Abstract
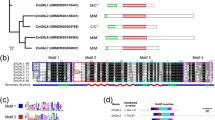
In this work, we retrace the evolutionary history of plant double-stranded RNA binding proteins (DRBs), a group of non-catalytic factors containing one or more double-stranded RNA binding motif (dsRBM) that play important roles in small RNA biogenesis and functions. Using a phylogenetic approach, we show that multiple dsRBM DRBs are systematically composed of two different types of dsRBMs evolving under different constraints and likely fulfilling complementary functions. In vascular plants, four distinct clades of multiple dsRBM DRBs are always present with the exception of Brassicaceae species, that do not possess member of the newly identified clade we named DRB6. We also identified a second new and highly conserved DRB family (we named DRB7) whose members possess a single dsRBM that shows concerted evolution with the most C-terminal dsRBM domain of the Dicer-like 4 (DCL4) proteins. Using a BiFC approach, we observed that Arabidopsis thaliana DRB7.2 (AtDRB7.2) can directly interact with AtDRB4 but not with AtDCL4 and we provide evidence that both AtDRB7.2 and AtDRB4 participate in the epigenetically activated siRNAs pathway.







Similar content being viewed by others
References
Adenot X, Elmayan T, Lauressergues D, Boutet S, Bouche N, Gasciolli V, Vaucheret H (2006) DRB4-dependent TAS3 trans-acting siRNAs control leaf morphology through AGO7. Curr Biol 16:927–932. doi:10.1016/j.cub.2006.03.035
Anisimova M, Gascuel O (2006) Approximate likelihood-ratio test for branches: a fast, accurate, and powerful alternative. Syst Biol 55:539–552
Azimzadeh J, Nacry P, Christodoulidou A, Drevensek S, Camilleri C, Amiour N, Parcy F, Pastuglia M, Bouchez D (2008) Arabidopsis TONNEAU1 proteins are essential for preprophase band formation and interact with centrin. Plant Cell 20:2146–2159. doi:10.1105/tpc.107.056812
Bernstein E, Caudy AA, Hammond SM, Hannon GJ (2001) Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 409:363–366. doi:10.1038/35053110
Blanc G, Hokamp K, Wolfe KH (2003) A recent polyploidy superimposed on older large-scale duplications in the Arabidopsis genome. Genome Res 13:137–144. doi:10.1101/gr.751803
Borsani O, Zhu J, Verslues PE, Sunkar R, Zhu JK (2005) Endogenous siRNAs derived from a pair of natural cis-antisense transcripts regulate salt tolerance in Arabidopsis. Cell 123:1279–1291
Bouche N, Lauressergues D, Gasciolli V, Vaucheret H (2006) An antagonistic function for Arabidopsis DCL2 in development and a new function for DCL4 in generating viral siRNAs. EMBO J 25:3347–3356. doi:10.1038/sj.emboj.7601217
Cerutti H, Ma X, Msanne J, Repas T (2011) RNA-mediated silencing in algae: biological roles and tools for analysis of gene function. Eukaryot Cell 10:1164–1172. doi:10.1128/EC.05106-11
Chang KY, Ramos A (2005) The double-stranded RNA-binding motif, a versatile macromolecular docking platform. FEBS J 272:2109–2117. doi:10.1111/j.1742-4658.2005.04652.x
Chendrimada TP, Gregory RI, Kumaraswamy E, Norman J, Cooch N, Nishikura K, Shiekhattar R (2005) TRBP recruits the Dicer complex to Ago2 for microRNA processing and gene silencing. Nature 436:740–744. doi:10.1038/nature03868
Clavel M et al (2015) Parallel action of AtDRB2 and RdDM in the control of transposable element expression. BMC Plant Biol 15:70. doi:10.1186/s12870-015-0455-z
Creasey KM, Zhai J, Borges F, Van Ex F, Regulski M, Meyers BC, Martienssen RA (2014) miRNAs trigger widespread epigenetically activated siRNAs from transposons in Arabidopsis. Nature 508:411–415. doi:10.1038/nature13069
Curtin SJ, Watson JM, Smith NA, Eamens AL, Blanchard CL, Waterhouse PM (2008) The roles of plant dsRNA-binding proteins in RNAi-like pathways. FEBS Lett 582:2753–2760. doi:10.1016/j.febslet.2008.07.004
Daniels SM et al (2009) Characterization of the TRBP domain required for dicer interaction and function in RNA interference. BMC Mol Biol 10:38. doi:10.1186/1471-2199-10-38
Denli AM, Tops BB, Plasterk RH, Ketting RF, Hannon GJ (2004) Processing of primary microRNAs by the Microprocessor complex. Nature 432:231–235. doi:10.1038/nature03049
Dugre-Brisson S, Elvira G, Boulay K, Chatel-Chaix L, Mouland AJ, DesGroseillers L (2005) Interaction of Staufen1 with the 5′ end of mRNA facilitates translation of these RNAs. Nucleic Acids Res 33:4797–4812. doi:10.1093/nar/gki794
Dunoyer P, Himber C, Ruiz-Ferrer V, Alioua A, Voinnet O (2007) Intra- and intercellular RNA interference in Arabidopsis thaliana requires components of the microRNA and heterochromatic silencing pathways. Nat Genet 39:848–856. doi:10.1038/ng2081
Eamens AL, Smith NA, Curtin SJ, Wang MB, Waterhouse PM (2009) The Arabidopsis thaliana double-stranded RNA binding protein DRB1 directs guide strand selection from microRNA duplexes. RNA 15:2219–2235. doi:10.1261/rna.1646909
Eamens AL, Kim KW, Curtin SJ, Waterhouse PM (2012a) DRB2 is required for microRNA biogenesis in Arabidopsis thaliana. PLoS One 7:e35933. doi:10.1371/journal.pone.0035933
Eamens AL, Wook Kim K, Waterhouse PM (2012b) DRB2, DRB3 and DRB5 function in a non-canonical microRNA pathway in Arabidopsis thaliana. Plant Signal Behav 7:1224–1229. doi:10.4161/psb.21518
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Ferrandon D, Elphick L, Nusslein-Volhard C, St Johnston D (1994) Staufen protein associates with the 3′UTR of bicoid mRNA to form particles that move in a microtubule-dependent manner. Cell 79:1221–1232
Fierro-Monti I, Mathews MB (2000) Proteins binding to duplexed RNA: one motif, multiple functions. Trends Biochem Sci 25:241–246
Fukudome A, Kanaya A, Egami M, Nakazawa Y, Hiraguri A, Moriyama H, Fukuhara T (2011) Specific requirement of DRB4, a dsRNA-binding protein, for the in vitro dsRNA-cleaving activity of Arabidopsis Dicer-like 4. RNA 17:750–760. doi:10.1261/rna.2455411
Gasciolli V, Mallory AC, Bartel DP, Vaucheret H (2005) Partially redundant functions of Arabidopsis DICER-like enzymes and a role for DCL4 in producing trans-acting siRNAs. Curr Biol 15:1494–1500. doi:10.1016/j.cub.2005.07.024
Gleghorn ML, Gong C, Kielkopf CL, Maquat LE (2013) Staufen1 dimerizes through a conserved motif and a degenerate dsRNA-binding domain to promote mRNA decay. Nat Struct Mol Biol 20:515–524. doi:10.1038/nsmb.2528
Gregory RI, Yan KP, Amuthan G, Chendrimada T, Doratotaj B, Cooch N, Shiekhattar R (2004) The Microprocessor complex mediates the genesis of microRNAs. Nature 432:235–240. doi:10.1038/nature03120
Guindon S, Gascuel O (2003) A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52:696–704
Han J, Lee Y, Yeom KH, Kim YK, Jin H, Kim VN (2004) The Drosha–DGCR8 complex in primary microRNA processing. Genes Dev 18:3016–3027. doi:10.1101/gad.1262504
Han J et al (2006) Molecular basis for the recognition of primary microRNAs by the Drosha–DGCR8 complex. Cell 125:887–901. doi:10.1016/j.cell.2006.03.043
Hiraguri A et al (2005) Specific interactions between Dicer-like proteins and HYL1/DRB-family dsRNA-binding proteins in Arabidopsis thaliana. Plant Mol Biol 57:173–188. doi:10.1007/s11103-004-6853-5
Hitti EG, Sallacz NB, Schoft VK, Jantsch MF (2004) Oligomerization activity of a double-stranded RNA-binding domain. FEBS Lett 574:25–30. doi:10.1016/j.febslet.2004.07.080
Hu CD, Chinenov Y, Kerppola TK (2002) Visualization of interactions among bZIP and Rel family proteins in living cells using bimolecular fluorescence complementation. Mol Cell 9:789–798. doi:10.1016/S1097-2765(02)00496-3
Hutvagner G, McLachlan J, Pasquinelli AE, Balint E, Tuschl T, Zamore PD (2001) A cellular function for the RNA-interference enzyme Dicer in the maturation of the let-7 small temporal RNA. Science 293:834–838. doi:10.1126/science.1062961
Jakubiec A, Yang SW, Chua NH (2012) Arabidopsis DRB4 protein in antiviral defense against Turnip yellow mosaic virus infection. Plant J 69:14–25. doi:10.1111/j.1365-313X.2011.04765.x
Jeddeloh JA, Stokes TL, Richards EJ (1999) Maintenance of genomic methylation requires a SWI2/SNF2-like protein. Nat Genet 22:94–97. doi:10.1038/8803
Kim YK, Furic L, Parisien M, Major F, DesGroseillers L, Maquat LE (2007) Staufen1 regulates diverse classes of mammalian transcripts. EMBO J 26:2670–2681. doi:10.1038/sj.emboj.7601712
Kiyota E, Okada R, Kondo N, Hiraguri A, Moriyama H, Fukuhara T (2011) An Arabidopsis RNase III-like protein, AtRTL2, cleaves double-stranded RNA in vitro. J Plant Res 124:405–414. doi:10.1007/s10265-010-0382-x
Kurihara Y, Watanabe Y (2004) Arabidopsis micro-RNA biogenesis through Dicer-like 1 protein functions. Proc Natl Acad Sci USA 101:12753–12758. doi:10.1073/pnas.0403115101
Kurihara Y, Takashi Y, Watanabe Y (2006) The interaction between DCL1 and HYL1 is important for efficient and precise processing of pri-miRNA in plant microRNA biogenesis. Rna 12:206–212
Laraki G, Clerzius G, Daher A, Melendez-Pena C, Daniels S, Gatignol A (2008) Interactions between the double-stranded RNA-binding proteins TRBP and PACT define the Medipal domain that mediates protein-protein interactions. RNA Biol 5:92–103
Le SQ, Gascuel O (2008) An improved general amino acid replacement matrix. Mol Biol Evol 25:1307–1320. doi:10.1093/molbev/msn067
Lee Y et al (2003) The nuclear RNase III Drosha initiates microRNA processing. Nature 425:415–419. doi:10.1038/nature01957
Lee Y, Hur I, Park SY, Kim YK, Suh MR, Kim VN (2006) The role of PACT in the RNA silencing pathway. EMBO J 25:522–532. doi:10.1038/sj.emboj.7600942
Liu F, Bakht S, Dean C (2012) Cotranscriptional role for Arabidopsis DICER-LIKE 4 in transcription termination. Science 335:1621–1623. doi:10.1126/science.1214402
Manavella PA, Hagmann J, Ott F, Laubinger S, Franz M, Macek B, Weigel D (2012) Fast-forward genetics identifies plant CPL phosphatases as regulators of miRNA processing factor HYL1. Cell 151:859–870. doi:10.1016/j.cell.2012.09.039
Masliah G, Barraud P, Allain FH (2013) RNA recognition by double-stranded RNA binding domains: a matter of shape and sequence. Cell Mol Life Sci CMLS 70:1875–1895. doi:10.1007/s00018-012-1119-x
McCue AD, Nuthikattu S, Reeder SH, Slotkin RK (2012) Gene expression and stress response mediated by the epigenetic regulation of a transposable element small RNA. PLoS Genet 8:e1002474. doi:10.1371/journal.pgen.1002474
McCue AD, Nuthikattu S, Slotkin RK (2013) Genome-wide identification of genes regulated in trans by transposable element small interfering RNAs. RNA Biol 10:1379–1395. doi:10.4161/rna.25555
McCue AD, Panda K, Nuthikattu S, Choudury SG, Thomas EN, Slotkin RK (2014) ARGONAUTE 6 bridges transposable element mRNA-derived siRNAs to the establishment of DNA methylation. EMBO J. doi:10.15252/embj.201489499
Micklem DR, Adams J, Grunert S (2000) St Johnston D. Distinct roles of two conserved Staufen domains in oskar mRNA localization and translation EMBO J 19:1366–1377. doi:10.1093/emboj/19.6.1366
Nakazawa Y, Hiraguri A, Moriyama H, Fukuhara T (2007) The dsRNA-binding protein DRB4 interacts with the Dicer-like protein DCL4 in vivo and functions in the trans-acting siRNA pathway. Plant Mol Biol 63:777–785. doi:10.1007/s11103-006-9125-8
Nanduri S, Rahman F, Williams BR, Qin J (2000) A dynamically tuned double-stranded RNA binding mechanism for the activation of antiviral kinase PKR. EMBO J 19:5567–5574. doi:10.1093/emboj/19.20.5567
Nuthikattu S, McCue AD, Panda K, Fultz D, DeFraia C, Thomas EN, Slotkin RK (2013) The initiation of epigenetic silencing of active transposable elements is triggered by RDR6 and 21–22 nucleotide small interfering RNAs. Plant Physiol 162:116–131. doi:10.1104/pp.113.216481
Pall GS, Codony-Servat C, Byrne J, Ritchie L, Hamilton A (2007) Carbodiimide-mediated cross-linking of RNA to nylon membranes improves the detection of siRNA, miRNA and piRNA by northern blot. Nucleic Acids Res 35:e60
Pélissier T, Bousquet-Antonelli C, Lavie L, Deragon JM (2004) Synthesis and processing of tRNA-related SINE transcripts in Arabidopsis thaliana. Nucleic Acids Res 32:3957–3966
Pélissier T, Clavel M, Chaparro C, Pouch-Pélissier MN, Vaucheret H, Deragon JM (2011) Double-stranded RNA binding proteins DRB2 and DRB4 have an antagonistic impact on polymerase IV-dependent siRNA levels in Arabidopsis. RNA 17:1502–1510. doi:10.1261/rna.2680711
Qin H, Chen F, Huan X, Machida S, Song J, Yuan YA (2010) Structure of the Arabidopsis thaliana DCL4 DUF283 domain reveals a noncanonical double-stranded RNA-binding fold for protein-protein interaction. RNA 16:474–481. doi:10.1261/rna.1965310
Raja P, Jackel JN, Li S, Heard IM, Bisaro DM (2014) Arabidopsis double-stranded RNA binding protein DRB3 participates in methylation-mediated defense against geminiviruses. J Virol 88:2611–2622. doi:10.1128/JVI.02305-13
Rajagopalan R, Vaucheret H, Trejo J, Bartel DP (2006) A diverse and evolutionarily fluid set of microRNAs in Arabidopsis thaliana. Genes Dev 20:3407–3425. doi:10.1101/gad.1476406
Saito K, Ishizuka A, Siomi H, Siomi MC (2005) Processing of pre-microRNAs by the Dicer-1–Loquacious complex in Drosophila cells. PLoS Biol 3:e235. doi:10.1371/journal.pbio.0030235
Sarazin A, Voinnet O (2014) Exploring new models of easiRNA biogenesis. Nat Genet 46:530–531. doi:10.1038/ng.2993
Schuldt AJ, Adams JH, Davidson CM, Micklem DR, Haseloff J, Johnston D, Brand AH (1998) Miranda mediates asymmetric protein and RNA localization in the developing nervous system. Genes Dev 12:1847–1857
Slotkin RK (2010) The epigenetic control of the Athila family of retrotransposons in Arabidopsis. Epigenetics 5:483–490
Slotkin RK, Vaughn M, Borges F, Tanurdzic M, Becker JD, Feijo JA, Martienssen RA (2009) Epigenetic reprogramming and small RNA silencing of transposable elements in pollen. Cell 136:461–472. doi:10.1016/j.cell.2008.12.038
Sohn SY, Bae WJ, Kim JJ, Yeom KH, Kim VN, Cho Y (2007) Crystal structure of human DGCR8 core. Nat Struct Mol Biol 14:847–853. doi:10.1038/nsmb1294
Song L, Han MH, Lesicka J, Fedoroff N (2007) Arabidopsis primary microRNA processing proteins HYL1 and DCL1 define a nuclear body distinct from the Cajal body. Proc Natl Acad Sci USA 104:5437–5442. doi:10.1073/pnas.0701061104
St Johnston D, Beuchle D, Nusslein-Volhard C (1991) Staufen, a gene required to localize maternal RNAs in the Drosophila egg. Cell 66:51–63
St Johnston D, Brown NH, Gall JG, Jantsch M (1992) A conserved double-stranded RNA-binding domain. Proc Natl Acad Sci USA 89:10979–10983
Steimer A, Amedeo P, Afsar K, Fransz P, Scheid OM, Paszkowski J (2000) Endogenous targets of transcriptional gene silencing in Arabidopsis. Plant Cell 12:1165–1178
Tabara H, Yigit E, Siomi H, Mello CC (2002) The dsRNA binding protein RDE-4 interacts with RDE-1, DCR-1, and a DExH-box helicase to direct RNAi in C. elegans. Cell 109:861–871
Tanurdzic M et al (2008) Epigenomic consequences of immortalized plant cell suspension culture. PLoS Biol 6:2880–2895. doi:10.1371/journal.pbio.0060302
Tian B, Bevilacqua PC, Diegelman-Parente A, Mathews MB (2004) The double-stranded-RNA-binding motif: interference and much more. Nat Rev Mol Cell Biol 5:1013–1023. doi:10.1038/nrm1528
Tomari Y, Matranga C, Haley B, Martinez N, Zamore PD (2004) A protein sensor for siRNA asymmetry. Science 306:1377–1380. doi:10.1126/science.1102755
Voinnet O, Rivas S, Mestre P, Baulcombe D (2003) An enhanced transient expression system in plants based on suppression of gene silencing by the p19 protein of tomato bushy stunt virus. Plant J 33:949–956. doi:10.1046/j.1365-313X.2003.01676.x
Xie Z et al (2004) Genetic and functional diversification of small RNA pathways in plants. PLoS Biol 2:E104. doi:10.1371/journal.pbio.0020104
Yang SW, Chen HY, Yang J, Machida S, Chua NH, Yuan YA (2010) Structure of Arabidopsis HYPONASTIC LEAVES1 and its molecular implications for miRNA processing. Structure 18:594–605. doi:10.1016/j.str.2010.02.006
Yeom KH, Lee Y, Han J, Suh MR, Kim VN (2006) Characterization of DGCR8/Pasha, the essential cofactor for Drosha in primary miRNA processing. Nucleic Acids Res 34:4622–4629. doi:10.1093/nar/gkl458
Zhao T, Li G, Mi S, Li S, Hannon GJ, Wang XJ, Qi Y (2007) A complex system of small RNAs in the unicellular green alga Chlamydomonas reinhardtii. Genes Dev 21:1190–1203. doi:10.1101/gad.1543507
Zhu S et al (2013) Double-stranded RNA-binding protein 4 is required for resistance signaling against viral and bacterial pathogens. Cell Rep 4:1168–1184. doi:10.1016/j.celrep.2013.08.018
Acknowledgments
This work was supported by l’Agence Nationale de la Recherche (ANR-06-BLAN-0203-02), by the CNRS, by the Université de Perpignan (UPVD). It was also published under the framework of the LABEX: ANR-10-LABX-0036_NETRNA and benefits from a funding from the state managed by the French National Research Agency as part of the Investments for the future program. M.C. and T.M. were supported by a grant from the French Ministry of Research and M-A.T. by a core grant from ETH-Z.
Author contributuion
MC and TP carried out the molecular biology studies and JMD the molecular evolution studies. VJ and JD provided general technical help. TM, PD and MAT made the BIFC work. CBA and MNPP helped with the immunoprecipitation experiments. MC, TP, PD and JMD conceived the study, and participated in its design. JMD coordinated the writing of the manuscript. All authors read and approved the final manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Clavel, M., Pélissier, T., Montavon, T. et al. Evolutionary history of double-stranded RNA binding proteins in plants: identification of new cofactors involved in easiRNA biogenesis. Plant Mol Biol 91, 131–147 (2016). https://doi.org/10.1007/s11103-016-0448-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-016-0448-9