Abstract
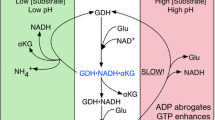
Mammalian glutamate dehydrogenase (GDH) catalyzes the reversible inter-conversion of glutamate to α-ketoglutarate and ammonia, interconnecting carbon skeleton and nitrogen metabolism. In addition, it functions as an energy switch by its ability to fuel the Krebs cycle depending on the energy status of the cell. As GDH lies at the intersection of several metabolic pathways, its activity is tightly regulated by several allosteric compounds that are metabolic intermediates. In contrast to other mammals that have a single GDH-encoding gene, humans and great apes possess two isoforms of GDH (hGDH1 and hGDH2, encoded by the GLUD1 and GLUD2 genes, respectively) with distinct regulation pattern, but remarkable sequence similarity (they differ, in their mature form, in only 15 of their 505 amino-acids). The GLUD2 gene is considered a very young gene, emerging from the GLUD1 gene through retro-position only recently (<23 million years ago). The new hGDH2 iso-enzyme, through random mutations and natural selection, is thought to have conferred an evolutionary advantage that helped its persistence through primate evolution. The properties of the two highly homologous human GDHs have been studied using purified recombinant hGDH1 and hGDH2 proteins obtained by expression of the corresponding cDNAs in Sf21 cells. According to these studies, in contrast to hGDH1 that maintains basal activity at 35–40 % of its maximal, hGDH2 displays low basal activity that is highly responsive to activation by rising levels of ADP and/or l-leucine which can also act synergistically. While hGDH1 is inhibited potently by GTP, hGDH2 shows remarkable GTP resistance. Furthermore, the two iso-enzymes are differentially inhibited by estrogens, polyamines and neuroleptics, and also differ in heat-lability. To elucidate the molecular mechanisms that underlie these different regulation patterns of the two iso-enzymes (and consequently the evolutionary adaptation of hGDH2 to a new functional role), we have performed mutagenesis at sites of difference in their amino acid sequence. Results showed that the low basal activity, heat-lability and estrogen sensitivity of hGDH2 could be, at least partially, ascribed to the Arg443Ser evolutionary change, whereas resistance to GTP inhibition has been attributed to the Gly456Ala change. Other amino acid substitutions studied thus far cannot explain all the remaining functional differences between the two iso-enzymes. Also, the Arg443Ser/Gly456Ala double mutation in hGDH1 approached the properties of wild-type hGDH2, without being identical to it. The insights into the structural mechanism of enzymatic regulation and the implications in cell biology provided by these findings are discussed.






Similar content being viewed by others
References
Hudson R, Daniel R (1993) L-glutamate dehydrogenases: distribution, properties and mechanism. Comp Biochem Physiol B 106:767–792
Smith TJ, Stanley CA (2008) Untangling the glutamate dehydrogenase allosteric nightmare. Trends Biochem Sci 33:557–564
McKenna MC, Tildon JT, Stevenson JH, Huang X (1996) New insights into the compartmentation of glutamate and glutamine in cultured rat brain astrocytes. Dev Neurosci 18:380–390
McKenna MC (2007) The glutamate–glutamine cycle is not stoichiometric: fates of glutamate in brain. J Neurosci Res 85:3347–3358
Raimundo N, Baysal BE, Shadel GS (2011) Revisiting the TCA cycle: signaling to tumor formation. Trends Mol Med 17:641–649
Lambeth DO (2002) What is the function of GTP produced in the Krebs citric acid cycle? IUBMB Life 54:143–144
Shashidharan P, Clarke DD, Ahmed N, Moschonas N, Plaitakis A (1997) Nerve tissue-specific human glutamate dehydrogenase that is thermolabile and highly regulated by ADP. J Neurochem 68:1804–1811
Owen OE, Kalhan SC, Hanson RW (2002) The key role of anaplerosis and cataplerosis for citric acid cycle function. J Biol Chem 277:30409–30412
Sener A, Malaisse-Lagae F, Malaisse W (1981) Stimulation of pancreatic islet metabolism and insulin release by a nonmetabolizable amino acid. Proc Natl Acad Sci USA 78:5460–5464
Plaitakis A, Zaganas I (2001) Regulation of human glutamate dehydrogenases: implications for glutamate, ammonia and energy metabolism in brain. J Neurosci Res 66:899–908
Cooper AL (2012) The role of glutamine synthetase and glutamate dehydrogenase in cerebral ammonia homeostasis. Neurochem Res 37:2439–2455
Li M, Li C, Allen A, Stanley C, Smith T (2014) Glutamate dehydrogenase: structure, allosteric regulation, and role in insulin homeostasis. Neurochem Res. doi:10.1007/s11064-013-1173-2
Peterson PE, Smith TJ (1999) The structure of bovine glutamate dehydrogenase provides insights into the mechanism of allostery. Struct Fold Des 7:769–782
Smith TJ, Schmidt T, Fang J, Wu J, Siuzdak G, Stanley CA (2002) The structure of apo human glutamate dehydrogenase details subunit communication and allostery. J Mol Biol 318:765–777
Smith TJ, Peterson PE, Schmidt T, Fang J, Stanley CA (2001) Structures of bovine glutamate dehydrogenase complexes elucidate the mechanism of purine regulation. J Mol Biol 307:707–720
Shashidharan P, Michaelidis TM, Robakis NK, Kresovali A, Papamatheakis J, Plaitakis A (1994) Novel human glutamate dehydrogenase expressed in neural and testicular tissues and encoded by an X-linked intronless gene. J Biol Chem 269:16971–16976
Burki F, Kaessmann H (2004) Birth and adaptive evolution of a hominoid gene that supports high neurotransmitter flux. Nat Genet 36:1061–1063
Mavrothalassitis G, Tzimagiorgis G, Mitsialis A, Zannis V, Plaitakis A, Papamatheakis J, Moschonas N (1988) Isolation and characterization of cDNA clones encoding human liver glutamate dehydrogenase: evidence for a small gene family. Proc Natl Acad Sci USA 85:3494–3498
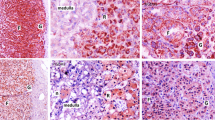
Spanaki C, Zaganas I, Kleopa KA, Plaitakis A (2010) Human GLUD2 glutamate dehydrogenase is expressed in neural and testicular supporting cells. J Biol Chem 285:16748–16756
Tomita T, Kuzuyama T, Nishiyama M (2011) Structural basis for leucine-induced allosteric activation of glutamate dehydrogenase. J Biol Chem 286:37406–37413
Zaganas I, Spanaki C, Plaitakis A (2012) Expression of human GLUD2 glutamate dehydrogenase in human tissues: functional implications. Neurochem Int 61:455–462
Ciomborowska J, Rosikiewicz W, Szklarczyk D, Makałowski W, Makałowska I (2013) Orphan retrogenes in the human genome. Mol Biol Evol 30:384–396
Plaitakis A, Latsoudis H, Kanavouras K, Ritz B, Bronstein JM, Skoula I, Mastorodemos V, Papapetropoulos S, Borompokas N, Zaganas I, Xiromerisiou G, Hadjigeorgiou GM, Spanaki C (2010) Gain-of-function variant in GLUD2 glutamate dehydrogenase modifies Parkinson’s disease onset. Eur J Hum Genet 18:336–341
Colon A, Plaitakis A, Perakis A, Berl S, Clarke D (1986) Purification and characterization of a soluble and a particulate glutamate dehydrogenase from rat brain. J Neurochem 46:1811–1819
Zaganas I, Plaitakis A (2002) Single amino acid substitution (G456A) in the vicinity of the GTP binding domain of human housekeeping glutamate dehydrogenase markedly attenuates GTP inhibition and abolishes the cooperative behavior of the enzyme. J Biol Chem 277:26422–26428
Mastorodemos V, Kotzamani D, Zaganas I, Arianoglou G, Latsoudis H, Plaitakis A (2009) Human GLUD1 and GLUD2 glutamate dehydrogenase localize to mitochondria and endoplasmic reticulum. Biochem Cell Biol 87:505–516
McCarthy AD, Walker JM, Tipton KF (1980) Purification of glutamate dehydrogenase from ox brain and liver. Evidence that commercially available preparations of the enzyme from ox liver have suffered proteolytic cleavage. Biochem J 191:605–611
Yang S-J, Huh J-W, Hong H-N, Kim TU, Cho S-W (2004) Important role of Ser443 in different thermal stability of human glutamate dehydrogenase isozymes. FEBS Lett 562:59–64
Kanavouras K, Mastorodemos V, Borompokas N, Spanaki C, Plaitakis A (2007) Properties and molecular evolution of human GLUD2 (neural and testicular tissue-specific) glutamate dehydrogenase. J Neurosci Res 85:3398–3406
Plaitakis A, Metaxari M, Shashidharan P (2000) Nerve tissue-specific (GLUD2) and housekeeping (GLUD1) human glutamate dehydrogenases are regulated by distinct allosteric mechanisms. Implications for biologic function. J Neurochem 75:1862–1869
Plaitakis A, Spanaki C, Mastorodemos V, Zaganas I (2003) Study of structure–function relationships in human glutamate dehydrogenases reveals novel molecular mechanisms for the regulation of the nerve tissue-specific (GLUD2) isoenzyme. Neurochem Int 43:401–410
Zaganas I, Kanavouras K, Mastorodemos V, Latsoudis H, Spanaki C, Plaitakis A (2009) The human GLUD2 glutamate dehydrogenase: localization and functional aspects. Neurochem Int 55:52–63
Choi M-M, Kim E-A, Yang S-J, Choi SY, Cho S-W, Huh J-W (2007) Amino acid changes within antenna helix are responsible for different regulatory preferences of human glutamate dehydrogenase isozymes. J Biol Chem 282:19510–19517
Plaitakis A, Latsoudis H, Spanaki C (2011) The human GLUD2 glutamate dehydrogenase and its regulation in health and disease. Neurochem Int 59:495–509
Spanaki C, Zaganas I, Kounoupa Z, Plaitakis A (2012) The complex regulation of human glud1 and glud2 glutamate dehydrogenases and its implications in nerve tissue biology. Neurochem Int 61:470–481
Zaganas I, Spanaki C, Karpusas M, Plaitakis A (2002) Substitution of Ser for Arg-443 in the regulatory domain of human housekeeping (GLUD1) glutamate dehydrogenase virtually abolishes basal activity and markedly alters the activation of the enzyme by ADP and l-leucine. J Biol Chem 277:46552–46558
Mastorodemos V, Zaganas I, Spanaki C, Bessa M, Plaitakis A (2005) Molecular basis of human glutamate dehydrogenase regulation under changing energy demands. J Neurosci Res 79:65–73
Stanley CA, Lieu YK, Hsu BYL, Burlina AB, Greenberg CR, Hopwood NJ, Perlman K, Rich BH, Zammarchi E, Poncz M (1998) Hyperinsulinism and hyperammonemia in infants with regulatory mutations of the glutamate dehydrogenase gene. N Engl J Med 338:1352–1357
Lee E-Y, Yoon H-Y, Ahn J-Y, Choi SY, Cho S-W (2001) Identification of the GTP binding site of human glutamate dehydrogenase by cassette mutagenesis and photoaffinity labeling. J Biol Chem 276:47930–47936
Borompokas N, Papachatzaki M-M, Kanavouras K, Mastorodemos V, Zaganas I, Spanaki C, Plaitakis A (2010) Estrogen modification of human glutamate dehydrogenases is linked to enzyme activation state. J Biol Chem 285:31380–31387
Choi M, Hwang E, Kim E, Huh J, Cho S (2008) Identification of amino acid residues responsible for different GTP preferences of human glutamate dehydrogenase isozymes. Biochem Biophys Res Commun 368:742–747
Kelly A, Stanley C (2001) Disorders of glutamate metabolism. Ment Retard Dev Disabil Res Rev 7:287–295
Fang J, Hsu B, MacMullen C, Poncz M, Smith T, Stanley C (2002) Expression, purification and characterization of human glutamate dehydrogenase (GDH) allosteric regulatory mutations. Biochem J 363:81–87
Lee E-Y, Huh J-W, Yang S-J, Choi SY, Cho S-W, Choi HJ (2003) Histidine 454 plays an important role in polymerization of human glutamate dehydrogenase. FEBS Lett 540:163–166
Kanavouras K, Borompokas N, Latsoudis H, Stagourakis A, Zaganas I, Plaitakis A (2009) Mutations in human GLUD2 glutamate dehydrogenase affecting basal activity and regulation. J Neurochem 109:167–173
Banerjee S, Schmidt T, Fang J, Stanley CA, Smith TJ (2003) Structural studies on ADP activation of mammalian glutamate dehydrogenase and the evolution of regulation. Biochemistry 42:3446–3456
Petersen RC, Smith GE, Waring SC, Ivnik RJ, Tangalos EG, Kokmen E (1999) Mild cognitive impairment: clinical characterization and outcome. Arch Neurol 56:303–308
Allen A, Kwagh J, Fang J, Stanley CA, Smith TJ (2004) Evolution of glutamate dehydrogenase regulation of insulin homeostasis is an example of molecular exaptation. Biochemistry 43:14431–14443
Stanley CA (2011) Two genetic forms of hyperinsulinemic hypoglycemia caused by dysregulation of glutamate dehydrogenase. Neurochem Int 59:465–472
Meshkini A, Yazdanparast R, Nouri K (2011) Intracellular GTP level determines cell’s fate toward differentiation and apoptosis. Toxicol Appl Pharmacol 253:188–196
Treberg JR, Clow KA, Greene KA, Brosnan ME, Brosnan JT (2010) Systemic activation of glutamate dehydrogenase increases renal ammoniagenesis: implications for the hyperinsulinism/hyperammonemia syndrome. Am J Physiol Endocrinol Metab 298:E1219–E1225
Wheeler LJ, Mathews CK (2011) Nucleoside triphosphate pool asymmetry in mammalian mitochondria. J Biol Chem 286:16992–16996
Attwell D, Laughlin SB (2001) An energy budget for signaling in the grey matter of the brain. J Cereb Blood Flow Metab 21:1133–1145
Schousboe A, Sickmann HM, Bak LK, Schousboe I, Jajo FS, Faek SAA, Waagepetersen HS (2011) Neuron–glia interactions in glutamatergic neurotransmission: roles of oxidative and glycolytic adenosine triphosphate as energy source. J Neurosci Res 89:1926–1934
Traut TW (1994) Physiological concentrations of purines and pyrimidines. Mol Cell Biochem 140:1–22
Metelkin E, Demin O, Kovacs Z, Chinopoulos C (2009) Modeling of ATP–ADP steady-state exchange rate mediated by the adenine nucleotide translocase in isolated mitochondria. FEBS J 276:6942–6955
Murthy MS, Pande SV (1985) Microcompartmentation of transported carnitine, acetylcarnitine and ADP occurs in the mitochondrial matrix. Implications for transport measurements and metabolism. Biochem J 230:657–663
Rottenberg H, Lee CP (1975) Energy dependent hydrogen ion accumulation in submitochondrial particles. Biochemistry 14:2675–2680
Schoolwerth A, LaNoue K, Hoover W (1984) Effect of pH on glutamate efflux from rat kidney mitochondria. Am J Physiol 246:F266–F271
Llopis J, McCaffery JM, Miyawaki A, Farquhar MG, Tsien RY (1998) Measurement of cytosolic, mitochondrial, and Golgi pH in single living cells with green fluorescent proteins. Proc Natl Acad Sci USA 95:6803–6808
Balut C, vandeVen M, Despa S, Lambrichts I, Ameloot M, Steels P, Smets I (2008) Measurement of cytosolic and mitochondrial pH in living cells during reversible metabolic inhibition. Kidney Int 73:226–232
Zaganas I, Pajecka K, Wendel Nielsen C, Schousboe A, Waagepetersen HS, Plaitakis A (2013) The effect of pH and ADP on ammonia affinity for human glutamate dehydrogenases. Metab Brain Dis 28:127–131
Schoolwerth AC, Nazar BL, LaNoue KF (1978) Glutamate dehydrogenase activation and ammonia formation by rat kidney mitochondria. J Biol Chem 253:6177–6183
Bouvier M, Szatkowski M, Amato A, Attwell D (1992) The glial cell glutamate uptake carrier countertransports pH-changing anions. Nature 360:471–474
Poitry S, Poitry-Yamate C, Ueberfeld J, MacLeish P, Tsacopoulos M (2000) Mechanisms of glutamate metabolic signaling in retinal glial (Mόller) cells. J Neurosci 20:1809–1821
Azarias G, Perreten H, Lengacher S, Poburko D, Demaurex N, Magistretti PJ, Chatton J-Y (2011) Glutamate transport decreases mitochondrial pH and modulates oxidative metabolism in astrocytes. J Neurosci 31:3550–3559
Yu AC, Schousboe A, Hertz L (1982) Metabolic fate of 14C-labeled glutamate in astrocytes in primary cultures. J Neurochem 39:954–960
McKenna MC, Sonnewald U, Huang X, Stevenson J, Zielke HR (1996) Exogenous glutamate concentration regulates the metabolic fate of glutamate in astrocytes. J Neurochem 66:386–393
Sonnewald U, Westergaard N, Schousboe A (1997) Glutamate transport and metabolism in astrocytes. Glia 21:56–63
Whitelaw BS, Robinson MB (2013) Inhibitors of glutamate dehydrogenase block sodium-dependent glutamate uptake in rat brain membranes. Front Endocrinol (Lausanne) 4:123
Palmieri F (2013) The mitochondrial transporter family SLC25: identification, properties and physiopathology. Mol Asp Med 34:465–484
Berkich DA, Ola MS, Cole J, Sweatt AJ, Hutson SM, LaNoue KF (2007) Mitochondrial transport proteins of the brain. J Neurosci Res 85:3367–3377
Dennis SC, Clark JB (1977) The pathway of glutamate metabolism in rat brain mitochondria. Biochem J 168:521–527
Aoki C, Milner TA, Berger SB, Sheu KF, Blass JP, Pickel VM (1987) Glial glutamate dehydrogenase: ultrastructural localization and regional distribution in relation to the mitochondrial enzyme, cytochrome oxidase. J Neurosci Res 18:305–318
Zaganas I, Waagepetersen HS, Georgopoulos P, Sonnewald U, Plaitakis A, Schousboe A (2001) Differential expression of glutamate dehydrogenase in cultured neurons and astrocytes from mouse cerebellum and cerebral cortex. J Neurosci Res 66:909–913
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948
Page R (1996) TreeView: an application to display phylogenetic trees on personal computers. Comput Appl Biosci 12:357–358
Acknowledgments
Most of the research described here has been performed under the guidance of Dr. Andreas Plaitakis, to whom this issue of Neurochemical Research is dedicated to. We (I.V.Z., K.K., V.M., H.L., N.B., G.A. and C.D.) are grateful for his help and inspiration during the last 18 years. This manuscript has been partially co-financed by the European Union (European Social Fund—ESF) and Greek national funds through the Operational Program “Education and Lifelong Learning” of the National Strategic Reference Framework (NSRF)—Research Funding Program: THALIS—UOC, Title “Mitochondrial dysfunction in neurodegenerative diseases” (Grant Code 377226).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zaganas, I.V., Kanavouras, K., Borompokas, N. et al. The Odyssey of a Young Gene: Structure–Function Studies in Human Glutamate Dehydrogenases Reveal Evolutionary-Acquired Complex Allosteric Regulation Mechanisms. Neurochem Res 39, 471–486 (2014). https://doi.org/10.1007/s11064-014-1251-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11064-014-1251-0