Abstract
Key message
A first creation of high oleic acid peanut varieties by using transcription activator-like effecter nucleases (TALENs) mediated targeted mutagenesis of Fatty Acid Desaturase 2 (FAD2).
Abstract
Transcription activator like effector nucleases (TALENs), which allow the precise editing of DNA, have already been developed and applied for genome engineering in diverse organisms. However, they are scarcely used in higher plant study and crop improvement, especially in allopolyploid plants. In the present study, we aimed to create targeted mutagenesis by TALENs in peanut. Targeted mutations in the conserved coding sequence of Arachis hypogaea fatty acid desaturase 2 (AhFAD2) were created by TALENs. Genetic stability of AhFAD2 mutations was identified by DNA sequencing in up to 9.52 and 4.11% of the regeneration plants at two different targeted sites, respectively. Mutation frequencies among AhFAD2 mutant lines were significantly correlated to oleic acid accumulation. Genetically, stable individuals of positive mutant lines displayed a 0.5–2 fold increase in the oleic acid content compared with non-transgenic controls. This finding suggested that TALEN-mediated targeted mutagenesis could increase the oleic acid content in edible peanut oil. Furthermore, this was the first report on peanut genome editing event, and the obtained high oleic mutants could serve for peanut breeding project.
Similar content being viewed by others
Introduction
Artificial sequence-specific nucleases, such as zinc-finger nucleases (ZFNs), transcription activator-like effector nucleases (TALENs) and CRISPR/cas system, can efficiently induce targeted mutagenesis, providing a powerful approach in plant biology research. All of those nucleases can make DNA double-stranded breaks (DSBs) at artificially defined genomic locus, subsequently relying on non-homologous end joining (NHEJ) and homology directed repair (HDR) to repair the damaged DNA at cleaved sites (Rodriguez-Leal et al. 2017). NHEJ directly rejoins the broken DNA end via an error-prone repair pathway, leading to deletions or insertions in homologous sections or frameshift mutation. In the HR pathway, homologous donor DNA functions as a template to provide precise information for DSB repair, facilitating DNA insertion or sequence replacement (Yin et al. 2017). TALENs consist of non-specific FokI cleavage domains fused with customizable TALE DNA binding repeat domains (Endo et al. 2016). The DNA binding central repeat domain of each TAL effector consists of 33.5 repeats, which are typically composed of 34 amino acids, and each repeat has a repeat-variable diresidue (RVD) at positions 12 and 13, specifically recognizing a single target nucleotide. TALEN-caused DSBs can be repaired by the NHEJ and HDR with distinct advantages. For example, over 30-bp target requirement results in less off-target effects, and no protospacer adjacent motif (PAM) restricts target arbitrary genome sequence compared with CRISPR/Cas9 system (Van Eck 2017).
As one of the five most important oilseed crops worldwide, peanut serves as an allotetraploid species (2n = 4x = 40, AABB), and about two-thirds of the global peanuts are utilized as edible oil in daily food commodity consumption (Chen et al. 2010; Clevenger et al. 2017). The peanut seed consists of around 50% oil, approximately 80% of which are oleic acids (36–67%) and linoleic acids (15–43%) (Pandey et al. 2014). Fatty acid composition of peanut oil is important for human health as well as shelf life of peanut products (Riveros et al. 2010). Higher content of polyunsaturated fatty acids (PUFAs) in oil leads to greater chance of oxidation, resulting in unpleasant tastes and short shelf life (Akhtar et al. 2014). Partial hydrogenation, which decreases PUFAs, reduces unsaturation and also generates trans fatty acids, which are not good for health (Broun et al. 1999; Zaloga et al. 2006). In plants, the conversion of oleic acid to linoleic acid is mainly catalyzed by the Δ12 fatty acid desaturase 2 (FAD2) (Okuley et al. 1994). FAD2 catalyzes the conversion of the monounsaturated fatty acid (oleic acid) to PUFA (linoleic acid) (Fig. 1a). In addition, genetic studies have revealed that high content of oleic acid in peanut is mainly regulated by two homozygous recessive mutants FAD2A and FAD2B (Janila et al. 2016). A previous study has demonstrated that the expression of ahFAD2B is significantly decreased or absent in high-oleate mutants, whereas both homeologous genes are expressed in normal oleate varieties (Chi et al. 2011). Therefore, it is a good strategy to create peanut lines with high level of oleic acid by reducing the activities of FAD2A and FAD2B genes. Conventional approaches include physical (X-rays or gama-raysetal) and chemical (MES or sodium) mutation breeding, which are time-consuming and not locus specific. RNAi-mediated suppression of FAD2 is subject to variation in transgene, and it requires the screening of a large number of transgenic events to identify candidates with stable phenotype over generations (McGinnis 2010).
TALENs targeting the AhFAD2 gene. a FAD2 is responsible for the conversion of oleic acid to linoleic acid. b Schematics of the AhFAD2 gene structures. The red triangle and green triangle represent the two potential TALEN target sites L1R1 and L2R2, respectively. c Structure of a TALEN binding to its target gene AhFAD2. The colored boxes denote the TAL effector repeats. Each color represents a different RVD. FokI endonuclease is fused to the C-terminal domains. NI, NG, HD and NN recognize A, T, C and G, respectively. Target sites are in red letters and underlined; the spacer region is in black letters between the target sites without underlined. d Structure of the last construct pCAMBIA1301-TALENs-FAD2 for Agrobacterium-induced peanut transformation
As an efficient genome editing tool, TALEN has been widely applied to enhance agronomic traits in crops. In previous studies, TALEN is used to improve the bacterial blight resistance in rice (Li et al. 2012; Blanvillain-Baufume et al. 2017), and it is also employed to reduce the lignin content in sugarcane via targeted mutagenesis of lignin biosynthetic gene COMT (Jung and Altpeter 2016). In addition, TALEN-mediated FAD2 mutants enhance the accumulation of oleic acid in soybean (Haun et al. 2014). In the present study, we aimed to create peanut varieties with high level of oleic acid using TALENs. TALENs were engineered to bind and simultaneously cleave specific DNA sequence in FAD2A and FAD2B genes. Our findings suggested that genome editing process could be tremendously accelerated in peanut by TALENs.
Materials and methods
Plasmid construction
TAL Effector-NuclecotideTargeter (TALE-NT) 2.0 program (Doyle et al. 2012) was employed to design TALENs, and FastTALE™ TALEN Assembly Kit (SIDANSAI) was used to construct TALEN vectors by one-step ligation. For peanut transformation, the final constructs were sub-cloned into a modified binary vector pCAMBIA 1301 for plant transformation. The constructs consisted of a TALEN expression cassette and a selectable transformation marker herbicide-resistant gene Bar.
Hairy root transformation of peanut
Each TALEN binary construct was independently transformed into Agrobacterium rhizogenes strain EHA105 for hairy root infection using a previously described protocol with minor modifications (Tiwari et al. 2015). Seeds of peanut (Arachis hypogaea) cv. YueYou NO. 7 were sequentially surface-sterilized by 75% (v/v) ethanol for 1 min and 5% (v/v) sodium hypochlorite solution for 5 min, and then they were rinsed with sterilized water for three times. The seeds were germinated in a growth chamber at 28 °C for 10 days under a 16-h light/8-h dark cycle. Excised leaves of 10-day-old peanut seedlings were used as explant materials for co-cultivation with Agrobacterium rhizogenes strain EHA105. The Agrobacterium solution (cultured until OD600 reached 0.8) was centrifuged and then suspended in liquid inoculation medium (MS salt and vitamins containing 30 g/L sucrose) to an OD600 of 0.5. Approximately 2 weeks after transformation, putative transgenic hairy roots were observed from the wound sites.
Peanut transformation
Transformation of peanut was conducted using a non-tissue culture-based transformation protocol (Sharma and Anjaiah 2000) using Agrobacterium tumefaciens strain EHA105. Embryo with one of the cotyledons from surface-sterilized seed of peanut cv. Yueyou 7 was cut off at the site attached to the primary axis as explant. The infection was performed at 28 °C for 18 h with gentle agitation. The seedlings were blot-dried and placed on autoclaved soil in capped bottles for germination under aseptic conditions. After 3 days, the seedlings were thoroughly washed with 300 µg/mL of cefotaxime and then blot-dried again. After 3–4 days, the seedlings were transferred to soilrite in pots under growth conditions for at least 2 weeks before transferred to greenhouse. The growth chamber was maintained at 28–30 °C under a 16-h light/8-h dark cycle with fluorescent light at an intensity of 35 µmol/m2 s.
Genomic DNA extraction and real-time PCR
Genomic DNA was extracted from approximately 1 g of leaf tissue or 300 mg of hairy roots using the DNeasy Plant Mini Kit (TIANGEN). To identify mutation in targeted genes by PCR, 50–100 ng of purified genomic DNA was used as template in the amplification.
Total RNA was extracted from 100 mg blade leaf of wild-type plant using Trizol Reagent (Invitrogen, Beijing, China). Concentration and integrity of purified RNA were assessed using a UV–visible spectrophotometer DNA NanoDrop (Thermo Fisher). RNase-free DNaseI (Fermentas, USA) was used to remove genomic DNA contaminants, and 1 µg of purified RNA was reversely transcribed into cDNA using PrimeScript RT reagent Kit (Takara, Dalian, China) according to the manufacturer’s instructions. PCR reaction was conducted in a 20-µL reaction system using SYBR Premix ExTaq™ (TaKaRa, Dalian, China) on an ABI StepOne Plus system. The relative expressions of target genes were calculated by the 2− ΔΔCT method and shown as fold changes relative to the wild-type plants. Ah18S was selected as the housekeeping gene. Each measurement was carried out in triplicate with three biological replicates, and data were expressed as means ± SE.
Profile analysis of fatty acids
Total fatty acid content was analyzed using 5 g of seeds collected from each FAD2 mutant lines. The fresh seeds of different stages were ground into a fine powder in liquid nitrogen. The total oil content was calculated using n-hexane following Soxhlet extraction method. The profile analysis of fatty acids in peanut seeds was performed by gas chromatography of methyl ester of fatty acid according to the National Standard of the People’s Republic of China (GB/T 17377-2008). Briefly, 100 mg peanut seeds were frozen and ground into a fine powder in liquid N2. The obtained powder was mixed with 1.5 mL chloroform/methanol (2:1, v/v) and 100 µL internal standard solution C17:0 (1 mg/mL). The mixture was extracted by ultrasound-assisted extraction for 30 min and centrifuged at 12,000 rpm for 6 min, and then 1 mL supernatant was transferred into a fresh tube. The fatty acid extracting solution was mixed with 0.2 mL KCL (75%, w/v) solution and centrifuged at 12,000 rpm for 6 min, and then 400 µL substratum chloroform extract phase was transferred into a new glass tube and prepared for methanol esterification. The extracting solution was mixed with 2 mL sulfuric acid methanol (5:95, v/v) and incubated in a water bath at 85 °C for 1.5 h. Subsequently, 1 mL H2O and 1 mL hexane were added into the tube, the mixture was centrifuged at 5000 rpm for 5 min, and 500 µL superstratum hexane extract phase was used for gas chromatography (Cat#YLSB076). The individual fatty acid contents were reported as the relative percentages of oleic acid and linoleic acid in the extracted oil.
Results
TALEN design and construction
To improve the quality of peanut oil, sequence-specific TALENs were designed to simultaneously inactivate (knockout) all alleles of the ahFAD2A and ahFAD2B genes in peanut. Both ahFAD2A and ahFAD2B encode enzymes of similar functions. TALENs’ arrays were designed by the TAL Effector-Nucleotide Targeter (TALE-NT) 2.0 (https://talent.cac.cornell.edu/), and we selected two potential TALEN target sites that were located in the 100-bp conserved domain of the both gene coding sequences following the start codon (Fig. 1b). TALEN repeat arrays were constructed using the FastTALETM TALEN Assembly Kit (SIDANSAI Biotechnology (Shanghai) CO., LTD) following the protocol (Fig. 1c). Plasmids L24 and R15 were selected to construct the left array and right array, respectively (Fig. S1). After the assembly was completed, we sequenced the complete repeat arrays of the left and right TALEN’s arms from forward direction. The sequencing results indicated that all of the designed repeat arrays were successfully assembled to recognize the TALEN target sites. Furthermore, to obtain the stable genetic transformation, the last TALEN pairs were sub-cloned into the plant binary vector pCAMBIA1301, in which the 35S promoter was used to drive the expressions of TALE binding domain and FokI, and the herbicide-resistant gene Bar served as a selective marker to replace the original hygromycin in the pCAMBIA1301 backbone (Fig. 1d). The pCAMBIA1301-TALENs were subsequently transformed into Agrobacterium strain EHA105 by the liquid nitrogen freeze/thaw method, and kanamycin selection was performed to select positive colonies.
Activity assessment of TALENs in hairy roots
The Agrobacterium rhizogenes hairy-root transformation method was used to assess the activity and efficiency of targeted mutagenesis by TALENs in peanut, by which transgenic hairy roots could be easily obtained within 4 weeks to provide an effective means to rapidly screen TALEN function at endogenous targets. A total of 321 regenerated roots were inoculated by two TALEN vectors, and PCR assays were carried out to detect mutations using the extracted genomic DNA. The number of hairy roots with mutations was divided by the total number of hairy roots inoculated by each vector in order to determine the mutagenesis frequency. In this assay, the mutagenesis frequency of transgenic peanut roots ranged from 8.33 to 12.38% (Table 1). In our case, most mutations were small deletions ranging from 1 to 10 bp (Fig. 2).
Generation of TALEN-induced FAD2 mutant lines
To obtain the stable genetic transformation, TALEN expression cassettes were introduced into peanut by Agrobacterium-mediated transformation via direct embryogenesis. TALEN integration was validated by PCR in regenerated plants, and a total of 63 and 72 independent TALEN-integrated FAD2 mutant lines were generated at the target site 1 and site 2 in the T0 generation, respectively (Table 2). In addition, we assessed the expression of the selective marker Bar gene by PCR, and 19 putative transformants exhibited the expression of Bar gene (500 bp). A representative agarose-gel electrophoresis suggested that the transgenic plants expressing the Bar gene were successfully established (Fig. S3). Finally, we obtained six and three FAD2 mutant lines with genetic stability in the T1 generation (Fig. S4), and the mutation frequency (T1/T0) was 9.52 and 4.11% at two different TALEN target sites, respectively (Table 2; Fig. 3a). To corroborate the correctness of those positive transgenic plants, we subsequently examined the relative expression of FAD2 in transgenic and wild-type plants by real-time PCR. The results indicated that the relative expression of FAD2 in TALEN-induced mutant lines was significantly decreased compared with the wild-type plants (Fig. 3b, c).
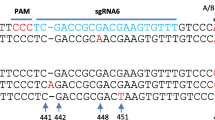
Nucleotide sequences of TALEN-induced mutations in genetic stability of the transgenic peanut at T1 generation. a The TALEN-binding sequences are in red letters, while deletions and insertions are indicated by dashes and green letters, respectively. b, c, Relative expression of FAD2 in wild-type, L1R1 and L2R2 plants. Values are mean ± SD of three biological replicates, and asterisks indicate a significant difference according to the t-test (P < 0.05) compared with wild-type
Fatty acid profiles of mutant seeds
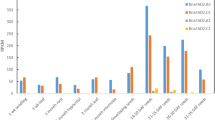
To evaluate whether the TALEN-induced FAD2 mutants had a great effect on peanut agronomic traits, the seeds of T2 generation were collected from the positive transgenic plants, and then their fatty acid composition was analyzed. The percentage of total fatty acid and protein content of wild-type plants was about 51 and 25%, respectively, and the transgenic seeds showed the similar characteristics. The composition of total fatty acid included approximately 43% oleic acid and 35.5% linoleic acid in wild-type seeds (Fig. 4a, b). However, the percentage of oleic acid was significantly increased in seeds of nine independent FAD2 mutant lines, ranging from 60 to 80% with a maximum value of 80.45% and an average increasing index up to 0.5–2 fold (Fig. 4c, d). On the contrary, the level of linoleic acid in fad2 T2 transgenic seeds was decreased by 3–19% compared with wild-type seeds (Fig. 4c, d).
The fatty acid profile and agronomic characters of TALEN-induced FAD2 mutant lines. a, b The percentages of total fatty acid and protein content in wild-type, L1R1 and L2R2 mutant lines in the T2 generation. Green bar indicates the wild-type, red bar indicates the L1R1, and blue bar represents the L2R2. Error bars represent the standard deviation from the mean (n = 3). c, d Relative contents of oleic acid and linoleic acid in wild-type, L1R1 and L2R2 plants. Error bars represent the standard deviation from the mean (n = 3). The asterisks indicate a significant difference according to the t-test (P < 0.05) compared with wild-type
In addition, we investigated multiple agronomic characters of 15 independent lines in wild-type and transgenic plants in order to further clarify whether the TALEN-mediated FAD2 mutants had a major influence on the plant growth (Fig. 5a). The results indicated that the pod number per plant was slightly decreased from 37 in wild-type plants to 31 and 33 in L1R1 and L2R2 transgenic lines, respectively (Fig. 5c). However, other agronomic traits, such as plant height, mature pod weight and kernel weight, were not altered when the whole plant was grown in the field, which were consistent with wild-type plants (Fig. 5b, c, d). This finding suggested that the FAD2 mutant was able to improve the accumulation of oleic acid, but it had a little influence on other agronomic characters, especially on per plant yield of peanut. Moreover, the TALEN-induced targeted mutagenesis of FAD2 was successfully established in this experiment.
Phenotype of the wild-type and FAD2 mutant lines. a The photograph of the wild-type and FAD2 mutant lines in T1 generation. b–d, The main agronomic traits in L1R1 and L2R2 mutant lines compared with wild-type plant in the T2 generation. The scale bars represents the standard deviation from the mean (n = 15), and asterisks indicate a significant difference according to the t-test (P < 0.05) compared with wild-type
Discussion
As a powerful tool, TALEN-mediated genome modification is widely used in genome engineering. Although TALE DNA binding array consists of alternative modules, it requires context-dependent specificity, and the synthesis of novel TALE arrays is time-consuming and costly (Tong et al. 2012). A single TALEN contains an N-terminal domain composed of a nuclear localization signal, a central domain typically containing tandem TALE repeats for the recognition of a specific DNA sequence (for natural TALEN, it is usually 14–20 bp), and a C-terminal domain of the functional endonuclease FokI. FokI usually functions in a dimeric fashion. Therefore, a pair of TALENs is required to make a cut at particular site of the genome. The pair of designed TALENs binds to their target DNA sequences flanking a spacer DNA, facilitating the FokI heterodimerization (Malzahn et al. 2017).
Genome editing plays critical roles in accelerating basic plant investigation and crop improvement (Mahfouz et al. 2014). Peanut oil with high level of oleic acid is desirable due to its longer shelf life and enhanced heat stability (Vassiliou et al. 2009). For targeting FAD2 gene, two TALEN target sites were selected at the same conserved region between FAD2-A and FAD2-B in order to simultaneously mutagenize both genes. All TALEN pairs showed good activity in transgenic peanut roots, while the mutation efficiencies were different (Table 1). L1R1 and L2R2 had different spacer lengths between TALEN targets, and such difference was mainly affected by the functional endonuclease FokI. In the present study, we employed TALENs to improve peanut oil quality by targeting FAD2 genes. Although the vector construction was costly and laborious, we, for the first time, successfully established targeted mutagenesis in peanut by TALENs. Valuably, the level of monounsaturated fatty (oleic acid) in transgenic seeds was obviously increased with the maximum value as high as 90.45%, and oil characteristic was altered by reducing PUFAs (linoleic) to 7.42%. Our findings offered a potential solution for the market demand of peanut oil with high level of oleic acid. TALEN-induced FAD2 mutants could be used to efficiently improve the quality of edible peanut oil. Actually, at least six FAD2 genes have been identified in peanut genome (Wang et al. 2015), and TALEN-mediated targeted mutation simultaneously occurred in the sequences of FAD2-2A (GenBank: KF741365.1) and FAD2-2B (GenBank: KF741366.1) in this study. However, the complete genome sequence of cultivated peanut was not available, therefore the detection of the potential off-target sites would be difficult.
Furthermore, a low mutation frequency was observed when the T1 plants were regenerated from the TALEN-induced T0 knockout plants under the normal circumstance. We supposed that this situation could be attributed to that peanut was an allotetraploid and cleistogamous species, and TALEN-induced DNA damage might be recovered when the microsporocyte undergoes simultaneous homologous chromosome synapsis and meiotic division stage (Ji et al. 2013; Haun et al. 2014). Additionally, the unstable chimera induced by TALEN in T0 generation was an unignorable problem for low frequency (Tian and Marcotrigiano 1993; Frank and Chitwood 2016). Indeed, genome editing techniques largely depend on genetic transformation and genetic stability of plants. Transformation efficiency in peanut remains still low compared with other crops. However, the genome editing efficiency in peanut might be enhanced by other potential sequence-specific nucleases, including ZFNs, CRISPR/Cas and CRISPR/Cpf1 systems (Lowder et al. 2016). Taken together, the TALEN-mediated targeted mutagenesis of FAD2 possessed a clear advantage over conventional breeding method, by which the genome editing process could be tremendously accelerated in peanut.
References
Akhtar S, Khalid N, Ahmed I, Shahzad A, Suleria HA (2014) Physicochemical characteristics, functional properties, and nutritional benefits of peanut oil: a review. Crit Rev Food Sci Nutr 54:1562–1575. https://doi.org/10.1080/10408398.2011.644353
Blanvillain-Baufume S, Reschke M, Sole M, Auguy F, Doucoure H, Szurek B, Meynard D, Portefaix M, Cunnac S, Guiderdoni E, Boch J, Koebnik R (2017) Targeted promoter editing for rice resistance to Xanthomonas oryzae pv. oryzae reveals differential activities for SWEET14-inducing TAL effectors. Plant Biotechnol J 15:306–317. https://doi.org/10.1111/pbi.12613
Broun P, Gettner S, Somerville C (1999) Genetic engineering of plant lipids. Annu Rev Nutr 19:197–216. https://doi.org/10.1146/annurev.nutr.19.1.197
Chen Z, Wang ML, Barkley NA, Pittman RN (2010) A simple allele-specific PCR assay for detecting FAD2 alleles in both A and B genomes of the cultivated peanut for high-oleate trait selection. Plant Mol Biol Rep 28:542–548. https://doi.org/10.1007/s11105-010-0181-5
Chi X, Yang Q, Pan L, Chen M, He Y, Yang Z, Yu S (2011). Isolation and characterization of fatty acid desaturase genes from peanut (Arachis hypogaea L.). Plant Cell Rep 30:1393–1404. https://doi.org/10.1007/s00299-011-1048-4
Clevenger J, Chu Y, Chavarro C, Agarwal G, Bertioli DJ, Leal-Bertioli S, Pandey MK, Vaughn J, Abernathy B, Barkley NA, Hovav R, Burow M, Nayak SN, Chitikineni A, Isleib TG, Holbrook CC, Jackson SA, Varshney RK, Ozias-Akins P (2017) Genome-wide SNP genotyping resolves signatures of selection and tetrasomic recombination in peanut. Mol Plant 10:309–322. https://doi.org/10.1016/j.molp.2016.11.015
Doyle EL, Booher NJ, Standage DS, Voytas DF, Brendel VP, Vandyk JK, Bogdanove AJ (2012) TAL effector-nucleotide targeter (TALE-NT) 2.0: tools for TAL effector design and target prediction. Nucleic Acids Res 40:W117-22. https://doi.org/10.1093/nar/gks608
Endo M, Nishizawa-Yokoi A, Toki S (2016). Targeted mutagenesis in rice using TALENs and the CRISPR/Cas9 system. Methods Mol Biol 1469:123–135. https://doi.org/10.1007/978-1-4939-4931-1_9
Frank MH, Chitwood DH (2016) Plant chimeras: the good, the bad, and the ‘Bizzaria’. Dev Biol 419:41–53. https://doi.org/10.1016/j.ydbio.2016.07.003
Haun W, Coffman A, Clasen BM, Demorest ZL, Lowy A, Ray E, Retterath A, Stoddard T, Juillerat A, Cedrone F, Mathis L, Voytas DF, Zhang F (2014). Improved soybean oil quality by targeted mutagenesis of the fatty acid desaturase 2 gene family. Plant Biotechnol J 12:934–940. https://doi.org/10.1111/pbi.12201
Janila P, Pandey MK, Shasidhar Y, Variath MT, Sriswathi M, Khera P, Manohar SS, Nagesh P, Vishwakarma MK, Mishra GP, Radhakrishnan T, Manivannan N, Dobariya KL, Vasanthi RP, Varshney RK (2016) Molecular breeding for introgression of fatty acid desaturase mutant alleles (ahFAD2A and ahFAD2B) enhances oil quality in high and low oil containing peanut genotypes. Plant Sci 242:203–213. https://doi.org/10.1016/j.plantsci.2015.08.013
Ji J, Tang D, Wang M, Li Y, Zhang L, Wang K, Li M, Cheng Z (2013). MRE11 is required for homologous synapsis and DSB processing in rice meiosis. Chromosoma 122:363–376. https://doi.org/10.1007/s00412-013-0421-1
Jung JH, Altpeter F (2016). TALEN mediated targeted mutagenesis of the caffeic acid O-methyltransferase in highly polyploid sugarcane improves cell wall composition for production of bioethanol. Plant Mol Biol 92:131–142. https://doi.org/10.1007/s11103-016-0499-y
Li T, Liu B, Spalding MH, Weeks DP, Yang B (2012) High-efficiency TALEN-based gene editing produces disease-resistant rice. Nat Biotechnol 30:390–392. https://doi.org/10.1038/nbt.2199
Lowder L, Malzahn A, Qi Y (2016) Rapid evolution of manifold CRISPR systems for plant genome editing. Front Plant Sci 7:1683. https://doi.org/10.3389/fpls.2016.01683
Mahfouz MM, Piatek A, Stewart CJ (2014) Genome engineering via TALENs and CRISPR/Cas9 systems: challenges and perspectives. Plant Biotechnol J 12:1006–1014. https://doi.org/10.1111/pbi.12256
Malzahn A, Lowder L, Qi Y (2017) Plant genome editing with TALEN and CRISPR. Cell Biosci 7:21. https://doi.org/10.1186/s13578-017-0148-4
McGinnis KM (2010) RNAi for functional genomics in plants. Brief Funct Genom 9:111–117. https://doi.org/10.1093/bfgp/elp052
Okuley J, Lightner J, Feldmann K, Yadav N, Lark E, Browse J (1994). Arabidopsis FAD2 gene encodes the enzyme that is essential for polyunsaturated lipid synthesis. Plant Cell 6:147–158. https://doi.org/10.1105/tpc.6.1.147
Pandey MK, Wang ML, Qiao L, Feng S, Khera P, Wang H, Tonnis B, Barkley NA, Wang J, Holbrook CC, Culbreath AK, Varshney RK, Guo B (2014) Identification of QTLs associated with oil content and mapping FAD2 genes and their relative contribution to oil quality in peanut (Arachis hypogaea L.). BMC Genet 15:133. https://doi.org/10.1186/s12863-014-0133-4
Riveros CG, Mestrallet MG, Gayol MF, Quiroga PR, Nepote V, Grosso NR (2010) Effect of storage on chemical and sensory profiles of peanut pastes prepared with high-oleic and normal peanuts. J Sci Food Agric 90:2694–2699. https://doi.org/10.1002/jsfa.4142
Rodriguez-Leal D, Lemmon ZH, Man J, Bartlett ME, Lippman ZB (2017) Engineering quantitative trait variation for crop improvement by genome editing. Cell 171:470–480.e8. https://doi.org/10.1016/j.cell.2017.08.030
Sharma KK, Anjaiah VV (2000) An efficient method for the production of transgenic plants of peanut (Arachis hypogaea L.) through Agrobacterium tumefaciens-mediated genetic transformation. Plant Sci 159:7–19
Tian HC, Marcotrigiano M (1993). Origin and development of adventitious shoot meristems initiated on plant chimeras. Dev Biol 155:259–269. https://doi.org/10.1006/dbio.1993.1023
Tiwari V, Chaturvedi AK, Mishra A, Jha B (2015). An efficient method of agrobacterium-mediated genetic transformation and regeneration in local Indian cultivar of groundnut (Arachis hypogaea) using grafting. Appl Biochem Biotechnol 175:436–453. https://doi.org/10.1007/s12010-014-1286-3
Tong C, Huang G, Ashton C, Wu H, Yan H, Ying QL (2012). Rapid and cost-effective gene targeting in rat embryonic stem cells by TALENs. J Genet Genom 39:275–280. https://doi.org/10.1016/j.jgg.2012.04.004
Van Eck J (2017) Genome editing and plant transformation of solanaceous food crops. Curr Opin Biotechnol 49:35–41. https://doi.org/10.1016/j.copbio.2017.07.012
Vassiliou EK, Gonzalez A, Garcia C, Tadros JH, Chakraborty G, Toney JH (2009) Oleic acid and peanut oil high in oleic acid reverse the inhibitory effect of insulin production of the inflammatory cytokine TNF-alpha both in vitro and in vivo systems. Lipids Health Dis 8:25. https://doi.org/10.1186/1476-511X-8-25
Wang Y, Zhang XG, Zhao YL, Prakash CS, He GH, Yin DM (2015) Insights into the novel members of the FAD2 gene family involved in high-oleate fluxes in peanut. Genome 58:375–383. https://doi.org/10.1139/gen-2015-0008
Yin K, Gao C, Qiu JL (2017) Progress and prospects in plant genome editing. Nat Plants 3:17107. https://doi.org/10.1038/nplants.2017.107
Zaloga GP, Harvey KA, Stillwell W, Siddiqui R (2006). Trans fatty acids and coronary heart disease. Nutr Clin Pract 21:505–512. https://doi.org/10.1177/0115426506021005505
Acknowledgements
This study was supported by the Science and Technology Planning Project of Guangdong Province (2015B020231006, 2016B020201003 and 2016LM3161), the National Natural Science Foundation of China (31501246), and the Modern Agro-industry Technology Research System (CARS-14).
Author information
Authors and Affiliations
Contributions
HL, QL, and YH conceived the original screening and research plans. XC supervised the experiments. SW, XL performed most of the experiments. HL designed the experiments and analyzed the data. QL conceived the project and wrote the article with contributions of all the authors. SW, HL supervised and complemented the writing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wen, S., Liu, H., Li, X. et al. TALEN-mediated targeted mutagenesis of fatty acid desaturase 2 (FAD2) in peanut (Arachis hypogaea L.) promotes the accumulation of oleic acid. Plant Mol Biol 97, 177–185 (2018). https://doi.org/10.1007/s11103-018-0731-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-018-0731-z