Abstract
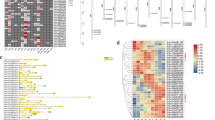
The tomato parthenocarpic fruit (pat) mutation associates a strong competence for parthenocarpy with homeotic transformation of anthers and aberrancy of ovules. To dissect this complex floral phenotype, genes involved in the pollination-independent fruit set of the pat mutant were investigated by microarray analysis using wild-type and mutant ovaries. Normalized expression data were subjected to one-way ANOVA and 2499 differentially expressed genes (DEGs) displaying a >1.5 log-fold change in at least one of the pairwise comparisons analyzed were detected. DEGs were categorized into 20 clusters and clusters classified into five groups representing transcripts with similar expression dynamics. The “regulatory function” group (685 DEGs) contained putative negative or positive fruit set regulators, “pollination-dependent” (411 DEGs) included genes activated by pollination, “fruit growth-related” (815 DEGs) genes activated at early fruit growth. The last groups listed genes with different or similar expression pattern at all stages in the two genotypes. qRT-PCR validation of 20 DEGs plus other four selected genes assessed the high reliability of microarray expression data; the average correlation coefficient for the 20 DEGs was 0.90. In all the groups were evidenced relevant transcription factors encoding proteins regulating meristem differentiation and floral organ development, genes involved in metabolism, transport and response of hormones, genes involved in cell division and in primary and secondary metabolism. Among pathways related to secondary metabolites emerged genes related to the synthesis of flavonoids, supporting the recent evidence that these compounds are important at the fruit set phase. Selected genes showing a de-regulated expression pattern in pat were studied in other four parthenocarpic genotypes either genetically anonymous or carrying lesions in known gene sequences. This comparative approach offered novel insights for improving the present molecular understanding of fruit set and parthenocarpy in tomato.




Similar content being viewed by others
References
Adamski NM, Anastasiou E, Eriksson S, O’Neill CM, Lenhard M (2009) Local maternal control of seed size by KLUH/CYP78A5-dependent growth signaling. Proc Natl Acad Sci USA 106:20115–20120
Ampomah-Dwamena C, Morris BA, Sutherland P, Veit B, Yao JL (2002) Down-regulation of TM29, a tomato SEPALLATA homolog, causes parthenocarpic fruit development and floral reversion. Plant Physiol 130:605–617
Ariizumi T, Shinozaki Y, Ezura H (2013) Genes that influence yield in tomato. Breed Sci 63:3–13
Arnaud N, Pautot V (2014) Ring the BELL and tie the KNOX: roles for TALEs in gynoecium development. Front Plant Sci 5:93
Audran-Delalande C, Bassa C, Mila I, Regad F, Zouine M, Bouzayen M (2012) Genome-wide identification, functional analysis and expression profiling of the Aux/IAA gene family in tomato. Plant Cell Physiol 53:659–672
Avivi Y, Lev-Yadun S, Morozova N, Libs L, Williams L, Zhao J, Varghese G, Grafi G (2000) Clausa, a tomato mutant with a wide range of phenotypic perturbations, displays a cell type-dependent expression of the homeobox gene LeT6/TKn2. Plant Physiol 124:541–552
Bachem CWB, Horvath B, Trindade L, Claassens M, Davelaar E, Jordi W, Visser RGF (2001) A potato tuber-expressed mRNA with homology to steroid dehydrogenases affects gibberellin levels and plant development. Plant J 25:595–604
Bowman JL, Smyth DR (1999) CRABS CLAW, a gene that regulates carpel and nectary development in Arabidopsis, encodes a novel protein with zinc finger and helix-loop-helix domains. Development 126:2387–2396
Carbonell-Bejerano P, Urbez C, Carbonell J, Granell A, Perez-Amador MA (2010) A fertilization-independent developmental program triggers partial fruit development and senescence processes in pistils of Arabidopsis. Plant Physiol 154:163–172
Chen QY, Atkinson A, Otsuga D, Christensen T, Reynolds L, Drews GN (1999) The Arabidopsis FILAMENTOUS FLOWER gene is required for flower formation. Development 126:2715–2726
de Jong M, Wolters-Arts M, Feron R, Mariani C, Vriezen WH (2009) The Solanum lycopersicum auxin response factor 7 (SlARF7) regulates auxin signaling during tomato fruit set and development. Plant J 57:160–170
de Jong M, Wolters-Arts M, García-Martínez JL, Mariani C, Vriezen WH (2011) The Solanum lycopersicum AUXIN RESPONSE FACTOR 7 (SlARF7) mediates cross-talk between auxin and gibberellin signalling during tomato fruit set and development. J Exp Bot 62:617–626
Ding J, Chen B, Xia X, Mao W, Shi K, Zhou Y, Yu J (2013) Cytokinin-induced parthenocarpic fruit development in tomato is partly dependent on enhanced gibberellin and auxin biosynthesis. PLoS ONE 8(7):e70080
Dorcey E, Urbez C, Blázquez M (2009) Fertilization-dependent auxin response in ovules triggers fruit development through the modulation of gibberellin metabolism in Arabidopsis. Plant J 58:318–332
Elliott RC, Betzner AS, Huttner E, Oakes MP, Tucker WQ, Gerentes D, Perez P, Smyth DR (1996) AINTEGUMENTA, an APETALA2-like gene of Arabidopsis with pleiotropic roles in ovule development and floral organ growth. Plant Cell 8:155–168
Expósito-Rodríguez M, Borges AA, Borges-Pérez A, Pérez JA (2008) Selection of internal control genes for quantitative real-time RT-PCR studies during tomato development process. BMC Plant Biol 8:131
Galon Y, Finkler A, Fromm H (2010) Calcium-regulated transcription in plants. Mol Plant 3:653–669
Geuten K, Irish V (2010) Hidden variability of floral homeotic B genes in Solanaceae provides a molecular basis for the evolution of novel functions. Plant Cell 22:2562–2578
Gillaspy G, Ben-David H, Gruissem W (1993) Fruits: a developmental perspective. Plant Cell 5:1439–1451
Goetz M, Hooper LC, Johnson SD, Rodrigues JCM, Vivian-Smith A, Koltunow AM (2007) Expression of aberrant forms of AUXIN RESPONSE FACTOR8 stimulates parthenocarpy in Arabidopsis and tomato. Plant Physiol 145:351–366
Gómez P, Jamilena M, Capel J, Zurita S (1999) Stamenless, a tomato mutant with homeotic conversions in petals and stamens. Planta 209:172–179
Gómez MD, Vera-Sirera F, Pérez-Amador MA (2014) Molecular programme of senescence in dry and fleshy fruits. J Exp Bot 65:4515–4526
Henriksson E, Olsson ASB, Johannesson H, Johansson H, Hanson J, Engström P, Söderman E (2005) Homeodomain leucine zipper class I genes in Arabidopsis. Expression patterns and phylogenetic relationships. Plant Physiol 139:509–518
Huang Z, Van Houten J, Gonzalez G, Xiao H, Esther van der Knaap E (2013) Genome-wide identification, phylogeny and expression analysis of SUN, OFP and YABBY gene family in tomato. Mol Genet Genomics 288:111–129
Ingrosso I, Bonsegna S, De Domenico S, Laddomada B, Blando F, Santino A, Giovinazzo G (2011) Over-expression of a grape stilbene synthase gene in tomato induces parthenocarpy and causes abnormal pollen development. Plant Physiol Biochem 49:1092–1099
Ito T, Meyerowitz ME (2000) Overexpression of a gene encoding a cytocrome p540, cyp78a9, induces large seedless fruit in Arabidopsis. Plant Cell 12:1541–1550
Joung J-G, Corbett AM, Fellman SM, Tieman DM, Klee HJ, Giovannoni JJ, Fei Z (2009) Plant MetGenMAP: an integrative analysis system for plant systems biology. Plant Physiol 151:1758–1768
Kiyohara S, Honda H, Shimizu N, Ejima C, Hamasaki R, Sawa S (2011) Tryptophan auxotroph mutants suppress the superroot2 phenotypes, modulating IAA biosynthesis in arabidopsis. Plant Signal Behav 6:1351–1355
Koltunow AM, Vivian-Smith A, Tucker MR, Peach N (2002) The central role of the ovule in apomixis and parthenocarpy. In: O’Neill SD, Roberts JA (eds) Plant reproduction. Academic Press, Sheffield, pp 221–256
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCt method. Methods 25:402–408
Lora J, Hormaza JI, Herrero M, Gasser CS (2011) Seedless fruits and the disruption of a conserved genetic pathway in angiosperm ovule development. Proc Natl Acad Sci USA 108:5461–5465
Maharjan PM, Dilkes BP, Fujioka S, Ales Pěnčík A, Ljung K, Burow M, Halkier BA, Choe S (2014) Arabidopsis gulliver1/superroot2-7 identifies a metabolic basis for auxin and brassinosteroid synergy. Plant J 80:797–808
Mapelli S, Frova C, Torti G, Soressi GP (1978) Relationship between set, development and activities of growth regulators in tomato fruits. Plant Cell Physiol 19:1281–1288
Martí C, Orzáez D, Ellul P, Moreno V, Carbonell J, Granell A (2007) Silencing of DELLA induces facultative parthenocarpy in tomato fruits. Plant J 52:865–876
Martinelli F, Uratsu SL, Reagan RL, Chen Y, Tricoli D, Fiehn O, Rocke DM, Gasser CS, Dandekar AM (2009) Gene regulation in parthenocarpic tomato fruit. J Exp Bot 60:3873–3890
Matsuo S, Kikuchi K, Fukuda M, Honda I, Imanishi S (2012) Roles and regulation of cytokinins in tomato fruit development. J Exp Bot 63:5569–5579
Mazzucato A, Taddei AR, Soressi GP (1998) The parthenocarpic fruit (pat) mutant of tomato (Lycopersicon esculentum Mill.) sets seedless fruits and has aberrant anther and ovule development. Development 125:107–114
Mazzucato A, Olimpieri I, Siligato F, Picarella ME, Soressi GP (2008) Characterization of genes controlling stamen identity and development in a parthenocarpic tomato mutant indicates a role for the DEFICIENS ortholog in the control of fruit set. Physiol Plant 132:526–537
Mazzucato A, Willems D, Bernini R, Picarella ME, Santangelo E, Ruiu F, Tilesi F, Soressi GP (2013) Novel phenotypes related to the breeding of purple-fruited tomatoes and effect of peel extracts on human cancer cell proliferation. Plant Physiol Biochem 72:125–133
Mazzucato A, Cellini F, Bouzayen M, Zouine M, Mila I, Minoia S, Petrozza A, Picarella ME, Ruiu F, Carriero F (2015) A TILLING allele of the tomato Aux/IAA9 gene offers new insights into fruit set mechanisms and perspectives for breeding seedless tomatoes. Mol Breed 35:22
Medina M, Roque E, Pineda B, Cañas L, Rodriguez-Concepción M, Beltrán JP, Gómez-Mena C (2013) Early anther ablation triggers parthenocarpic fruit development in tomato. Plant Biotechnol J 11:770–779
Mejía N, Soto B, Guerrero M et al (2011) Molecular, genetic and transcriptional evidence for a role of VvAGL11 in stenospermocarpic seedlessness in grapevine. BMC Plant Biol 11:57
Molesini B, Pandolfini T, Rotino GL, Dani V, Spena A (2009) Aucsia gene silencing causes parthenocarpic fruit development in tomato. Plant Physiol 149:534–548
Mounet F, Moing A, Kowalczyk M et al (2012) Down-regulation of a single auxin efflux transport protein in tomato induces precocious fruit development. J Exp Bot 63:4901–4917
Muños S, Ranc N, Botton E, Bérard AL, Rolland S, Duffé P, Carretero Y, Le Paslier M-C, Delalande C, Bouzayen M, Brunel D, Causse M (2011) Increase in tomato locule number is controlled by two single-nucleotide polymorphisms located near WUSCHEL. Plant Physiol 156:2244–2254
Nesi N, Debeaujon I, Jond C, Stewart AJ, Jenkins GI, Caboche M, Lepiniec LC (2002) The TRANSPARENT TESTA16 locus encodes the ARABIDOPSIS BSISTER MADS domain protein and is required for proper development and pigmentation of the seed coat. Plant Cell 14:2463–2479
Nwafor C, Gribaudo I, Schneider A, Wehrens R, Grando M, Costantini L (2014) Transcriptome analysis during berry development provides insights into co-regulated and altered gene expression between a seeded wine grape variety and its seedless somatic variant. BMC Genomics 15:1030
Olimpieri I, Mazzucato A (2008) Phenotypic and genetic characterization of the pistillate mutation in tomato. Theor Appl Genet 118:151–163
Olimpieri I, Siligato F, Caccia R, Mariotti L, Ceccarelli N, Soressi GP, Mazzucato A (2007) Tomato fruit set driven by pollination or by the parthenocarpic fruit allele are mediated by transcriptionally regulated gibberellin biosynthesis. Planta 226:877–888
Otsuga D, DeGuzman B, Prigge MJ, Drews GN, Clark SE (2001) REVOLUTA regulates meristem initiation at lateral positions. Plant J 25:223–236
Park SO, Zheng Z, Oppenheimer DG, Hauser BA (2005) The PRETTY FEW SEEDS2 gene encodes an Arabidopsis homeodomain protein that regulates ovule development. Development 132:841–849
Pascual L, Blanca JM, Cañizares J, Nuez F (2009) Transcriptomic analysis of tomato carpel development reveals alterations in ethylene and gibberellin synthesis during pat3/pat4 parthenocarpic fruit set. BMC Plant Biol 9:67
Pattison RJ, Catalá C (2012) Evaluating auxin distribution in tomato (Solanum lycopersicum) through an analysis of the PIN and AUX/LAX gene families. Plant J 70:585–598
Philouze J (1991) Description of isogenic lines, except for one, or two, monogenically controlled morphological traits in tomato, Lycopersicon esculentum Mill. Euphytica 56:121–131
Philouze J, Maisonneuve B (1978) Heredity of the natural ability to set parthenocarpic fruits in the soviet variety Severianin. Tomato Genet Coop 28:12–13
Pnueli L, Hareven D, Broday L, Hurwitz C, Lifschitz E (1994) The TM5 MADS box gene mediates organ differentiation in the three inner whorls of tomato flowers. Plant Cell 6:175–186
Popescu SC, Popescu GV, Bachan S, Zhang Z, Seay M, Gerstein M, Snyder M, Dinesh-Kumar SP (2007) Differential binding of calmodulin related proteins to their targets revealed through high-density Arabidopsis protein microarrays. Proc Natl Acad Sci USA 104:4730–4735
Preston JC, Hileman LC (2013) Functional evolution in the plant SQUAMOSA-PROMOTER BINDING PROTEIN-LIKE (SPL) gene family. Front Plant Sci. doi:10.3389/fpls.2013.00080
Rieu I, Ruiz-Rivero O, Fernandez-Garcia N et al (2008) The gibberellin biosynthetic genes AtGA20ox1 and AtGA20ox2 act, partially redundantly, to promote growth and development throughout the Arabidopsis life cycle. Plant J 53:488–504
Ruan Y-L, Patrick JW, Bouzayen M, Osorio S, Fernie AR (2012) Molecular regulation of seed and fruit set. Trends Plant Sci 17:656–665
Rubin G, Tohge T, Matsuda F, Saito K, Scheible W-R (2009) Members of the LBD family of transcription factors repress anthocyanin synthesis and affect additional nitrogen responses in Arabidopsis. Plant Cell 21:3567–3584
Sanders PM, Bui AQ, Weterings K, McIntire KN, Hsu Y-C, Lee PY, Truong MT, Beals TP, Goldberg RB (1999) Anther developmental defects in Arabidopsis thaliana male-sterile mutants. Sex Plant Reprod 11:297–322
Schijlen EGWM, De Vos CHR, Martens S, Jonker HH, Rosin FM, Molthoff JW, Tikunov YM, Angenent GC, Van Tunen AJ, Bovy AG (2007) RNA interference silencing of chalcone synthase, the first step in the flavonoid biosynthesis pathway, leads to parthenocarpic tomato fruits. Plant Physiol 144:1520–1530
Schoenbohm C, Martens S, Eder C, Forkmann G, Weisshaar B (2000) Identification of the Arabidopsis thaliana flavonoid 3′-hydroxylase gene and functional expression of the encoded P450 enzyme. Biol Chem 381:749–753
Schwabe WW, Mills JJ (1981) Hormones and parthenocarpic fruit set: a literature survey. Hort Abstr 51:661–698
Serrani JC, Ruiz-Rivero O, Fos M, García-Martínez JL (2008) Auxin-induced fruit-set in tomato is mediated in part by gibberellins. Plant J 56:922–934
Shinozaki Y, Hao S, Kojima M, Sakakibara H, Ozeki-Iida Y, Zheng Y, Fei Z, Zhong S, Giovannoni JJ, Rose JKC, Okabe Y, Heta Y, Ezura H, Ariizumi T (2015) Ethylene suppresses tomato (Solanum lycopersicum) fruit set through modification of gibberellin metabolism. Plant J 83:237–251
Shuai B, Reynaga-Peña C, Springer P (2002) The lateral organ boundaries gene defines a novel, plant specific gene family. Plant Physiol 129:747–761
Sotelo-Silveira M, Cucinotta M, Chauvin A-L, Chávez Montes RA, Colombo L, Marsch-Martínez N, de Folter S (2013) Cytochrome P450 CYP78A9 is involved in Arabidopsis reproductive development. Plant Physiol 162:779–799
Tabata R, Ikezaki M, Fujibe T, Aida M, Tian C, Ueno Y, Yamamoto KT, Machida Y, Nakamura K, Ishiguro S (2010) Arabidopsis AUXIN RESPONSE FACTOR6 and 8 regulate jasmonic acid biosynthesis and floral organ development via repression of class 1 KNOX genes. Plant Cell Physiol 51:164–175
Tadege M, Lin H, Bedair M et al (2011) STENOFOLIA regulates blade outgrowth and leaf vascular patterning in Medicago truncatula and Nicotiana sylvestris. Plant Cell 23:2125–2142
Tan X, Calderon-Villalobos LIA, Sharon M, Zheng C, Robinson CV, Estelle M, Zheng N (2007) Mechanism of auxin perception by the TIR1 ubiquitin ligase. Nature 446:640–645
Tang N, Deng W, Hu G, Hu N, Li Z (2015) Transcriptome profiling reveals the regulatory mechanism underlying pollination dependent and parthenocarpic fruit set mainly mediated by auxin and gibberellin. PLoS One 10:e0125355
Testa G, Caccia R, Tilesi F, Soressi G, Mazzucato A (2002) Sequencing and characterization of tomato genes putatively involved in fruit set and early development. Sex Plant Reprod 14:269–277
Tiwari A, Vivian-Smith A, Voorrips RE, Habets MEJ, Xue LB, Offringa R, Heuvelink EP (2011) Parthenocarpic potential in Capsicum annuum L. is enhanced by carpelloid structures and controlled by a single recessive gene. BMC Plant Biol 11:143
Tsai W-C, Lee P-F, Chen H-I, Hsiao Y-Y, Wei W-J, Pan Z-J, Chuang M-H, Kuoh C-S, Chen W-H, Chen H-H (2005) PeMADS6, a GLOBOSA/PISTILLATA-like gene in Phalaenopsis equestris involved in petaloid formation, and correlated with flower longevity and ovary development. Plant Cell Physiol 46:1125–1139
Varaud E, Brioudes F, Szécsi J, Leroux J, Brown S, Perrot-Rechenmann C, Bendahmane M (2011) AUXIN RESPONSE FACTOR8 regulates Arabidopsis petal growth by interacting with the bHLH transcription factor BIGPETALp. Plant Cell 23:973–983
Vivian-Smith A, Luo M, Chaudhury A, Koltunow A (2001) Fruit development is actively restricted in the absence of fertilization in Arabidopsis. Development 128:2321–2331
Vriezen WH, Feron R, Maretto F, Keijman J, Mariani C (2008) Changes in tomato ovary transcriptome demonstrate complex hormonal regulation of fruit set. New Phytol 177:60–76
Wang H, Jones B, Li Z, Frasse P, Delalande C, Regad F, Chaabouni S, Latché A, Pech J-C, Bouzayen M (2005) The tomato Aux/IAA transcription factor IAA9 is involved in fruit development and leaf morphogenesis. Plant Cell 17:2676–2692
Wang Y, Zhang W-Z, Song L-F, Zou J-J, Su Z, Wu W-H (2008) Transcriptome analyses show changes in gene expression to accompany pollen germination and tube growth in Arabidopsis. Plant Physiol 148:1201–1211
Wang H, Schauer N, Usadel B, Frasse P, Zouine M, Hernould M, Latché A, Pech J-C, Fernie AR, Bouzayen M (2009) Regulatory features underlying pollination-dependent and -independent tomato fruit set revealed by transcript and primary metabolite profiling. Plant Cell 21:1428–1452
Weigel D, Meyerowitz E (1994) The ABCs of floral homeotic genes. Cell 78:203–209
Wu J, Wang F, Cheng L, Kong F, Peng Z, Liu S, Yu X, Lu G (2011) Identification, isolation and expression analysis of auxin response factor (ARF) genes in Solanum lycopersicum. Plant Cell Rep 30:2059–2073
Yao J, Dong Y, Morris BA (2001) Parthenocarpic apple fruit production conferred by transposon insertion mutations in a MADS-box transcription factor. Proc Natl Acad Sci USA 98:1306–1311
Acknowledgments
The authors thank Joaquín Cañizares, Filomena Carriero, Celestina Mariani and Ivo Rieu for sharing material of the parthenocarpic systems studied. J. Cañizares is also acknowledged for critical reading of the manuscript; Luigi Selleri, Marco Cirilli and Pietro Mosconi for expert technical assistance and two anonymous reviewers for substantial help in improving the paper.
Author contributions statement
F.R. and A.M. conceived and designed the research. F.R. and M.E.P. performed phenotypic and gene validation analysis, S.I. performed the microarray hybridization, F.R., M.E.P. and S.I. analysed the data, F.R. and A.M. wrote the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ruiu, F., Picarella, M.E., Imanishi, S. et al. A transcriptomic approach to identify regulatory genes involved in fruit set of wild-type and parthenocarpic tomato genotypes. Plant Mol Biol 89, 263–278 (2015). https://doi.org/10.1007/s11103-015-0367-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-015-0367-1