Abstract
Aims
Aluminium (Al) stress is one of the most adverse abiotic factors limiting the growth and productivity of crops in acidic soils. Fine-mapping and cloning of quantitative trait loci (QTLs) provides an effective tool in analysing the genetic mechanisms underlying Al tolerance and in breeding Al-tolerant soybean varieties.
Methods
Soybean cultivar Huachun2 in South China has been reported to be highly tolerant to multiple abiotic stresses in acidic soils, including Al stress. Here, we employ a recombinant inbred line (RIL) population derived from a cross of Huachun2 and Wayao to investigate the Al-tolerance QTLs. The prioritization method and qRT-PCR were applied to predict candidate genes in each QTL. Additionally, the functions of GmGSTU9 and GmPrx145 were investigated in transgenic soybean hairy roots.
Results
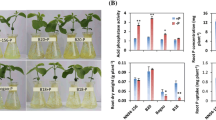
In total, five QTLs associated with relative root elongation and Al content were identified by using the high-density genetic map in hydroponics. GmGSTU9, which encodes a glutathione S-transferase gene in qAl06, and GmPrx145, which encodes a class III peroxidase gene in qAl-HC2, were selected to further study the gene functions by using transgenic soybean hairy roots. In transgenic soybean hairy roots, the MDA, H2O2 and O2− contents in GmGSTU9- and GmPrx145-overexpressing hairy roots were lower than those in the control and RNA-interference-exposed hairy roots under Al stress.
Conclusions
GmGSTU9 and GmPrx145 detected in qAl06 and qAl-HC2, respectively, positively regulate Al tolerance in soybean hairy roots by improving the antioxidant activity. These Al tolerance genes and molecular markers will be useful for marker-assisted selection to improve the Al tolerance of soybeans in acidic soils.







Similar content being viewed by others
Data availability
The data sets supporting the results of this study are included in the manuscript. Soybean seeds are available from the Guangdong Subcenter of the National Center for Soybean Improvement, PR China.
Abbreviations
- CIM:
-
composite interval mapping method
- ICP-AES:
-
Inductively coupled plasma-atomic emission spectrometry
- GST:
-
glutathione S-transferase
- LOD:
-
log-likelihood
- MAS:
-
marker-assisted selection
- NCBI:
-
National Center for Biotechnology Information
- qRT-PCR:
-
real-time quantitative polymerase chain reaction
- QTL:
-
quantitative trait loci
- RAD-seq:
-
restriction-site associated DNA sequencing
- RIL:
-
Recombinant inbred line
- RT-PCR:
-
reverse transcription-polymerase chain reaction
- SNP:
-
single nucleotide polymorphism
References
Abdel-Haleem H, Carter TE, Rufty TW et al (2014) Quantitative trait loci controlling aluminum tolerance in soybean: candidate gene and single nucleotide polymorphism marker discovery. Mol Breed 33(4):851–862
Alam SM (1981) Influence of aluminium on plant growth and mineral nutrition of barley. Commun Soil Sci Plant Anal 12(2):121–138
Amenós M, Corrales I, Poschenrieder C et al (2009) Different effects of aluminum on the actin cytoskeleton and brefeldin A-sensitive vesicle recycling in root apex cells of two maize varieties differing in root elongation rate and aluminum tolerance. Plant Cell Physiol 50(3):528–540
Bai L, Liu Y, Mu Y et al (2018) Heterotrimeric G-protein γ subunit CsGG3. 2 positively regulates the expression of CBF genes and chilling tolerance in cucumber. Front Plant Sci 9:488
Balakumar T, Sivaguru M, James MR et al (1992) Impact of aluminium toxicity on growth and efficiency of nutrient metabolism in some tropical rice cultivars. Trop Agric (Trinidad) 69:211–216
Bargsten JW, Nap JP, Sanchez-Perez GF et al (2014) Prioritization of candidate genes in QTL regions based on associations between traits and biological processes. BMC Plant Biol 14(1):330
Bela K, Horváth E, Gallé Á et al (2015) Plant glutathione peroxidases: emerging role of the antioxidant enzymes in plant development and stress responses. J Plant Physiol 176:192–201
Bianchi-Hall CM, Carter TE, Ruffy TW et al (1998) Heritability and resource allocation of aluminum tolerance derived from soybean PI 416937. Crop Sci 38(2):513–522
Bianchi-Hall CM, Carter TE, Bailey MA et al (2000) Aluminum tolerance associated with quantitative trait loci derived from soybean PI 416937 in hydroponics. Crop Sci 40(2):538–545
Cai Z, Cheng Y, Xian P et al (2018) Acid phosphatase gene GmHAD1 linked to low phosphorus tolerance in soybean, through fine mapping. Theor Appl Genet 131(8):1715–1728
Castrejón SE, Yatsimirsky AK (1997) Cyclodextrin enhanced fluorimetric determination of malonaldehyde by the thiobarbituric acid method. Talanta 44(6):951–957
Chen L, Cai Y, Liu X et al (2018) GmGRP-like gene confers Al tolerance in Arabidopsis. Sci Rep 8(1):13601
Cheng F, Cao G, Wang X et al (2008) The discovery and application of the efficient strains of soybean rhizobia in the acid low phosphorus soil in South China. Chin Sci Bull 23:2903–2910
Cheng Y, Ma Q, Ren H et al (2017) Fine mapping of a Phytophthora-resistance gene RpsWY in soybean (Glycine max L.) by high-throughput genome-wide sequencing. Theor Appl Genet 130(5):1041–1051
Ding ZJ, Yan JY, Xu XY et al (2013) WRKY 46 functions as a transcriptional repressor of ALMT 1, regulating aluminum-induced malate secretion in Arabidopsis. Plant J 76(5):825–835
Ding C, Chen T, Yang Y et al (2015) Molecular cloning and characterization of an S-adenosylmethionine synthetase gene from Chorispora bungeana. Gene 572(2):205–213
Dong D, Thornton I, Ramsey MH (1993) Influence of soil-extractable aluminium and pH on the uptake of aluminium from soil into the soybean plant (Glycine max). Environ Geochem Health 15(2–3):105–111
Eggert E, Obata T, Gerstenberger A et al (2016) A sucrose transporter-interacting protein disulphide isomerase affects redox homeostasis and links sucrose partitioning with abiotic stress tolerance. Plant Cell Environ 39(6):1366–1380
Ezaki B, Suzuki M, Motoda H et al (2004) Mechanism of gene expression of Arabidopsis glutathione S-transferase, AtGST1, and AtGST11 in response to aluminum stress. Plant Physiol 134(4):1672–1682
Fageria NK, Wright RJ, Ballgar VC (1988) Rice cultivar response to aluminum in nutrient solution. Commun Soil Sci Plant Anal 19(7–12):1133–1142
Foy CD (1984) Physiological Effects of Hydrogen, Aluminum, and Manganese Toxicities in Acid Soil 1. Soil acidity and liming, (soilacidityandl): 57–97
Garcia O, Dasilva WJ, Massei MAS. (1979) Efficient method for screening maize inbreds for aluminum tolerance. Maydica
Gratão PL, Polle A, Lea PJ et al (2005) Making the life of heavy metal-stressed plants a little easier. Funct Plant Biol 32(6):481–494
Grimme H. (1984) Aluminium tolerance of soybean plants as related to magnesium nutrition[C]//6. Colloque international pour l'optimisation de la nutrition des plantes, Montpellier (France), 2-8 Sep 1984. GERDAT
Gudys K, Guzy-Wrobelska J, Janiak A, et al. (2018) Prioritization of candidate genes in qtl regions for physiological and biochemical traits underlying drought response in barley (Hordeum vulgare L.). Frontiers in plant science, 9
Harper JE (1974) Soil and symbiotic nitrogen requirements for optimum soybean production 1. Crop Sci 14(2):255–260
Hiraga S, Sasaki K, Ito H et al (2001) A large family of class III plant peroxidases. Plant Cell Physiol 42(5):462–468
Hoekenga OA, Vision TJ, Shaff JE et al (2003) Identification and characterization of aluminum tolerance loci in Arabidopsis (Landsberg erecta × Columbia) by quantitative trait locus mapping. A physiologically simple but genetically complex trait. Plant Physiol 132(2):936–948
Horst WJ, Wang Y, Eticha D (2010) The role of the root apoplast in aluminium-induced inhibition of root elongation and in aluminium resistance of plants: a review. Ann Bot 106(1):185–197
Kale SM, Jaganathan D, Ruperao P et al (2015) Prioritization of candidate genes in “QTL-hotspot” region for drought tolerance in chickpea (Cicer arietinum L.). Sci Rep 5:15296
Kereszt A, Li D, Indrasumunar A, et al (2007) Agrobacterium rhizogenes-mediated transformation of soybean to study root biology. Nature protocols 2(4):948
Kopittke PM, Moore KL, Lombi E et al (2015) Identification of the primary lesion of toxic aluminum in plant roots. Plant Physiol 167(4):1402–1411
Kopittke PM, Menzies NW, Wang P et al (2016) Kinetics and nature of aluminium rhizotoxic effects: a review. J Exp Bot 67(15):4451–4467
Korir PC, Qi B, Wang Y et al (2011) A study on relative importance of additive, epistasis and unmapped QTL for Aluminium tolerance at seedling stage in soybean. Plant Breed 130(5):551–562
Korir PC, Zhang J, Wu K et al (2013) Association mapping combined with linkage analysis for aluminum tolerance among soybean cultivars released in yellow and Changjiang River valleys in China. Theor Appl Genet 126(6):1659–1675
Kumar J, Gupta DS, Gupta S et al (2017) Quantitative trait loci from identification to exploitation for crop improvement. Plant Cell Rep 36(8):1187–1213
Li X, Li Y, Qu M et al (2016) Cell wall pectin and its methyl-esterification in transition zone determine Al resistance in cultivars of pea (Pisum sativum). Front Plant Sci 7:39
Li H, Mollier A, Ziadi N et al (2017) Soybean root traits after 24 years of different soil tillage and mineral phosphorus fertilization management. Soil Tillage Res 165:258–267
Li GZ, Wang ZQ, Yokosho K et al (2018) Transcription factor WRKY 22 promotes aluminum tolerance via activation of OsFRDL 4 expression and enhancement of citrate secretion in rice (Oryza sativa). New Phytol 219(1):149–162
Little R (1988) Plant soil interactions at low pH problem solving-the genetic approach. Commun Soil Sci Plant Anal 19(7–12):1239–1257
Ma HX, Bai GH, Carver BF et al (2005) Molecular mapping of a quantitative trait locus for aluminum tolerance in wheat cultivar atlas 66. Theor Appl Genet 112(1):51–57
Ma Y, Li C, Ryan PR et al (2016) A new allele for aluminium tolerance gene in barley (Hordeum vulgare L.). BMC Genomics 17(1):186
Men Y, Wang D, Li B et al (2018) Effects of drought stress on the antioxidant system, osmolytes and secondary metabolites of Saposhnikovia divaricata seedlings. Acta Physiol Plant 40(11):191
Meng L, Wang B, Zhao X et al (2017) Association mapping of ferrous, zinc, and aluminum tolerance at the seedling stage in indica rice using MAGIC populations. Front Plant Sci 8:1822
Mugwira LM, Elgawhary SM, Patel SU (1978) Aluminium tolerance in triticale, wheat and rye as measured by root growth characteristics and aluminium concentration. Plant Soil 50(1–3):681–690
Pandey S, Ceballos H, Magnavaca R et al (1994) Genetics of tolerance to soil acidity in tropical maize. Crop Sci 34(6):1511–1514
Passardi F, Cosio C, Penel C et al (2005) Peroxidases have more functions than a Swiss army knife. Plant Cell Rep 24(5):255–265
Patil G, Do T, Vuong TD et al (2016) Genomic-assisted haplotype analysis and the development of high-throughput SNP markers for salinity tolerance in soybean. Sci Rep 6:19199
Patil G, Vuong TD, Kale S et al (2018) Dissecting genomic hotspots underlying seed protein, oil, and sucrose content in an interspecific mapping population of soybean using high-density linkage mapping. Plant Biotechnol J 16(11):1939–1953
Polle E, Konzak CF, Kattrick JA (1978) Visual detection of Aluminum tolerance levels in wheat by hematoxylin staining of seedling roots 1. Crop Sci 18(5):823–827
Qi B, Korir P, Zhao T et al (2008) Mapping quantitative trait loci associated with aluminum toxin tolerance in NJRIKY recombinant inbred line population of soybean (Glycine max). J Integr Plant Biol 50(9):1089–1095
Rhue RD, Grogan CO, Stockmeyer EW et al (1978) Genetic control of Aluminum tolerance in corn 1. Crop Sci 18(6):1063–1067
Richards KD, Schott EJ, Sharma YK et al (1998) Aluminum induces oxidative stress genes in Arabidopsis thaliana. Plant Physiol 116(1):409–418
Ryan PR, Ditomaso JM, Kochian LV (1993) Aluminium toxicity in roots: an investigation of spatial sensitivity and the role of the root cap. J Exp Bot 44(2):437–446
Ryan PR, Tyerman SD, Sasaki T et al (2010) The identification of aluminium-resistance genes provides opportunities for enhancing crop production on acid soils. J Exp Bot 62(1):9–20
Salinas J G, Gourley L M. (1987) Workshop on Evaluating Sorghum for Tolerance to Al-toxic Tropical Soils in Latin America (1984, Cali, Colombia). Sorghum for acid soils: Proceedings
Sánchez PA, Salinas JG (1981) Low-input technology for managing Oxisols and Ultisols in tropical America [M]//Advances in agronomy. Academic Press 34:279–406
Sartain J B. (1974) Differential effects of aluminum on top and root growth, nutrient accumulation and nodulation of several soybean varieties [D]. North Carolina State University at Raleigh
Schindler DW, Hecky RE (2009) Eutrophication: more nitrogen data needed. Science 324(5928):721–722
Sentelhas PC, Battisti R, Câmara GMS et al (2015) The soybean yield gap in Brazil–magnitude, causes and possible solutions for sustainable production. J Agric Sci 153(8):1394–1411
Sharma AD, Sharma H, Lightfoot DA (2011) The genetic control of tolerance to aluminum toxicity in the ‘Essex’ by ‘Forrest’ recombinant inbred line population. Theor Appl Genet 122(4):687–694
Shigeto J, Tsutsumi Y (2016) Diverse functions and reactions of class III peroxidases. New Phytol 209(4):1395–1402
Siecińska J, Nosalewicz A (2016) Aluminium toxicity to plants as influenced by the properties of the root growth environment affected by other co-stressors: a review [M]//reviews of environmental contamination and toxicology volume 243. Springer, Cham, 1–26
Tabaldi LA, Cargnelutti D, Gonçalves JF et al (2009) Oxidative stress is an early symptom triggered by aluminum in Al-sensitive potato plantlets. Chemosphere 76(10):1402–1409
Tao Y, Niu Y, Wang Y et al (2018) Genome-wide association mapping of aluminum toxicity tolerance and fine mapping of a candidate gene for Nrat1 in rice. PLoS One 13(6):e0198589
Valentine MF, De Tar JR, Mookkan M et al (2017) Silencing of soybean raffinose synthase gene reduced raffinose family oligosaccharides and increased true metabolizable energy of poultry feed. Front Plant Sci 8:692
Wang J, Raman H, Zhang G et al (2006) Aluminium tolerance in barley (Hordeum vulgare L.): physiological mechanisms, genetics and screening methods. J Zhejiang Univ Sci B 7(10):769–787
Wang X, Yan X, Liao H (2010) Genetic improvement for phosphorus efficiency in soybean: a radical approach. Ann Bot 106(1):215–222
Wang X, Pan Q, Chen F et al (2011) Effects of co-inoculation with arbuscular mycorrhizal fungi and rhizobia on soybean growth as related to root architecture and availability of N and P. Mycorrhiza 21(3):173–181
Wang S, Basten CJ, Zeng B (2012) Windows QTL cartographer 2.5. Department of Statistics, North Carolina State University, Raleigh, NC. http://statgen.ncsu.edu/qtlcart/WQTLCart.htm. Accessed 20 June 2018
Xie Z, Nolan TM, Jiang H et al (2019) AP2/ERF transcription factor regulatory networks in hormone and abiotic stress responses in Arabidopsis. Front Plant Sci 10
Yamamoto Y, Kobayashi Y, Devi SR et al (2002) Aluminum toxicity is associated with mitochondrial dysfunction and the production of reactive oxygen species in plant cells. Plant Physiol 128(1):63–72
Yao Y, He RJ, Xie QL et al (2017) ETHYLENE RESPONSE FACTOR 74 (ERF74) plays an essential role in controlling a respiratory burst oxidase homolog D (RbohD)-dependent mechanism in response to different stresses in Arabidopsis. New Phytol 213(4):1667–1681
Yu Y, Zhou W, Zhou K et al (2018) Polyamines modulate aluminum-induced oxidative stress differently by inducing or reducing H2O2 production in wheat. Chemosphere 212:645–653
Zhao LN, Zhao Q, Ao GM et al (2009) The foxtail millet Si69 gene is a Wali7 (wheat aluminum-induced protein 7) homologue and may function in aluminum tolerance. Chin Sci Bull 54(10):1697–1706
Zhao M, Song J, Wu A et al (2018) Mining beneficial genes for aluminum tolerance within a core collection of rice landraces through genome-wide association mapping with high density SNPs from specific-locus amplified fragment sequencing. Front Plant Sci 9
Zhu XF, Shi YZ, Lei GJ et al (2012) XTH31, encoding an in vitro XEH/XET-active enzyme, regulates aluminum sensitivity by modulating in vivo XET action, cell wall xyloglucan content, and aluminum binding capacity in Arabidopsis. Plant Cell 24(11):4731–4747
Zhu C, Luo N, He M et al (2014) Molecular characterization and expression profiling of the protein disulfide isomerase gene family in Brachypodium distachyon L. PLoS One 9(4):e94704
Acknowledgements
This work was supported by the Projects of Science and Technology of Guangzhou (201804020015); the National Key R&D Program of China (2018YFD0201006); the China Agricultural Research System (CARS-04-PS09) and the Research Project of the State Key Laboratory of Agricultural and Biological Resources Protection and Utilization in Subtropics.
Author information
Authors and Affiliations
Contributions
Y. C., T. L., Q. M. and H. N. provided the soybean materials used in this study. P. X., R. L., X. H., Q. L., and Z. C. performed the experiments and date analyses. Q. X. performed QTL mapping. Z. C. and H. N. prepared the manuscript. H. N. planned, supervised and financed this work, as well as edited the manuscript. All authors have read and approved the final version of the manuscript to be published.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Responsible Editor: Jian Feng Ma.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Cai, Z., Cheng, Y., Xian, P. et al. Fine-mapping QTLs and the validation of candidate genes for Aluminum tolerance using a high-density genetic map. Plant Soil 444, 119–137 (2019). https://doi.org/10.1007/s11104-019-04261-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-019-04261-0