Abstract
Introduction
Ependymoma is the third most common malignant pediatric brain tumor. Although the biology that drives ependymoma is slowly being unraveled, the ability to translate these findings to clinical care remains an ongoing challenge. Epigenetic alterations appear to play a central role in the development of molecular classification of ependymoma.
Methods
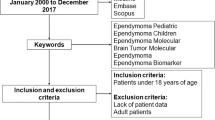
We reviewed the published literature available describing genetic and epigenetic underpinnings of ependymoma that have been reported to date and have summarized the information regarding genetic drivers of ependymoma that may point us toward therapeutic strategies.
Results
Ependymoma is a molecularly heterogeneous disease which has now been divided into at least nine distinct molecular subtypes based on DNA methylation and gene expression profiling. DNA methylation has emerged as an effective tool for classification of brain tumors alongside histopathology and other molecular diagnostics. There have been large retrospective cohorts describing molecular subgroup identity as a powerful independent predictor of outcome. There is limited published data on prospective trials to date however this is forthcoming which will lead to molecular stratification in the next generation of clinical studies.
Conclusion
This is a review of recent advancements in our understanding of the epigenetic basis of ependymoma and discussion of how these findings reveal potential therapeutic opportunities.
Similar content being viewed by others
References
Lee SH, Chung CK, Kim CH, Yoon SH, Hyun SJ, Kim KJ et al (2013) Long-term outcomes of surgical resection with or without adjuvant radiation therapy for treatment of spinal ependymoma: a retrospective multicenter study by the Korea Spinal Oncology Research Group. Neuro-Oncology 15(7):921–929
Metellus P, Barrie M, Figarella-Branger D, Chinot O, Giorgi R, Gouvernet J et al (2007) Multicentric French study on adult intracranial ependymomas: prognostic factors analysis and therapeutic considerations from a cohort of 152 patients. Brain : J Neurol 130(Pt 5):1338–1349
Ostrom QT, Gittleman H, Liao P, Rouse C, Chen Y, Dowling J et al (2014) CBTRUS statistical report: primary brain and central nervous system tumors diagnosed in the United States in 2007–2011. Neuro-Oncology. https://doi.org/10.1093/neuonc/nou223
Taylor MD, Poppleton H, Fuller C, Su X, Liu Y, Jensen P et al (2005) Radial glia cells are candidate stem cells of ependymoma. Cancer Cell 8(4):323–335
Wu J, Armstrong TS, Gilbert MR (2016) Biology and management of ependymomas. Neuro-Oncology 18(7):902–913
Merchant TE, Li C, Xiong X, Kun LE, Boop FA, Sanford RA (2009) Conformal radiotherapy after surgery for paediatric ependymoma: a prospective study. Lancet Oncol 10(3):258–266
Pajtler KW, Mack SC, Ramaswamy V, Smith CA, Witt H, Smith A et al (2017) The current consensus on the clinical management of intracranial ependymoma and its distinct molecular variants. Acta Neuropathol 133(1):5–12
Mack SC, Witt H, Piro RM, Gu L, Zuyderduyn S, Stutz AM et al (2014) Epigenomic alterations define lethal CIMP-positive ependymomas of infancy. Nature 506(7489):445–450
Ramaswamy V, Hielscher T, Mack SC, Lassaletta A, Lin T, Pajtler KW et al (2016) Therapeutic impact of cytoreductive surgery and irradiation of posterior fossa ependymoma in the molecular era: a retrospective multicohort analysis. J Clin Oncol 34(21):2468–2477
Pajtler KW, Wen J, Sill M, Lin T, Orisme W, Tang B et al (2018) Molecular heterogeneity and CXorf67 alterations in posterior fossa group A (PFA) ependymomas. Acta Neuropathol 136(2):211–226
Pajtler KW, Witt H, Sill M, Jones DT, Hovestadt V, Kratochwil F et al (2015) Molecular classification of ependymal tumors across all CNS compartments, histopathological grades, and age groups. Cancer Cell 27(5):728–743
Capper D, Jones DTW, Sill M, Hovestadt V, Schrimpf D, Sturm D et al (2018) DNA methylation-based classification of central nervous system tumours. Nature 555(7697):469–474
Mack SC, Pajtler KW, Chavez L, Okonechnikov K, Bertrand KC, Wang X et al (2018) Therapeutic targeting of ependymoma as informed by oncogenic enhancer profiling. Nature 553(7686):101–105
Ozawa T, Arora S, Szulzewsky F, Juric-Sekhar G, Miyajima Y, Bolouri H et al (2018) A de novo mouse model of C11orf95-RELA fusion-driven ependymoma identifies driver functions in addition to NF-kappaB. Cell Rep 23(13):3787–3797
Parker M, Mohankumar KM, Punchihewa C, Weinlich R, Dalton JD, Li Y et al (2014) C11orf95-RELA fusions drive oncogenic NF-kappaB signalling in ependymoma. Nature 506(7489):451–455
Griesinger AM, Witt DA, Grob ST, Georgio Westover SR, Donson AM, Sanford B et al (2017) NF-κB upregulation through epigenetic silencing of LDOC1 drives tumor biology and specific immunophenotype in Group A ependymoma. Neuro-Oncology 19(10):1350–1360
Chen LF, Mu Y, Greene WC (2002) Acetylation of RelA at discrete sites regulates distinct nuclear functions of NF-kappaB. EMBO J 21(23):6539–6548
Hoesel B, Schmid JA (2013) The complexity of NF-κB signaling in inflammation and cancer. Molecular Cancer 12:86
Huang B, Yang XD, Lamb A, Chen LF (2010) Posttranslational modifications of NF-kappaB: another layer of regulation for NF-kappaB signaling pathway. Cell Signal 22(9):1282–1290
Andreiuolo F, Varlet P, Tauziède-Espariat A, Jünger ST, Dörner E, Dreschmann V et al (2019) Childhood supratentorial ependymomas with YAP1-MAMLD1 fusion: an entity with characteristic clinical, radiological, cytogenetic and histopathological features. Brain Pathol (Zurich, Switzerland) 29(2):205–216
Pajtler KW, Wei Y, Okonechnikov K, Silva PBG, Vouri M, Zhang L et al (2019) YAP1 subgroup supratentorial ependymoma requires TEAD and nuclear factor I-mediated transcriptional programmes for tumorigenesis. Nature Commun 10(1):3914
Lester A, McDonald KL (2020) Intracranial ependymomas: molecular insights and translation to treatment. Brain Pathol (Zurich, Switzerland) 30(1):3–12
Totaro A, Panciera T, Piccolo S (2018) YAP/TAZ upstream signals and downstream responses. Nat Cell Biol 20(8):888–899
Yu FX, Zhang Y, Park HW, Jewell JL, Chen Q, Deng Y et al (2013) Protein kinase a activates the hippo pathway to modulate cell proliferation and differentiation. Genes Dev 27(11):1223–1232
Zhao B, Tumaneng K, Guan KL (2011) The hippo pathway in organ size control, tissue regeneration and stem cell self-renewal. Nat Cell Biol 13(8):877–883
Hübner JM, Müller T, Papageorgiou DN, Mauermann M, Krijgsveld J, Russell RB et al (2019) EZHIP/CXorf67 mimics K27M mutated oncohistones and functions as an intrinsic inhibitor of PRC2 function in aggressive posterior fossa ependymoma. Neuro-Oncology 21(7):878–889
Jain SU, Do TJ, Lund PJ, Rashoff AQ, Diehl KL, Cieslik M et al (2019) PFA ependymoma-associated protein EZHIP inhibits PRC2 activity through a H3 K27M-like mechanism. Nature Commun 10(1):2146
Piunti A, Smith ER, Morgan MAJ, Ugarenko M, Khaltyan N, Helmin KA et al (2019) CATACOMB: An endogenous inducible gene that antagonizes H3K27 methylation activity of Polycomb repressive complex 2 via an H3K27M-like mechanism. Sci Adv. https://doi.org/10.1126/sciadv.aax2887
Panwalkar P, Clark J, Ramaswamy V, Hawes D, Yang F, Dunham C et al (2017) Immunohistochemical analysis of H3K27me3 demonstrates global reduction in group-A childhood posterior fossa ependymoma and is a powerful predictor of outcome. Acta Neuropathol 134(5):705–714
Pratt D, Quezado M, Abdullaev Z, Hawes D, Yang F, Garton HJL et al (2020) Diffuse intrinsic pontine glioma-like tumor with EZHIP expression and molecular features of PFA ependymoma. Acta Neuropathologica Commun 8(1):37
Wu G, Broniscer A, McEachron TA, Lu C, Paugh BS, Becksfort J et al (2012) Somatic histone H3 alterations in pediatric diffuse intrinsic pontine gliomas and non-brainstem glioblastomas. Nat Genet 44(3):251–253
Schwartzentruber J, Korshunov A, Liu XY, Jones DT, Pfaff E, Jacob K et al (2012) Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma. Nature 482(7384):226–231
de Koning AP, Gu W, Castoe TA, Batzer MA, Pollock DD (2011) Repetitive elements may comprise over two-thirds of the human genome. PLoS Genet 7(12):e1002384
Henssen AG, Koche R, Zhuang J, Jiang E, Reed C, Eisenberg A et al (2017) PGBD5 promotes site-specific oncogenic mutations in human tumors. Nat Genet 49(7):1005–1014
Henssen AG, Kentsis A (2018) Emerging functions of DNA transposases and oncogenic mutators in childhood cancer development. JCI Insight 3(20):e123–172
Krug B, De Jay N, Harutyunyan AS, Deshmukh S, Marchione DM, Guilhamon P et al (2019) Pervasive H3K27 acetylation leads to ERV expression and a therapeutic vulnerability in H3K27M gliomas. Cancer Cell 35(5):782–97.e8
Acknowledgements
S.C.M. is supported by a Cancer Prevention Research Institute of Texas (CPRIT) scholar award (RR170023), an Alex s Lemonade Stand Foundation (ALSF) A award and Young Investigator award, a Pediatric Brain Tumor Foundation award, a Chad Tough Young Investigator award, a Cookies for Cancer research grant, a RALLY research grant, a BEAR Necessities Pediatric Cancer Foundation grant, a Children's Cancer Research Fund award, a Children's Brain Tumor Foundation award, and a Baylor College of Medicine Junior Faculty award.
Author information
Authors and Affiliations
Contributions
All authors contributed to the review conception and design. Material preparation, literature search, and data analysis was performed by AS, KB, PW, AS, and SM. The first draft of the manuscript was written by AS, KB, and PW and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors of this manuscript have no conflicts of interest to declare.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Stuckert, A., Bertrand, K.C., Wang, P. et al. Weighing ependymoma as an epigenetic disease. J Neurooncol 150, 57–61 (2020). https://doi.org/10.1007/s11060-020-03562-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-020-03562-0