Abstract
Background and aims
Considering the global demands in sustaining agriculture, use of organic amendments is gradually increasing. An improved understanding of the biological process is essential to evaluate the performance of organic amendments on agro-ecosystem.
Methods
Soils subjected to different fertilization regimes were collected from a field experiment. Microbial community compositions are assessed with 16S and ITS rRNA gene sequencing and subsequent bioinformatics analysis. Microbial functions are characterized with the geometric mean of the assayed enzyme activities (GMea) and the microbial carbon-use efficiency:nitrogen-use efficiency ratio (CUE:NUE).
Results
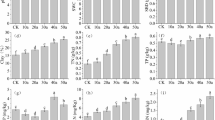
Compared with the chemically fertilized soil, the GMea significantly increased in organically amended soils. In contrast, the CUE:NUE was highest in chemically treated soil. These changes of microbial functional indicators were associated with shifts in the bacterial and not the fungal community composition, despite the fact that both the bacterial and fungal community compositions were significantly affected by the fertilization regimes. The abundances of specific soil bacterial taxa, especially the genera Luteimonas and Gemmatimona, were enriched by organic amendments. Soil organic carbon emerged as the major determinant of the bacterial community composition.
Conclusions
Soil microbial activities and nutrient-use efficiencies were dramatically changed along with the alteration of bacterial community composition. Relatively greater abundance of Luteimonas and Gemmatimona taxa in soils might be useful indicators for soil amelioration. Our research could be helpful to provide better strategies for the maintenance of soil fertility.






Similar content being viewed by others
References
Abarenkov K, Henrik Nilsson R, Larsson KH, Alexander IJ, Eberhardt U, Erland S, Høiland K, Kjøller R, Larsson E, Pennanen T (2010) The UNITE database for molecular identification of fungi–recent updates and future perspectives. New Phytol 186:281–285
Ai C, Liang GQ, Sun JW, Wang XB, Zhou W (2012) Responses of extracellular enzyme activities and microbial community in both the rhizosphere and bulk soil to long-term fertilization practices in a fluvo-aquic soil. Geoderma 173:330–338. https://doi.org/10.1016/j.geoderma.2011.07.020
Alden L, Demoling F, Baath E (2001) Rapid method of determining factors limiting bacterial growth in soil. Appl Environ Microbiol 67:1830–1838. https://doi.org/10.1128/AEM.67.4.1830-1838.2001
Banerjee S, Kirkby CA, Schmutter D, Bissett A, Kirkegaard JA, Richardson AE (2016) Network analysis reveals functional redundancy and keystone taxa amongst bacterial and fungal communities during organic matter decomposition in an arable soil. Soil Biol Biochem 97:188–198. https://doi.org/10.1016/j.soilbio.2016.03.017
Benitez E, Sainz H, Nogales R (2005) Hydrolytic enzyme activities of extracted humic substances during the vermicomposting of a lignocellulosic olive waste. Bioresour Technol 96:785–790. https://doi.org/10.1016/j.biortech.2004.08.010
Blackwood CB, Waldrop MP, Zak DR, Sinsabaugh RL (2007) Molecular analysis of fungal communities and laccase genes in decomposing litter reveals differences among forest types but no impact of nitrogen deposition. Environ Microbiol 9:1306–1316. https://doi.org/10.1111/j.1462-2920.2007.01250.x
Bowles TM, Acosta-Martínez V, Calderón F, Jackson LE (2014) Soil enzyme activities, microbial communities, and carbon and nitrogen availability in organic agroecosystems across an intensively-managed agricultural landscape. Soil Biol Biochem 68:252–262. https://doi.org/10.1016/j.soilbio.2013.10.004
Brookes PC, Landman A, Pruden G, Jenkinson DS (1985) Chloroform fumigation and the release of soil-nitrogen - a rapid direct extraction method to measure microbial biomass nitrogen in soil. Soil Biol Biochem 17:837–842. https://doi.org/10.1016/0038-0717(85)90144-0
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Pena AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336. https://doi.org/10.1038/nmeth.f.303
Cederlund H, Wessen E, Enwall K, Jones CM, Juhanson J, Pell M, Philippot L, Hallin S (2014) Soil carbon quality and nitrogen fertilization structure bacterial communities with predictable responses of major bacterial phyla. Appl Soil Ecol 84:62–68. https://doi.org/10.1016/j.apsoil.2014.06.003
Chen C, Zhang J, Lu M, Qin C, Chen Y, Yang L, Huang Q, Wang J, Shen Z, Shen Q (2016) Microbial communities of an arable soil treated for 8 years with organic and inorganic fertilizers. Biol Fertil Soils 52:455–467. https://doi.org/10.1007/s00374-016-1089-5
Curlevski NJA, Xu Z, Anderson IC, Cairney JWG (2010) Converting Australian tropical rainforest to native Araucariaceae plantations alters soil fungal communities. Soil Biol Biochem 42:14–20. https://doi.org/10.1016/j.soilbio.2009.08.001
DeForest JL (2009) The influence of time, storage temperature, and substrate age on potential soil enzyme activity in acidic forest soils using MUB-linked substrates and L-DOPA. Soil Biol Biochem 41:1180–1186. https://doi.org/10.1016/j.soilbio.2009.02.029
Demoling F, Figueroa D, Baath E (2007) Comparison of factors limiting bacterial growth in different soils. Soil Biol Biochem 39:2485–2495. https://doi.org/10.1016/j.soilbio.2007.05.002
Edgar RC (2013) UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10:996–998. https://doi.org/10.1038/nmeth.2604
Fan F, Yin C, Tang Y, Li Z, Song A, Wakelin SA, Zou J, Liang Y (2014) Probing potential microbial coupling of carbon and nitrogen cycling during decomposition of maize residue by 13C-DNA-SIP. Soil Biol Biochem 70:12–21. https://doi.org/10.1016/j.soilbio.2013.12.002
Fierer N, Bradford MA, Jackson RB (2007) Toward an ecological classification of soil bacteria. Ecology 88:1354–1364
Fierer N, Lauber CL, Ramirez KS, Zaneveld J, Bradford MA, Knight R (2012) Comparative metagenomic, phylogenetic and physiological analyses of soil microbial communities across nitrogen gradients. ISME J 6:1007–1017. https://doi.org/10.1038/ismej.2011.159
Fontaine S, Bardoux G, Abbadie L, Mariotti A (2004) Carbon input to soil may decrease soil carbon content. Ecol Lett 7:314–320. https://doi.org/10.1111/j.1461-0248.2004.00579.x
Francioli D, Schulz E, Lentendu G, Wubet T, Buscot F, Reitz T (2016) Mineral vs. organic amendments: microbial community structure, activity and abundance of agriculturally relevant microbes are driven by long-term fertilization strategies. Front Microbiol 7:1446. https://doi.org/10.3389/fmicb.2016.01446
García-Ruiz R, Ochoa V, Hinojosa MB, Carreira JA (2008) Suitability of enzyme activities for the monitoring of soil quality improvement in organic agricultural systems. Soil Biol Biochem 40:2137–2145. https://doi.org/10.1016/j.soilbio.2008.03.023
Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for basidiomycetes--application to the identification of mycorrhizae and rusts. Mol Ecol 2:113–118. https://doi.org/10.1111/j.1365-294X.1993.tb00005.x
Johnston AE, Poulton PR, Coleman K (2009) Soil organic matter: its importance in sustainable agriculture and carbon dioxide fluxes. Adv Agron 101:1–57. https://doi.org/10.1016/S0065-2113(08)00801-8
Kallenbach C, Grandy AS (2011) Controls over soil microbial biomass responses to carbon amendments in agricultural systems: a meta-analysis. Agric Ecosyst Environ 144:241–252. https://doi.org/10.1016/j.agee.2011.08.020
Kamolmanit B, Vityakon P, Kaewpradit W, Cadisch G, Rasche F (2013) Soil fungal communities and enzyme activities in a sandy, highly weathered tropical soil treated with biochemically contrasting organic inputs. Biol Fertil Soils 49:905–917. https://doi.org/10.1007/s00374-013-0785-7
Krepski ST, Hanson TE, Chan CS (2012) Isolation and characterization of a novel biomineral stalk-forming iron-oxidizing bacterium from a circumneutral groundwater seep. Environ Microbiol 14:1671–1680. https://doi.org/10.1111/j.1462-2920.2011.02652.x
Kuzyakov Y (2010) Priming effects: interactions between living and dead organic matter. Soil Biol Biochem 42:1363–1371. https://doi.org/10.1016/j.soilbio.2010.04.003
Lal R (2008) Soils and sustainable agriculture. A review. Agron Sustain Dev 28:57–64. https://doi.org/10.1051/agro:2007025
Lane D (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics
Lazcano C, Gómez-Brandón M, Revilla P, Domínguez J (2012) Short-term effects of organic and inorganic fertilizers on soil microbial community structure and function. Biol Fertil Soils 49:723–733. https://doi.org/10.1007/s00374-012-0761-7
Letunic I, Bork P (2016) Interactive tree of life (iTOL) v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res 44:W242–W245. https://doi.org/10.1093/nar/gkw290
Li F, Chen L, Zhang J, Yin J, Huang S (2017) Bacterial community structure after long-term organic and inorganic fertilization reveals important associations between soil nutrients and specific taxa involved in nutrient transformations. Front Microbiol 8:187. https://doi.org/10.3389/fmicb.2017.00187
Liu J, Sui Y, Yu Z, Shi Y, Chu H, Jin J, Liu X, Wang G (2014) High throughput sequencing analysis of biogeographical distribution of bacterial communities in the black soils of northeast China. Soil Biol Biochem 70:113–122. https://doi.org/10.1016/j.soilbio.2013.12.014
Loeppmann S, Blagodatskaya E, Pausch J, Kuzyakov Y (2016) Substrate quality affects kinetics and catalytic efficiency of exo-enzymes in rhizosphere and detritusphere. Soil Biol Biochem 92:111–118. https://doi.org/10.1016/j.soilbio.2015.09.020
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:550. https://doi.org/10.1186/s13059-014-0550-8
Maillard E, Angers DA (2014) Animal manure application and soil organic carbon stocks: a meta-analysis. Glob Chang Biol 20:666–679. https://doi.org/10.1111/gcb.12438
Manzoni S, Taylor P, Richter A, Porporato A, Agren GI (2012) Environmental and stoichiometric controls on microbial carbon-use efficiency in soils. New Phytol 196:79–91. https://doi.org/10.1111/j.1469-8137.2012.04225.x
Mooshammer M, Wanek W, Hammerle I, Fuchslueger L, Hofhansl F, Knoltsch A, Schnecker J, Takriti M, Watzka M, Wild B, Keiblinger KM, Zechmeister-Boltenstern S, Richter A (2014) Adjustment of microbial nitrogen use efficiency to carbon:nitrogen imbalances regulates soil nitrogen cycling. Nat Commun 5:3694. https://doi.org/10.1038/ncomms4694
Muyzer G, de Waal EC, Uitterlinden AG (1993) Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol 59:695–700
Nannipieri P, Ascher J, Ceccherini M, Landi L, Pietramellara G, Renella G (2003) Microbial diversity and soil functions. Eur J Soil Sci 54:655–670
Philippot L, Andersson SG, Battin TJ, Prosser JI, Schimel JP, Whitman WB, Hallin S (2010) The ecological coherence of high bacterial taxonomic ranks. Nat Rev Microbiol 8:523–529. https://doi.org/10.1038/nrmicro2367
Romanenko LA, Tanaka N, Svetashev VI, Kurilenko VV, Mikhailov VV (2012) Luteimonas vadosa sp. nov., isolated from seashore sediment. Int J Syst Evol Microbiol 63:1261–1266. https://doi.org/10.1099/ijs.0.043273-0
Saiya-Cork KR, Sinsabaugh RL, Zak DR (2002) The effects of long term nitrogen deposition on extracellular enzyme activity in an Acer saccharum forest soil. Soil Biol Biochem 34:1309–1315. https://doi.org/10.1016/S0038-0717(02)00074-3
Shanks OC, Kelty CA, Archibeque S, Jenkins M, Newton RJ, McLellan SL, Huse SM, Sogin ML (2011) Community structures of fecal bacteria in cattle from different animal feeding operations. Appl Environ Microbiol 77:2992–3001. https://doi.org/10.1128/AEM.02988-10
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504. https://doi.org/10.1101/gr.1239303
Smith J, Paul E (1990) The significance of soil microbial biomass estimations
Soman C, Li D, Wander MM, Kent AD (2016) Long-term fertilizer and crop-rotation treatments differentially affect soil bacterial community structure. Plant Soil 413:145–159. https://doi.org/10.1007/s11104-016-3083-y
Strickland MS, Rousk J (2010) Considering fungal:bacterial dominance in soils – methods, controls, and ecosystem implications. Soil Biol Biochem 42:1385–1395. https://doi.org/10.1016/j.soilbio.2010.05.007
Sul WJ, Asuming-Brempong S, Wang Q, Tourlousse DM, Penton CR, Deng Y, Rodrigues JLM, Adiku SGK, Jones JW, Zhou JZ, Cole JR, Tiedje JM (2013) Tropical agricultural land management influences on soil microbial communities through its effect on soil organic carbon. Soil Biol Biochem 65:33–38. https://doi.org/10.1016/j.soilbio.2013.05.007
Sun RB, Zhang XX, Guo XS, Wang DZ, Chu HY (2015) Bacterial diversity in soils subjected to long-term chemical fertilization can be more stably maintained with the addition of livestock manure than wheat straw. Soil Biol Biochem 88:9–18. https://doi.org/10.1016/j.soilbio.2015.05.007
Sun R, Dsouza M, Gilbert JA, Guo X, Wang D, Guo Z, Ni Y, Chu H (2016) Fungal community composition in soils subjected to long-term chemical fertilization is most influenced by the type of organic matter. Environ Microbiol 18:5137–5150. https://doi.org/10.1111/1462-2920.13512
Swift MJ, Heal OW, Anderson JM (1979) Decomposition in terrestrial ecosystems. Q Rev Biol 83:2772–2774
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729. https://doi.org/10.1093/molbev/mst197
Tian W, Wang L, Li Y, Zhuang KM, Li G, Zhang JB, Xiao XJ, Xi YG (2015) Responses of microbial activity, abundance, and community in wheat soil after three years of heavy fertilization with manure-based compost and inorganic nitrogen. Agric Ecosyst Environ 213:219–227. https://doi.org/10.1016/j.agee.2015.08.009
Tu C, Ristaino JB, Hu SJ (2006) Soil microbial biomass and activity in organic tomato farming systems: Effects of organic inputs and straw mulching. Soil Biol Biochem 38:247–255. https://doi.org/10.1016/j.soilbio.2005.05.002
Vilgalys R, Hester M (1990) Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several cryptococcus species. J Bacteriol 172:4238–4246
Voroney R, Winter J, Beyaert R (1993) Soil microbial biomass C and N. Soil sampling and methods of analysis. 277–286
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267
Wang X, Hong Q, Zhang C-F, Yang H-X, Hu G, Zhu S-J, Zhang Y-K, Liu X-W, Zhang H, Zhao C-R (2015a) Luteimonas soli sp. nov., isolated from farmland soil. Int J Syst Evol Microbiol 65:4809–4815. https://doi.org/10.1099/ijsem.0.000652
Wang Y, Ji H, Gao C (2015b) Differential responses of soil bacterial taxa to long-term P, N, and organic manure application. J Soils Sediments 16:1046–1058. https://doi.org/10.1007/s11368-015-1320-2
Wei WL, Yan Y, Cao J, Christie P, Zhang FS, Fan MS (2016) Effects of combined application of organic amendments and fertilizers on crop yield and soil organic matter: an integrated analysis of long-term experiments. Agric Ecosyst Environ 225:86–92. https://doi.org/10.1016/j.agee.2016.04.004
Wessén E, Hallin S, Philippot L (2010) Differential responses of bacterial and archaeal groups at high taxonomical ranks to soil management. Soil Biol Biochem 42:1759–1765. https://doi.org/10.1016/j.soilbio.2010.06.013
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR Protocols. Academic Press, San Diego
WRB IWG (2015) World reference base for soil resources 2014, update 2015 International soil classification system for naming soils and creating legends for soil maps. World Soil Resources Reports 106
Wu J, Joergensen RG, Pommerening B, Chaussod R, Brookes PC (1990) Measurement of soil microbial biomass C by fumigation extraction - an automated procedure. Soil Biol Biochem 22:1167–1169. https://doi.org/10.1016/0038-0717(90)90046-3
Zak DR, Tilman D, Parmenter RR, Rice CW, Fisher FM, Vose J, Milchunas D, Martin CW (1994) Plant-production and soil-microorganisms in late-successional ecosystems - a continental-scale study. Ecology 75:2333–2347. https://doi.org/10.2307/1940888
Zak DR, Blackwood CB, Waldrop MP (2006) A molecular dawn for biogeochemistry. Trends Ecol Evol 21:288–295. https://doi.org/10.1016/j.tree.2006.04.003
Zechmeister-Boltenstern S, Keiblinger KM, Mooshammer M, Peñuelas J, Richter A, Sardans J, Wanek W (2016) The application of ecological stoichiometry to plant–microbial–soil organic matter transformations. Ecol Monogr 85:133–155
Zhang L, Wang X, Yu M, Qiao Y, Zhang X-H (2015) Genomic analysis of Luteimonas abyssi XH031T: insights into its adaption to the subseafloor environment of South Pacific Gyre and ecological role in biogeochemical cycle. BMC Genomics 16. https://doi.org/10.1186/s12864-015-2326-2
Zhong Y, Yan W, Shangguan Z (2015) Impact of long-term N additions upon coupling between soil microbial community structure and activity, and nutrient-use efficiencies. Soil Biol Biochem 91:151–159. https://doi.org/10.1016/j.soilbio.2015.08.030
Zhong WH, Bian BY, Gao N, Min J, Shi WM, Lin XG, Shen WS (2016) Nitrogen fertilization induced changes in ammonia oxidation are attributable mostly to bacteria rather than archaea in greenhouse-based high N input vegetable soil. Soil Biol Biochem 93:150–159. https://doi.org/10.1016/j.soilbio.2015.11.003
Acknowledgements
This work was supported by the National Basic Research Program of China (2015CB150500), the National Key Research and Development Program of China (2017YFD0200206) and the Special Fund for Agro-scientific Research in the Public Interest (20150312205).
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Hans Lambers.
Electronic supplementary material
ESM 1
(DOCX 25 kb)
Rights and permissions
About this article
Cite this article
Guo, J., Liu, W., Zhu, C. et al. Bacterial rather than fungal community composition is associated with microbial activities and nutrient-use efficiencies in a paddy soil with short-term organic amendments. Plant Soil 424, 335–349 (2018). https://doi.org/10.1007/s11104-017-3547-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-017-3547-8