Abstract
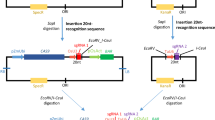
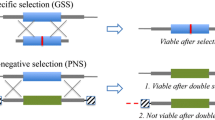
We have demonstrated that targeted mutagenesis can be accomplished in maize plants by excision, activation, and subsequent elimination of an endonuclease in the progeny of genetic crosses. The yeast FLP/FRT site-specific recombination system was used to excise and transiently activate the previously integrated yeast I-SceI homing endonuclease in maize zygotes and/or developing embryos. An artificial I-SceI recognition sequence integrated into genomic DNA was analyzed for mutations to indicate the I-SceI endonuclease activity. Targeted mutagenesis of the I-SceI site occurred in about 1% of analyzed F1 plants. Short deletions centered on the I-SceI-produced double-strand break were the predominant genetic lesions observed in the F1 plants. The I-SceI expression cassette was not detected in the mutant F1 plants and their progeny. However, the original mutations were faithfully transmitted to the next generation indicating that the mutations occurred early during the F1 plant development. The procedure offers simultaneous production of double-strand breaks and delivery of DNA template combined with a large number of progeny plants for future gene targeting experiments.






Similar content being viewed by others
References
An G, Mitra A, Choi HK, Costa MA, An K, Thornburg RW, Ryan CA (1989) Functional analysis of the 3′ control region of the potato wound-inducible proteinase inhibitor II gene. Plant Cell 1:115–122
Arnould S, Perez C, Cabaniols JP, Smith J, Gouble A, Grizot S, Epinat JC, Duclert A, Duchateau P, Paques F (2007) Engineered I-CreI derivatives cleaving sequences from the human XPC gene can induce highly efficient gene correction in mammalian cells. J Mol Biol 371:49–65. doi:10.1016/j.jmb.2007.04.079
Beumer K, Bhattacharyya G, Bibikova M, Trautman JK, Carroll D (2006) Efficient gene targeting in Drosophila with zinc-finger nucleases. Genetics 172:2391–2403. doi:10.1534/genetics.105.052829
Bibikova M, Golic M, Golic KG, Carroll D (2002) Targeted chromosomal cleavage and mutagenesis in Drosophila using zinc-finger nucleases. Genetics 161:1169–1175
Bibikova M, Beumer K, Trautman JK, Carroll D (2003) Enhancing gene targeting with designed zinc finger nucleases. Science 300:764. doi:10.1126/science.1079512
Carroll D, Beumer KJ, Morton JJ, Bozas A, Trautman JK (2008) Gene targeting in Drosophila and Caenorhabditis elegans with zinc-finger nucleases. Methods Mol Biol 435:63–77. doi:10.1007/978-1-59745-232-8_5
Cheng P-C, Pareddy DR (1994) Morphology and development of the tassel and ear. In: Freeling M, Walbot V (eds) The maize handbook. Springer-Verlag, New York, Berlin, Heidelberg, London, Paris, pp 37–47
Chiurazzi M, Ray A, Viret JF, Perera R, Wang XH, Lloyd AM, Signer ER (1996) Enhancement of somatic intrachromosomal homologous recombination in Arabidopsis by the HO endonuclease. Plant Cell 8:2057–2066
Choulika A, Perrin A, Dujon B, Nicolas JF (1995) Induction of homologous recombination in mammalian chromosomes by using the I-SceI system of Saccharomyces cerevisiae. Mol Cell Biol 15:1968–1973
Cornu TI, Thibodeau-Beganny S, Guhl E, Alwin S, Eichtinger M, Joung JK, Cathomen T (2008) DNA-binding specificity is a major determinant of the activity and toxicity of zinc-finger nucleases. Mol Ther 16:352–358. doi:10.1038/sj.mt.6300357
D’Halluin K, Vanderstraeten C, Stals E, Cornelissen M, Ruiter R (2008) Homologous recombination: a basis for targeted genome optimization in crop species such as maize. Plant Biotechnol J 6:93–102
Djukanovic V, Orczyk W, Gao H, Sun X, Garrett N, Zhen S, Gordon-Kamm W, Barton J, Lyznik LA (2006) Gene conversion in transgenic maize plants expressing FLP/FRT and Cre/loxP site-specific recombination systems. Plant Biotechnol J 4:345–357. doi:10.1111/j.1467-7652.2006.00186.x
Djukanovic V, Lenderts B, Bidney D, Lyznik LA (2008) A Cre:FLP fusion protein recombines FRT or loxP sites in transgenic maize plants. Plant Biotechnol J 6:770–781. doi:10.1111/j.1467-7652.2008.00357.x
Doyon JB, Pattanayak V, Meyer CB, Liu DR (2006) Directed evolution and substrate specificity profile of homing endonuclease I-SceI. J Am Chem Soc 128:2477–2484. doi:10.1021/ja057519l
Doyon Y, McCammon JM, Miller JC, Faraji F, Ngo C, Katibah GE, Amora R, Hocking TD, Zhang L, Rebar EJ, Gregory PD, Urnov FD, Amacher SL (2008) Heritable targeted gene disruption in zebrafish using designed zinc-finger nucleases. Nat Biotechnol 26:702–708. doi:10.1038/nbt1409
Epinat JC, Arnould S, Chames P, Rochaix P, Desfontaines D, Puzin C, Patin A, Zanghellini A, Paques F, Lacroix E (2003) A novel engineered meganuclease induces homologous recombination in yeast and mammalian cells. Nucleic Acids Res 31:2952–2962. doi:10.1093/nar/gkg375
Gao H, Yang M, Falco SC, Lyznik A (2008) Locus-specific mutations induced by homing endonucleases in maize. FAO/IAEA International Symposium on induced mutations in plants. Vienna, Austria, pp 86
Gidoni D, Bar M, Leshem B, Gilboa N, Mett A, Feiler J (2001) Embryonal recombination and germline inheritance of recombined FRT loci mediated by constitutively expressed FLP in tobacco. Euphytica 121:145–156. doi:10.1023/A:1012081125631
Gisler B, Salomon S, Puchta H (2002) The role of double-strand break-induced allelic homologous recombination in somatic plant cells. Plant J 32:277–284. doi:10.1046/j.1365-313X.2002.01421.x
Lloyd A, Plaisier CL, Carroll D, Drews GN (2005) Targeted mutagenesis using zinc-finger nucleases in Arabidopsis. Proc Natl Acad Sci USA 102:2232–2237. doi:10.1073/pnas.0409339102
Lyznik LA, McGee JD, Tung PY, Bennetzen JL, Hodges TK (1991) Homologous recombination between plasmid DNA molecules in maize protoplasts. Mol Gen Genet 230:209–218. doi:10.1007/BF00290670
Maeder ML, Thibodeau-Beganny S, Osiak A, Wright DA, Anthony RM, Eichtinger M, Jiang T, Foley JE, Winfrey RJ, Townsend JA, Unger-Wallace E, Sander JD, Muller-Lerch F, Fu F, Pearlberg J, Gobel C, Dassie JP, Pruett-Miller SM, Porteus MH, Sgroi DC, Iafrate AJ, Dobbs D, McCray PB Jr, Cathomen T, Voytas DF, Joung JK (2008) Rapid “open-source” engineering of customized zinc-finger nucleases for highly efficient gene modification. Mol Cell 31:294–301. doi:10.1016/j.molcel.2008.06.016
Matz MV, Fradkov AF, Labas YA, Savitsky AP, Zaraisky AG, Markelov ML, Lukyanov SA (1999) Fluorescent proteins from nonbioluminescent Anthozoa species. Nat Biotechnol 17:969–973. doi:10.1038/13657
McElroy D, Blowers AD, Jenes B, Wu R (1991) Construction of expression vectors based on the rice actin 1 (Act1) 5′ region for use in monocot transformation. Mol Gen Genet 231:150–160. doi:10.1007/BF00293832
Meng X, Noyes MB, Zhu LJ, Lawson ND, Wolfe SA (2008) Targeted gene inactivation in zebrafish using engineered zinc-finger nucleases. Nat Biotechnol 26:695–701. doi:10.1038/nbt1398
Moehle EA, Rock JM, Lee YL, Jouvenot Y, DeKelver RC, Gregory PD, Urnov FD, Holmes MC (2007) Targeted gene addition into a specified location in the human genome using designed zinc finger nucleases. Proc Natl Acad Sci USA 104:3055–3060. doi:10.1073/pnas.0611478104
Morton J, Davis MW, Jorgensen EM, Carroll D (2006) Induction and repair of zinc-finger nuclease-targeted double-strand breaks in Caenorhabditis elegans somatic cells. Proc Natl Acad Sci USA 103:16370–16375. doi:10.1073/pnas.0605633103
Orel N, Kyryk A, Puchta H (2003) Different pathways of homologous recombination are used for the repair of double-strand breaks within tandemly arranged sequences in the plant genome. Plant J 35:604–612. doi:10.1046/j.1365-313X.2003.01832.x
Porteus MH, Baltimore D (2003) Chimeric nucleases stimulate gene targeting in human cells. Science 300:763. doi:10.1126/science.1078395
Puchta H (2005) The repair of double-strand breaks in plants: mechanisms and consequences for genome evolution. J Exp Bot 56:1–14. doi:10.1093/jxb/eri123
Redondo P, Prieto J, Munoz IG, Alibes A, Stricher F, Serrano L, Cabaniols JP, Daboussi F, Arnould S, Perez C, Duchateau P, Paques F, Blanco FJ, Montoya G (2008) Molecular basis of Xeroderma pigmentosum group C DNA recognition by engineered meganucleases. Nature 456:107–111. doi:10.1038/nature07343
Rong YS, Titen SW, Xie HB, Golic MM, Bastiani M, Bandyopadhyay P, Olivera BM, Brodsky M, Rubin GM, Golic KG (2002) Targeted mutagenesis by homologous recombination in D. melanogaster. Genes Dev 16:1568–1581. doi:10.1101/gad.986602
Rosen LE, Morrison HA, Masri S, Brown MJ, Springstubb B, Sussman D, Stoddard BL, Seligman LM (2006) Homing endonuclease I-CreI derivatives with novel DNA target specificities. Nucleic Acids Res 34:4791–4800. doi:10.1093/nar/gkl645
Rouet P, Smih F, Jasin M (1994) Introduction of double-strand breaks into the genome of mouse cells by expression of a rare-cutting endonuclease. Mol Cell Biol 14:8096–8106
Salomon S, Puchta H (1998) Capture of genomic and T-DNA sequences during double-strand break repair in somatic plant cells. EMBO J 17:6086–6095. doi:10.1093/emboj/17.20.6086
Sorrell DA, Kolb AF (2005) Targeted modification of mammalian genomes. Biotechnol Adv 23:431–469. doi:10.1016/j.biotechadv.2005.03.003
Szczepek M, Brondani V, Buchel J, Serrano L, Segal DJ, Cathomen T (2007) Structure-based redesign of the dimerization interface reduces the toxicity of zinc-finger nucleases. Nat Biotechnol 25:786–793. doi:10.1038/nbt1317
Truett GE, Heeger P, Mynatt RL, Truett AA, Walker JA, Warman ML (2000) Preparation of PCR-quality mouse genomic DNA with hot sodium hydroxide and tris (HotSHOT). Biotechniques 29:52–54
Ugaki M, Ueda T, Timmermans MC, Vieira J, Elliston KO, Messing J (1991) Replication of a geminivirus derived shuttle vector in maize endosperm cells. Nucleic Acids Res 19:371–377. doi:10.1093/nar/19.2.371
Urnov FD, Miller JC, Lee YL, Beausejour CM, Rock JM, Augustus S, Jamieson AC, Porteus MH, Gregory PD, Holmes MC (2005) Highly efficient endogenous human gene correction using designed zinc-finger nucleases. Nature 435:646–651. doi:10.1038/nature03556
Vanderschuren H, Stupak M, Futterer J, Gruissem W, Zhang P (2007) Engineering resistance to geminiviruses–review and perspectives. Plant Biotechnol J 5:207–220. doi:10.1111/j.1467-7652.2006.00217.x
Wright DA, Townsend JA, Winfrey RJ Jr, Irwin PA, Rajagopal J, Lonosky PM, Hall BD, Jondle MD, Voytas DF (2005) High-frequency homologous recombination in plants mediated by zinc-finger nucleases. Plant J 44:693–705. doi:10.1111/j.1365-313X.2005.02551.x
Xiao YL, Li X, Peterson T (2000) Ac insertion site affects the frequency of transposon-induced homologous recombination at the maize p1 locus. Genetics 156:2007–2017
Zhang W, Subbarao S, Addae P, Shen A, Armstrong C, Peschke V, Gilbertson L (2003) Cre/lox-mediated marker gene excision in transgenic maize (Zea mays L.) plants. Theor Appl Genet 107:1157–1168. doi:10.1007/s00122-003-1368-z
Zhao X, Coats I, Fu P, Gordon-Kamm B, Lyznik LA (2003) T-DNA recombination and replication in maize cells. Plant J 33:149–159. doi:10.1046/j.1365-313X.2003.016016.x
Acknowledgments
The authors thank Shifu Zhen and Susan Nilles for technical assistance in producing transgenic plants. We acknowledge Carl Simmons’ contribution to this work by designing the maize codon-optimized I-SceI. We thank Cellectis, S.A. for providing the I-Sce endonuclease and for helpful research discussions.
Novel materials described in this publication may be available for non-commercial research purposes upon acceptance and signing of a material transfer agreement. In some cases such materials may contain or be derived from materials obtained from a third party. In such cases, distribution of material will be subject to the requisite permission from any third-party owners, licensors or controllers of all or parts of the material. Plant germplasm and transgenic material will not be made available except at the discretion of the owner and then only in accordance with all applicable governmental regulations. Obtaining any permissions will be the sole responsibility of the requestor.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yang, M., Djukanovic, V., Stagg, J. et al. Targeted mutagenesis in the progeny of maize transgenic plants. Plant Mol Biol 70, 669–679 (2009). https://doi.org/10.1007/s11103-009-9499-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-009-9499-5