine and histidine residues in the identyfied binding sites. These residues constitute about 25% of all amino acid residues as described here binding places (see Fig. 6). This result suggest, that cation–π interactions are of a high importance for structural models and they are probably the main common feature for all of them. It was observed that in proteins, statistically, cation–π interactions are almost two times more probable than van der Waals contacts between amino acids so-called chains [87]. Brocchieri et al. reported that lysine and mainly arginine are the most common neighbors of tyrosine and tryptophan in protein structures [88]. These observations suggest that fullerene may be competitive to tyrosine and tryptophan in this kind of interactions, when adsorbed on a protein surface. It may be a reason why we observed two times more possible cation–π than π–π contact spots. It is worth to mention that cation–π interactions which involve neutral side chains of asparagine or glutamine are much weaker [89] than the same interactions created by positively charged groups of arginine or lysine [90]. Interestingly, cation–π interactions contribute to α-helix stability [91], and it was observed that fullerenol decreased the content of α-helix without significantly interrupting the whole secondary structure in complexes with lysozyme [50] and HSA [58]. This fact may also indirectly indicate that cation–π interactions exist between C60 and a protein in their complex.
Amino acid residues participating in detected binding sites: blue—residues creating first binding sites (for the most probable binding pockets, see Table 1) for all proteins, orange—residues creating all presented here binding sites
Another type of fullerene–protein contact which should be taken into consideration are anion–π interactions [92]. These binding forces have been defined between negatively charged species and electron-deficient aromatic rings [93, 94]. The C60–protein complexes presented herein, reveal high number of glutamic acid and aspartic acid residues in observed binding sites (~14% of all residues), and additionally, glutamic acid is the most numerous amino acid residue of all (see Fig. 6). Fullerene molecules are rich in dislocated electrons and double bonds, and such structures seem to promote the creation of anion–π interactions. Fullerene molecules can also create a weak hydrogen bond interaction with amino acid residues, since localized double bonds of C60 may act as weak hydrogen bond acceptor with the hydroxyl group of tyrosine [95] or N-H of an amide group [96, 97]. The small statistics as shown in Fig. 6 pointed that the most preferred residues for fullerene interactions are hydrophilic residues: GLU, LYS, ARG, ASN, and ASP. In the case of hydrophobic residues, surprisingly, these are ALA and LEU (Fig. 7).
The results of SMD simulations. The work and the time are shown on vertical and horizontal axes, respectively. The gray lines correspond to single simulation; the average work was drawn as black, bold line. The final results are summarized in Table 1
Conclusions
Docking results presented here were obtained from rigid structures of proteins, and possible conformational changes triggered by fullerene molecules were not performed in our models. It may have an influence on the accuracy of described binding pockets and protein–ligand interactions. But yet, available literature data shows that there are no large-scale conformational changes in the secondary structure of the protein even after binding more than one fullerene molecule [58, 98].
Data presented here indicates that protein surface used to be highly receptive for fullerene C60 molecules and that these ligands tend to interact with active sites of proteins. Besides, numerous ligand particles attached to receptor surface must strongly interrupt protein biological functions. Fullerene may not only block binding sites but also interfere with conformational changes of the protein required in its activation process, compete with original substrates having, for example, the same surface charge as fullerene, or affect a product release. They also may interrupt creating functional protein–protein complexes. Observed fullerene–protein interactions may have a toxic influence on alive organisms caused by malfunctioning of proteins crucial in biochemical processes.
Our results indicated that in most of the cases, the simple docking model is able to predict, in a very good affinity, the protein–fullerene interface and binding affinity. However, in some of the cases, in which the binding pocket is buried in the protein, the observed differences in comparison to SMD simulations are high. Our results confirmed that the most preferable residues for protein fullerene interactions are hydrophilic: GLU, LYS, ARG, ASN, and ASP; in the case of hydrophobic residues, these are ALA and LEU.
References
Giles J (2003) Nanotechnology: what is there to fear from something so small? Nature 426:750–750. doi:10.1038/426750a
Maynard AD, Kuempel ED (2005) Airborne nanostructured particles and occupational health. J Nanopart Res 7:587–614. doi:10.1007/s11051-005-6770-9
Aydın A, Sipahi H, Charehsaz M (2012) Pharmacology, Toxicology and Pharmaceutical Science, Pharmacology, "Recent Advances in Novel Drug Carrier Systems", Ali Demir Sezer (ed) , ISBN 978-953-51-0810-8
Flesken-Nikitin A, Toshkov I, Naskar J, et al (2007) Toxicity and biomedical imaging of layered nanohybrids in the mouse. Toxicol Pathol 35:804–810. doi:10.1080/01926230701584239
Quadros ME, Marr LC (2010) Environmental and human health risks of aerosolized silver nanoparticles. J Air Waste Manage Assoc 60:770–781. doi:10.3155/1047-3289.60.7.770
Li JJ, Muralikrishnan S, Ng C-T, et al (2010) Nanoparticle-induced pulmonary toxicity. Exp Biol Med 235:1025–1033. doi:10.1258/ebm.2010.010021
Dailey LA, Jekel N, Fink L, et al (2006) Investigation of the proinflammatory potential of biodegradable nanoparticle drug delivery systems in the lung. Toxicol Appl Pharmacol 215:100–108. doi:10.1016/j.taap.2006.01.016
Peter HM, Hoet PHM, Brüske-Hohlfeld I, Salata OV (2004) Nanoparticles—known and unknown health risks. Nanobiotechnol 2 12:1–15. doi:10.1186/1477-3155-2-12
Warheit DB (2003) Comparative pulmonary toxicity assessment of single-wall carbon nanotubes in rats. Toxicol Sci 77:117–125. doi:10.1093/toxsci/kfg228
Oberdörster G, Oberdörster E, Oberdörster J (2005) Nanotoxicology: an emerging discipline evolving from studies of ultrafine particles. Environ Health Perspect 113:823–839. doi:10.1289/ehp.7339
Nemmar A, Vanbilloen H, Hoylaerts MF, et al (2001) Passage of intratracheally instilled ultrafine particles from the lung into the systemic circulation in hamster. Am J Respir Crit Care Med 164:1665–1668. doi:10.1164/ajrccm.164.9.2101036
Medina C, Santos-Martinez MJ, Radomski A, et al (2009) Nanoparticles: pharmacological and toxicological significance: nanoparticles. Br J Pharmacol 150:552–558. doi:10.1038/sj.bjp.0707130
Zhang Z, Kleinstreuer C, Donohue JF, Kim CS (2005) Comparison of micro- and nano-size particle depositions in a human upper airway model. J Aerosol Sci 36:211–233. doi:10.1016/j.jaerosci.2004.08.006
Yah CS, Simate GS, Iyuke SE (2012) Nanoparticles toxicity and their routes of exposures. Pak J Pharm Sci 25(2):477–491
Monopoli MP, Walczyk D, Campbell A, et al (2011) Physical−chemical aspects of protein corona: relevance to in vitro and in vivo biological impacts of nanoparticles. J Am Chem Soc 133:2525–2534. doi:10.1021/ja107583h
Dowling AP (2004) Development of nanotechnologies. Mater Today 7:30–35. doi:10.1016/S1369-7021(04)00628-5
Fleischer T, Grunwald A (2008) Making nanotechnology developments sustainable. A role for technology assessment? J Clean Prod 16:889–898. doi:10.1016/j.jclepro.2007.04.018
Owen R, Depledge M (2005) Nanotechnology and the environment: risks and rewards. Mar Pollut Bull 50:609–612. doi:10.1016/j.marpolbul.2005.05.001
Kroto HW, Heath JR, O’Brien SC, et al (1985) C60: Buckminsterfullerene. Nature 318:162–163. doi:10.1038/318162a0
Sayes CM, Fortner JD, Guo W, et al (2004) The differential cytotoxicity of water-soluble fullerenes. Nano Lett 4:1881–1887. doi:10.1021/nl0489586
Fortner JD, Lyon DY, Sayes CM, et al (2005) C60 in water: nanocrystal formation and microbial response. Environmental Science & Technology 39:4307–4316. doi:10.1021/es048099n
Lyon DY, Fortner JD, Sayes CM, et al (2005) Bacterial cell association and antimicrobial activity of a C60 water suspension. Environ Toxicol Chem 24:2757. doi:10.1897/04-649R.1
Calvaresi M, Furini S, Domene C, et al (2015) Blocking the passage: C60 geometrically clogs K+ channels. ACS Nano 9:4827–4834. doi:10.1021/nn506164s
Prato M (1997) [60]Fullerene chemistry for materials science applications. J Mater Chem 7:1097–1109. doi:10.1039/a700080d
Brant J, Lecoanet H, Wiesner MR (2005) Aggregation and deposition characteristics of fullerene nanoparticles in aqueous systems. J Nanopart Res 7:545–553. doi:10.1007/s11051-005-4884-8
Scharff P, Risch K, Carta-Abelmann L, et al (2004) Structure of C60 fullerene in water: spectroscopic data. Carbon 42:1203–1206. doi:10.1016/j.carbon.2003.12.053
Cheng X, Kan AT, Tomson MB (2004) Naphthalene adsorption and desorption from aqueous C60 fullerene. J Chem Eng Data 49:675–683. doi:10.1021/je030247m
Deguchi S, Alargova RG, Tsujii K (2001) Stable dispersions of fullerenes, C60 and C70, in water. Preparation and characterization. Langmuir 17:6013–6017. doi:10.1021/la010651o
Sayes CM, Gobin AM, Ausman KD, et al (2005) Nano-C60 cytotoxicity is due to lipid peroxidation. Biomaterials 26:7587–7595. doi:10.1016/j.biomaterials.2005.05.027
Deguchi S, Yamazaki T, Mukai S, et al (2007) Stabilization of C60 nanoparticles by protein adsorption and its implications for toxicity studies. Chem Res Toxicol 20:854–858. doi:10.1021/tx6003198
Benyamini H, Shulman-Peleg A, Wolfson HJ, et al (2006) Interaction of C60-fullerene and carboxyfullerene with proteins: docking and binding site alignment. Bioconjug Chem 17:378–386. doi:10.1021/bc050299g
Song M, Liu S, Yin J, Wang H (2011) Interaction of human serum albumin and C60 aggregates in solution. Int J Mol Sci 12:4964–4974. doi:10.3390/ijms12084964
Calvaresi M, Zerbetto F (2010) Baiting proteins with C 60. ACS Nano 4:2283–2299. doi:10.1021/nn901809b
Guex N, Peitsch MC (1997) SWISS-MODEL and the Swiss-Pdb viewer: an environment for comparative protein modeling. Electrophoresis 18:2714–2723. doi:10.1002/elps.1150181505
Pettersen EF, Goddard TD, Huang CC, et al (2004) UCSF chimera. A visualization system for exploratory research and analysis Journal of Computational Chemistry 25:1605–1612. doi:10.1002/jcc.20084
Berman H, Henrick K, Nakamura H (2003) Announcing the worldwide protein data bank. Nat Struct Biol 10:980–980. doi:10.1038/nsb1203-980
Sugio S, Kashima A, Mochizuki S, et al (1999) Crystal structure of human serum albumin at 2.5 A resolution. Protein Eng Des Sel 12:439–446. doi:10.1093/protein/12.6.439
Patschull AOM, Segu L, Nyon MP, et al (2011) Therapeutic target-site variability in α1-antitrypsin characterized at high resolution. Acta Crystallographica Section F Structural Biology and Crystallization Communications 67:1492–1497. doi:10.1107/S1744309111040267
Samygina VR, Sokolov AV, Bourenkov G, et al (2013) Ceruloplasmin: macromolecular assemblies with iron-containing acute phase proteins. PLoS One 8:e67145. doi:10.1371/journal.pone.0067145
Kumar J, Weber W, Munchau S, Yadav A, Sarvanan K, Paramsivam M, Sharma S, Kaur P, Bhushan A, Srinivasan A, Betzel C, Singh TOP (2003) Crystal structure of human seminal diferric lactoferrin at 3.4 Å resolution. Indian J Biochem Biophys 40:14–21
Chiba-Kamoshida K, Matsui T, Chatake T, Ohhara T, Ostermann A, Tanaka I, Yutani K, Niimura N (2008) Site-specific softening of peptide bonds by localized deuterium observed by neutron crystallography of human lysozyme hydrogen. X-ray crystal structure of wild type human lysozyme in D2O. doi: 10.2210/pdb3fe0/pdb
Yang N, Zhang H, Wang M, et al (2012) Iron and bismuth bound human serum transferrin reveals a partially-opened conformation in the N-lobe. Scientific Reports. doi:10.1038/srep00999
Hanwell MD, Curtis DE, Lonie DC, et al (2012) Avogadro: an advanced semantic chemical editor, visualization, and analysis platform. Journal of Cheminformatics 4:17. doi:10.1186/1758-2946-4-17
Morris GM, Huey R, Lindstrom W, et al (2009) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 30:2785–2791. doi:10.1002/jcc.21256
Pearlman DA, Case DA, Caldwell JW, et al (1995) AMBER, a package of computer programs for applying molecular mechanics, normal mode analysis, molecular dynamics and free energy calculations to simulate the structural and energetic properties of molecules. Comput Phys Commun 91:1–41. doi:10.1016/0010-4655(95)00041-D
Huey R, Morris GM, Olson AJ, Goodsell DS (2007) A semiempirical free energy force field with charge-based desolvation. J Comput Chem 28:1145–1152. doi:10.1002/jcc.20634
Park H, Lee J, Lee S (2006) Critical assessment of the automated AutoDock as a new docking tool for virtual screening. Proteins: Structure, Function, and Bioinformatics 65:549–554. doi:10.1002/prot.21183
Kankanala R, Ganugapati J, Vutukuru SS, Sai KSSR (2013) Molecular docking analysis of fullerene (C60) with human antioxidant enzymes: implications in inhibition of enzymes. J Comput Theor Nanosci 10:1403–1407. doi:10.1166/jctn.2013.2861
Anil N, S. Vutukuru S, S. R. Sivasai K, Ganugapati J (2013) In silico studies of fullerene C60 with zebrafish proteins prostacyclin synthase and S100z calcium binding protein. International Journal of Computer Applications 71:6–10. doi: 10.5120/12376-8720
Yang S-T, Wang H, Guo L, et al (2008) Interaction of fullerenol with lysozyme investigated by experimental and computational approaches. Nanotechnology 19:395101. doi:10.1088/0957-4484/19/39/395101
Lee VS, Nimmanpipug P, Aruksakunwong O, et al (2007) Structural analysis of lead fullerene-based inhibitor bound to human immunodeficiency virus type 1 protease in solution from molecular dynamics simulations. J Mol Graph Model 26:558–570. doi:10.1016/j.jmgm.2007.03.013
Krieger E, Joo K, Lee J, et al (2009) Improving physical realism, stereochemistry, and side-chain accuracy in homology modeling: four approaches that performed well in CASP8. Proteins: Structure, Function, and Bioinformatics 77:114–122. doi:10.1002/prot.22570
Jarzynski C (1997) Nonequilibrium equality for free energy differences. Phys Rev Lett 78:2690–2693. doi:10.1103/PhysRevLett.78.2690
Izrailev S, Stepaniants S, Isralewitz B, Kosztin D, Lu H, Molnar F, Wriggers W, Schulten K (1998) Steered molecular dynamics. Computational Molecular Dynamics: Challenges, Methods, Ideas 4:36–62
Park S, Khalili-Araghi F, Tajkhorshid E, Schulten K (2003) Free energy calculation from steered molecular dynamics simulations using Jarzynski’s equality. J Chem Phys 119:3559–3566. doi:10.1063/1.1590311
Cornell WD, Cieplak P, Bayly CI, et al (1995) A second generation force field for the simulation of proteins, nucleic acids, and organic molecules. J Am Chem Soc 117:5179–5197. doi:10.1021/ja00124a002
Belgorodsky B, Fadeev L, Ittah V, et al (2005) Formation and characterization of stable human serum albumin−Tris-malonic acid [C 60 ]fullerene complex. Bioconjug Chem 16:1058–1062. doi:10.1021/bc050103c
Li S, Zhao X, Mo Y, et al (2013) Human serum albumin interactions with C60 fullerene studied by spectroscopy, small-angle neutron scattering, and molecular dynamics simulations. J Nanopart Res. doi:10.1007/s11051-013-1769-0
Calvaresi M, Arnesano F, Bonacchi S, et al (2014) C60 @lysozyme: direct observation by nuclear magnetic resonance of a 1:1 fullerene protein adduct. ACS Nano 8:1871–1877. doi:10.1021/nn4063374
Schick C, Bromme D, Bartuski AJ, et al (1998) The reactive site loop of the serpin SCCA1 is essential for cysteine proteinase inhibition. Proc Natl Acad Sci 95:13465–13470. doi:10.1073/pnas.95.23.13465
Carrell RW, Jeppsson J-O, Laurell C-B, et al (1982) Structure and variation of human α1–antitrypsin. Nature 298:329–334. doi:10.1038/298329a0
Bento I, Peixoto C, Zaitsev VN, Lindley PF (2007) Ceruloplasmin revisited: structural and functional roles of various metal cation-binding sites. Acta Crystallographica Section D Biological Crystallography 63:240–248. doi:10.1107/S090744490604947X
Sokolov AV, Pulina MO, Ageeva KV, et al (2007) Identification of leukocyte cationic proteins that interact with ceruloplasmin. Biochem Mosc 72:872–877. doi:10.1134/S0006297907080093
Sokolov AV, Ageeva KV, Pulina MO, et al (2008) Ceruloplasmin and myeloperoxidase in complex affect the enzymatic properties of each other. Free Radic Res 42:989–998. doi:10.1080/10715760802566574
Zaitsev VN, Zaitseva I, Papiz M, Lindley PF (1999) An X-ray crystallographic study of the binding sites of the azide inhibitor and organic substrates to ceruloplasmin, a multi-copper oxidase in the plasma. J Biol Inorg Chem 4:579–587. doi:10.1007/s007750050380
Sokolov AV, Pulina MO, Zakharova ET, et al (2006) Identification and isolation from breast milk of ceruloplasmin-lactoferrin complex. Biochem Mosc 71:160–166. doi:10.1134/S0006297906020076
Baker EN, Anderson BF, Baker HM, et al (1990) Metal and anion binding sites in lactoferrin and related proteins. Pure Appl Chem. doi:10.1351/pac199062061067
Pulina MO, Zakharova ET, Sokolov AV, et al (2002) Studies of the ceruloplasmin-lactoferrin complex. Biochem Cell Biol 80:35–39. doi:10.1139/o01-206
Strynadka NCJ, James MNG (1991) Lysozyme revisited: crystallographic evidence for distortion of an N-acetylmuramic acid residue bound in site D. J Mol Biol 220:401–424. doi:10.1016/0022-2836(91)90021-W
Vocadlo DJ, Davies GJ, Laine R, Withers SG (2001) Catalysis by hen egg-white lysozyme proceeds via a covalent intermediate. Nature 412:835–838. doi:10.1038/35090602
Harata K, Muraki M, Jigami Y (1993) Role of Arg115 in the catalytic action of human lysozyme. J Mol Biol 233:524–535. doi:10.1006/jmbi.1993.1529
Madhusudan VM (1992) Additional binding sites in lysozyme. X-ray analysis of lysozyme complexes with bromophenol red and bromophenol blue. Protein engineering. Design and Selection 5:399–404. doi:10.1093/protein/5.5.399
Gomme PT, McCann KB, Bertolini J (2005) Transferrin: structure, function and potential therapeutic actions. Drug Discov Today 10:267–273. doi:10.1016/S1359-6446(04)03333-1
Irving JA, PikeRN LAM, Whisstock JC (2000) Phylogeny of the serpin superfamily: implications of patterns of amino acid conservation for structure and function. Genome Res 10:1845–1864. doi:10.1101/gr.147800
Lomas DA, LI-Evans D, Finch JT, Carrell RW (1992) The mechanism of Z α1-antitrypsin accumulation in the liver. Nature 357:605–607. doi: 10.1038/357605a0
T. Peters, Jr. (ed) (1996) All about albumin: biochemistry, genetics, and medical applications, Academic Press, San Diego. ISBN 0-12-552110-3
Sabatucci A, Vachette P, Vasilyev VB, et al (2007) Structural Characterization of the Ceruloplasmin: Lactoferrin complex in solution. J Mol Biol 371:1038–1046. doi:10.1016/j.jmb.2007.05.089
Kretchmar SA, Raymond KN (1986) Biphasic kinetics and temperature dependence of iron removal from transferrin by 3,4-LICAMS. J Am Chem Soc 108:6212–6218. doi:10.1021/ja00280a017
Calvaresi M, Bottoni A, Zerbetto F (2015) Thermodynamics of binding between proteins and carbon nanoparticles: the case of C 60 @lysozyme. J Phys Chem C 119:28077–28082. doi:10.1021/acs.jpcc.5b09985
Drug E, Fadeev L, Gozin M (2011) The impact of highly hydrophobic material on the structure of transferrin and its ability to bind iron. Toxicol Lett 203:33–39. doi:10.1016/j.toxlet.2011.02.017
Dougherty DA (1996) Cation-pi interactions in chemistry and biology: a new view of benzene, Phe, Tyr, and Trp. Science 271:163–168. doi:10.1126/science.271.5246.163
Scrutton NS, Raine ARC (1996) Cation-π bonding and amino-aromatic interactions in the biomolecular recognition of substituted ammonium ligands. Biochem J 319:1–8. doi:10.1042/bj3190001
Ma JC, Dougherty DA (1997) The cation−π interaction. Chem Rev 97:1303–1324. doi:10.1021/cr9603744
Faust R, Vollhardt KPC (1993) Semibuckminsterfullerene: MNDO study of a hemispherical triindenotriphenylene. J Chem Soc Chem Commun 1471. doi:10.1039/c39930001471
Sygula A, Rabideau PW (1994) Bowl-to-bowl inversion in polynuclear aromatic hydrocarbons with curved surfaces: an ab initio study. Journal of the Chemical Society, Chemical Communications 1497. doi: 10.1039/c39940001497
Cioslowski J, Lin Q (1995) Guest discrimination in complexes of alkali metal cations with the C36H36 spheriphane: an ab initio electronic structure study. J Am Chem Soc 117:2553–2556. doi:10.1021/ja00114a018
Burley SK, Petsko GA (1988) Weakly polar interactions in proteins. In: Advances in protein chemistry. Elsevier, pp 125–189. doi: 10.1016/S0065-3233(08)60376-9
Brocchieri L, Karlin S (1994) Geometry of interplanar residue contacts in protein structures. Proc Natl Acad Sci U S A 91(20):9297–9301
Rodham DA, Suzuki S, Suenram RD, et al (1993) Hydrogen bonding in the benzene–ammonia dimer. Nature 362:735–737. doi:10.1038/362735a0
Meot-Ner M, Deakyne CA (1985) Unconventional ionic hydrogen bonds. 2. NH+.cntdot..cntdot..cntdot..pi.. complexes of onium ions with olefins and benzene derivatives. J Am Chem Soc 107:474–479. doi:10.1021/ja00288a034
Shoemaker KR, Fairman R, Schultz DA, et al (1990) Side-chain interactions in the C-peptide helix: Phe 8 His 12+. Biopolymers 29:1–11. doi:10.1002/bip.360290104
Wang D-X, Wang M-X (2013) Anion−π interactions: generality, binding strength, and structure. J Am Chem Soc 135:892–897. doi:10.1021/ja310834w
Berryman OB, Bryantsev VS, Stay DP, et al (2007) Structural criteria for the design of anion receptors: the interaction of halides with electron-deficient arenes. J Am Chem Soc 129:48–58. doi:10.1021/ja063460m
Hay BP, Bryantsev VS (2008) Anion–arene adducts: C–H hydrogen bonding, anion–π interaction, and carbon bonding motifs. Chem Commun 2417. doi:10.1039/b800055g
Braden BC, Goldbaum FA, Chen B-X, et al (2000) X-ray crystal structure of an anti-buckminsterfullerene antibody fab fragment: biomolecular recognition of C60. Proc Natl Acad Sci 97:12193–12197. doi:10.1073/pnas.210396197
Levitt M, Perutz MF (1988) Aromatic rings act as hydrogen bond acceptors. J Mol Biol 201:751–754. doi:10.1016/0022-2836(88)90471-8
Perutz MF (1993) The role of aromatic rings as hydrogen-bond acceptors in molecular recognition. Philos Trans R Soc A Math Phys Eng Sci 345:105–112. doi:10.1098/rsta.1993.0122
Liu S, Sui Y, Guo K, et al (2012) Spectroscopic study on the interaction of pristine C60 and serum albumins in solution. Nanoscale Res Lett 7:433. doi:10.1186/1556-276X-7-433
Acknowledgements
This material is based on research sponsored by the Polish National Science Center (grant no. UMO-2011/01/M/NZ7/01445).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical statement
The authors confirm that the manuscript has been read and approved by all named authors and that there are no other persons who satisfied the criteria for authorship but are not listed. The authors further confirm that the order of authors listed in the manuscript has been approved by all of us.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Giełdoń, A., Witt, M.M., Gajewicz, A. et al. Rapid insight into C60 influence on biological functions of proteins. Struct Chem 28, 1775–1788 (2017). https://doi.org/10.1007/s11224-017-0957-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11224-017-0957-4