Abstract
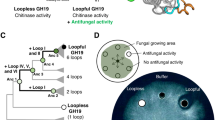
Chitinases help plants defend themselves against fungal attack, and play roles in other processes, including development. The catalytic modules of most plant chitinases belong to glycoside hydrolase family 19. We report here x-ray structures of such a module from a Norway spruce enzyme, the first for any family 19 class IV chitinase. The bi-lobed structure has a wide cleft lined by conserved residues; the most interesting for catalysis are Glu113, the proton donor, and Glu122, believed to be a general base that activate a catalytic water molecule. Comparisons to class I and II enzymes show that loop deletions in the class IV proteins make the catalytic cleft shorter and wider; from modeling studies, it is predicted that only three N-acetylglucosamine-binding subsites exist in class IV. Further, the structural comparisons suggest that the family 19 enzymes become more closed on substrate binding. Attempts to solve the structure of the complete protein including the associated chitin-binding module failed, however, modeling studies based on close relatives indicate that the binding module recognizes at most three N-acetylglucosamine units. The combined results suggest that the class IV enzymes are optimized for shorter substrates than the class I and II enzymes, or alternatively, that they are better suited for action on substrates where only small regions of chitin chain are accessible. Intact spruce chitinase is shown to possess antifungal activity, which requires the binding module; removing this module had no effect on measured chitinase activity.







Similar content being viewed by others
References
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Andersen OA, Dixon MJ, Eggleston IM, van Aalten DM (2005) Natural product family 18 chitinase inhibitors. Nat Prod Rep 22:563–579
Arlorio M, Ludwig A, Boller T, Bonfante P (1992) Inhibition of fungal growth by plant chitinases and beta-1, 3-glucanases—a morphological-study. Protoplasma 171:34–43
Berger S, Menudier A, Julien R, Karamanos Y (1995) Do de-N-glycosylation enzymes have an important role in plant cells? Biochimie 77:751–760
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The protein data bank. Nucleic Acids Res 28:235–242
Boraston AB, Bolam DN, Gilbert HJ, Davies GJ (2004) Carbohydrate-binding modules: fine-tuning polysaccharide recognition. Biochem J 382:769–781
Brameld KA, Goddard WA 3rd (1998) The role of enzyme distortion in the single displacement mechanism of family 19 chitinases. Proc Natl Acad Sci USA 95:4276–4281
Broglie K, Chet I, Holliday M, Cressman R, Biddle P, Knowlton S, Mauvais CJ, Broglie R (1991) Transgenic plants with enhanced resistance to the fungal pathogen rhizoctonia solani. Science 254:1194–1197
Brunner F, Stintzi A, Fritig B, Legrand M (1998) Substrate specificities of tobacco chitinases. Plant J 14:225–234
Collaborative Computational Project, Number 4 (1994) The CCP4 Suite: programs for protein crystallography. Acta Cryst D50:760–763
Engh RA, Huber R (1991) Accurate bond and angle parameters for x-ray protein-structure refinement. Acta Crystallogr A 47:392–400
Evans PR (1997) Scaling of MAD data. In: Wilson KS, Davies G, Ashton AW, Bailey S (eds) Proceedings of the CCP4 study weekend. CCLRC Daresbury Laboratory, Warrington, pp 97–102
Frettinger P, Herrmann S, Lapeyrie F, Oelmuller R, Buscot F (2006) Differential expression of two class III chitinases in two types of roots of Quercus robur during pre-mycorrhizal interactions with Piloderma croceum. Mycorrhiza 16:219–223
Fukamizo T (2000) Chitinolytic enzymes: catalysis, substrate binding, and their application. Curr Protein Pept Sci 1:105–124
Hahn M, Hennig M, Schlesier B, Hohne W (2000) Structure of jack bean chitinase. Acta Crystallogr D Biol Crystallogr 56(Pt 9):1096–1099
Harris M, Jones TA (2001) Molray-a web interface between O and the POV-ray ray tracer. Acta Crystallogr D Biol Crystallogr 57:1201–1203
Hart PJ, Pfluger HD, Monzingo AF, Hollis T, Robertus JD (1995) The refined crystal structure of an endochitinase from Hordeum vulgare L. seeds at 1.8 A resolution. J Mol Biol 248:402–413
Henrissat B, Bairoch A (1993) New families in the classification of glycosyl hydrolases based on amino-acid-sequence similarities. Biochem J 293:781–788
Hietala AM, Kvaalen H, Schmidt A, Johnk N, Solheim H, Fossdal CG (2004) Temporal and spatial profiles of chitinase expression by norway spruce in response to bark colonization by Heterobasidion annosum. Appl Environ Microbiol 70:3948–3953
Hoell IA, Dalhus B, Heggset EB, Aspmo SI, Eijsink VG (2006) Crystal structure and enzymatic properties of a bacterial family 19 chitinase reveal differences from plant enzymes. FEBS J 273:4889–4900
Huet J, Rucktooa P, Clantin B, Azarkan M, Looze Y, Villeret V, Wintjens R (2008) X-ray structure of papaya chitinase reveals the substrate binding mode of glycosyl hydrolase family 19 chitinases. Biochemistry 47:8283–8291
Jones TA, Zou JY, Cowan SW, Kjeldgaard M (1991) Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr A 47(Pt 2):110–119
Karasuda S, Tanaka S, Kajihara H, Yamamoto Y, Koga D (2003) Plant chitinase as a possible biocontrol agent for use instead of chemical fungicides. Biosci Biotechnol Biochem 67:221–224
Kasprzewska A (2003) Plant chitinases–regulation and function. Cell Mol Biol Lett 8:809–824
Kawase T, Yokokawa S, Saito A, Fujii T, Nikaidou N, Miyashita K, Watanabe T (2006) Comparison of enzymatic and antifungal properties between family 18 and 19 chitinases from S. coelicolor A3(2). Biosci Biotechnol Biochem 70:988–998
Kezuka Y, Ohishi M, Itoh Y, Watanabe J, Mitsutomi M, Watanabe T, Nonaka T (2006) Structural studies of a two-domain chitinase from Streptomyces griseus HUT6037. J Mol Biol 358:472–484
Kim JS, Kim YO, Ryu HJ, Kwak YS, Lee JY, Kang H (2003) Isolation of stress-related genes of rubber particles and latex in fig tree (Ficus carica) and their expressions by abiotic stress or plant hormone treatments. Plant Cell Physiol 44:412–414
Kleywegt GJ, Jones TA (1996) Phi/psi-chology: Ramachandran revisited. Structure 4:1395–1400
Kleywegt GJ, Jones TA (1997) Detecting folding motifs and similarities in protein structures. In: Carter JCW, Sweet RM (eds) Methods in enzymology, macromolecular crystallography, Pt B. Academic Press, New York, pp 525–545
Kleywegt GJ, Zou JY, Kjeldgaard M, Jones TA (2001) Around O. In: Rossmann MG, Arnold E (eds) International tables for crystallography. Vol. F crystallography of biological macromolecules. Kluwer, Dordrecht, pp 353–356, 366–367
Kraulis PJ (1991) Molscript—a program to produce both detailed and schematic plots of protein structures. J Appl Crystallogr 24:946–950
Leslie AGW (1992) Recent changes to the MOSFLM package for processing film and image plate data. Joint CCP4 + ESF-EAMCB newsletter on protein crystallography
Matthews BW (1968) Solvent content of protein crystals. J Mol Biol 33:491–497
McCarter JD, Withers SG (1994) Mechanisms of enzymatic glycoside hydrolysis. Curr Opin Struct Biol 4:885–892
Meins F, Fritig B, Linthorst HJM, Mikkelsen JD, Neuhaus JM, Ryals J (1994) Plant chitinase genes. Plant Mol Biol Rep 12:S22–S28
Melchers LS, Apotheker-de Groot M, van der Knaap JA, Ponstein AS, Sela-Buurlage MB, Bol JF, Cornelissen BJ, van den Elzen PJ, Linthorst HJ (1994) A new class of tobacco chitinases homologous to bacterial exo-chitinases displays antifungal activity. Plant J 5:469–480
Mizuno R, Fukamizo T, Sugiyama S, Nishizawa Y, Kezuka Y, Nonaka T, Suzuki K, Watanabe T (2008) Role of the loop structure of the catalytic domain in rice class I chitinase. J Biochem 143:487–495
Murshudov GN, Vagin AA, Dodson EJ (1997) Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr D Biol Crystallogr 53:240–255
Neuhaus JM, Fritig B, Linthorst HJM, Meins F, Mikkelsen JD, Ryals J (1996) A revised nomenclature for chitinase genes. Plant Mol Biol Rep 14:102–104
Otwinowski Z, Minor W (1997) Processing of X-ray diffraction data collected in oscillation mode. In: Carter JCW, Sweet RM (eds) Methods in enzymology, macromolecular crystallography, Pt A. Academic Press, New York, pp 307–326
Patil RS, Ghormade V, Deshpande MV (2000) Chitinolytic enzymes: an exploration. Enzyme Microb Technol 26:473–483
Potterton E, Briggs P, Turkenburg M, Dodson E (2003) A graphical user interface to the CCP4 program suite. Acta Crystallogr D Biol Crystallogr 59:1131–1137
Robertus JD, Monzingo AF (1999) The structure and action of chitinases. EXS 87:125–135
Schultze M, Staehelin C, Brunner F, Genetet I, Legrand M, Fritig B, Kondorosi E, Kondorosi A (1998) Plant chitinase/lysozyme isoforms show distinct substrate specificity and cleavage site preference towards lipochitooligosaccharide nod signals. Plant J 16:571–580
Scorer CA, Clare JJ, McCombie WR, Romanos MA, Sreekrishna K (1994) Rapid selection using G418 of high copy number transformants of Pichia pastoris for high-level foreign gene expression. Bio-Technology 12:181–184
Song HK (1996) Refined structure of the chitinase from barley seeds at 2.0 A resolution. Acta Crystallogr D Biol Crystallogr 52:289–298
Tang CM, Chye ML, Ramalingam S, Ouyang SW, Zhao KJ, Ubhayasekera W, Mowbray SL (2004) Functional analyses of the chitin-binding domains and the catalytic domain of Brassica juncea chitinase BjCHI1. Plant Mol Biol 56:285–298
Ubhayasekera W, Tang CM, Ho SW, Berglund G, Bergfors T, Chye ML, Mowbray SL (2007) Crystal structures of a family 19 chitinase from Brassica juncea show flexibility of binding cleft loops. FEBS J 274:3695–3703
Vagin A, Teplyakov A (1997) MOLREP: an automated program for molecular replacement. J Appl Crystallogr 30:1022–1025
van Aalten DM, Komander D, Synstad B, Gaseidnes S, Peter MG, Eijsink VG (2001) Structural insights into the catalytic mechanism of a family 18 exo-chitinase. Proc Natl Acad Sci USA 98:8979–8984
van Hengel AJ, Tadesse Z, Immerzeel P, Schols H, van Kammen A, de Vries SC (2001) N-acetylglucosamine and glucosamine-containing arabinogalactan proteins control somatic embryogenesis. Plant Physiol 125:1880–1890
Vierheilig H, Alt-Hug M, Wiemken A, Boller T (2001) Hyphal in vitro growth of the arbuscular mycorrhizal fungus Glomus mosseae is affected by chitinase but not by beta-1,3-glucanase. Mycorrhiza 11:279–282
Wirth SJ, Wolf GA (1990) Dye-labelled substrates for the assay and detection of chitinase and lysozyme activity. J Microbiol Methods 12:197–205
Wiweger M, Farbos I, Ingouff M, Lagercrantz U, Von Arnold S (2003) Expression of Chia4-Pa chitinase genes during somatic and zygotic embryo development in Norway spruce (Picea abies): similarities and differences between gymnosperm and angiosperm class IV chitinases. J Exp Bot 54:2691–2699
Wright CS (1990) 2.2 A resolution structure analysis of two refined N-acetylneuraminyl-lactose–wheat germ agglutinin isolectin complexes. J Mol Biol 215:635–651
Yan R, Hou J, Ding D, Guan W, Wang C, Wu Z, Li M (2008) In vitro antifungal activity and mechanism of action of chitinase against four plant pathogenic fungi. J Basic Microbiol 48:293–301
Acknowledgments
The authors would like to thank Dr. Fred Asiegbu (Swedish University of Agricultural Sciences) for providing Heterobasidion annosum (strain FP5), and Dr. Mark Harris (Uppsala University) for photographing the chitinase-inhibited fungal plate. The work was supported by the Swedish Research Council (VR) and the Swedish Foundation for Strategic Research via the Glycoconjugates in Biological Systems network, GLIBS (SLM), as well as by the University of Hong Kong (ORA10208034) (MLC).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ubhayasekera, W., Rawat, R., Ho, S.W.T. et al. The first crystal structures of a family 19 class IV chitinase: the enzyme from Norway spruce. Plant Mol Biol 71, 277–289 (2009). https://doi.org/10.1007/s11103-009-9523-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-009-9523-9