Abstract
Introduction
Temozolomide (TMZ) is a life prolonging DNA alkylating agent active against glioblastomas (GBM) in which the O6-methylguanine-DNA methyltransferase (MGMT) gene is silenced by promoter methylation. Unfortunately acquired TMZ resistance severely undermines its clinical efficacy. Using an in vitro model, we tested whether poly (ADP-ribose) polymerase-1 and -2 (PARP) inhibition could suppress the emergence of resistance to enhance the effectiveness of TMZ.
Methods
Using the MGMT-methylated GBM line U251N, in which TMZ resistance can be induced, we developed a method to rapidly recreate mechanisms of TMZ resistance seen in GBMs, including MMR mutations and MGMT re-expression. We then assessed whether TMZ resistant U251N sub-clones could be re-sensitized to TMZ by co-treatment with the PARP inhibitor ABT-888, and also whether the emergence of resistance could be suppressed by PARP inhibition.
Results
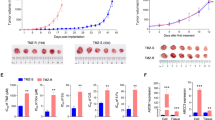
U251N cultures chronically exposed to TMZ developed discrete colonies that expanded during TMZ treatment. These colonies were isolated, expanded further as sub-clones, and assessed for mechanisms of TMZ resistance. Most resistant sub-clones had detectable mutations in one or more mismatch repair (MMR) genes, frequently MSH6, and displayed infrequent re-expression of MGMT. TMZ resistance was associated with isolated poly(ADP-ribose) (pADPr) up-regulation in one sub-clone and was unexplained in several others. TMZ resistant sub-clones regressed during co-treatment with TMZ and ABT-888, and early co-treatment of U251N parental cultures suppressed the emergence of TMZ resistant colonies.
Conclusion
In a model of acquired resistance, co-treatment with TMZ and a PARP inhibitor had two important benefits: re-sensitization of TMZ resistant cells and suppression of TMZ resistance.





Similar content being viewed by others
References
Stupp R, Mason WP, van den Bent MJ et al (2005) Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N Engl J Med 352:987–996
Stupp R, Hegi ME, Mason WP et al (2009) Effects of radiotherapy with concomitant and adjuvant temozolomide versus radiotherapy alone on survival in glioblastoma in a randomised phase III study: 5-year analysis of the EORTC-NCIC trial. Lancet Oncol 10:459–466
Hegi ME, Diserens A, Gorlia T et al (2005) MGMT gene silencing and benefit from temozolomide in glioblastoma. N Engl J Med 352:997–1003
Omuro A, DeAngelis LM (2013) Glioblastoma and other malignant gliomas: a clinical review. JAMA 310:1842–1850
Johnson BE, Mazor T, Hong C et al (2014) Mutational analysis reveals the origin and therapy-driven evolution of recurrent glioma. Science 343:189–193
van Thuijl HF, Mazor T, Johnson BE et al (2015) Evolution of DNA repair defects during malignant progression of low-grade gliomas after temozolomide treatment. Acta Neuropathol 129:597–607
Wang J, Cazzato E, Ladewig E et al (2016) Clonal evolution of glioblastoma under therapy. Nat Genet 48:768–776
Yip S, Miao J, Cahill DP et al (2009) MSH6 mutations arise in glioblastomas during temozolomide therapy and mediate temozolomide resistance. Clin Cancer Res 15:4622–4629. [Correction in (2013) Clin Cancer Res 19:4543–4544]
Maxwell JA, Johnson SP, McLendon RE et al (2008) Mismatch repair deficiency does not mediate clinical resistance to temozolomide in malignant glioma. Clin Cancer Res 14:4859–4868
Felsberg J, Thon N, Eigenbrod S et al (2011) Promoter methylation and expression of MGMT and the DNA mismatch repair genes MLH1, MSH2, MSH6 and PMS2 in paired primary and recurrent glioblastomas. Int J Cancer 129:659–670
Cahill DP, Levine KK, Betensky RA et al (2007) Loss of the mismatch repair protein MSH6 in human glioblastomas is associated with tumor progression during temozolomide treatment. Clin Cancer Res 13:2038–2045
Shinsato Y, Furukawa T, Yunoue S et al (2013) Reduction of MLH1 and PMS2 confers temozolomide resistance and is associated with recurrence of glioblastoma. Oncotarget 4:2261–2270
McFaline-Figueroa JL, Braun CJ, Stanciu M et al (2015) Minor changes in expression of the mismatch repair protein MSH2 exert a major impact on glioblastoma response to temozolomide. Cancer Res 75:3127–3138
Jung TY, Jung S, Moon KS et al (2010) Changes of the O6-methylguanine-DNA methyltransferase promoter methylation and MGMT protein expression after adjuvant treatment in glioblastoma. Oncol Rep 23:1269–1276
Brandes AA, Franceschi E, Tosoni A et al (2010) O(6)-methylguanine DNA-methyltransferase methylation status can change between first surgery for newly diagnosed glioblastoma and second surgery for recurrence: clinical implications. Neuro Oncology 12:283–288
Bobola MS, Kolstoe DD, Blank A et al (2012) Repair of 3-methyladenine and abasic sites by base excision repair mediates glioblastoma resistance to temozolomide. Front Oncol 2:176
Kaur S, Ramdzan ZM, Guiot MC et al (2018) CUX1 stimulates APE1 enzymatic activity and increases the resistance of glioblastoma cells to the mono-alkylating agent temozolomide. Neuro Oncology 20:484–493
Yuan AL, Ricks CB, Bohm AK et al (2018) ABT-888 restores sensitivity in temozolomide resistant glioma cells and xenografts. PLoS ONE 13:e0202860
Higuchi F, Nagashima H, Ning J et al (2020) Restoration of temozolomide sensitivity by PARP inhibitors in mismatch repair deficient glioblastoma is independent of base excision repair. Clin Cancer Res 26:1690–1699
Nguyen SA, Stechishin ODM, Luchman HA et al (2014) Novel MSH6 mutations in treatment-naïve glioblastoma and anaplastic oligodendroglioma contribute to temozolomide resistance independently of MGMT promoter methylation. Clin Cancer Res 20:4894–4903
Choi S, Yu Y, Grimmer MR et al (2018) Temozolomide-associated hypermutation in gliomas. Neuro Oncology 20:1300–1309
National Library of Medicine, National Centre for Biotechnology Information (2018) Reference SNP(rs) report rs2228006. NIH. https://www.ncbi.nlm.nih.gov/snp/rs2228006. Accessed 8 Oct 2018
National Library of Medicine, National Centre for Biotechnology Information (2018) Reference SNP(rs) report rs1802683. NIH. https://www.ncbi.nlm.nih.gov/snp/rs1802683. Accessed 8 Oct 2018
Touat M, Li YY, Ligon KL (2020) Mechanisms and therapeutic implications of hypermutation in gliomas. Nature 580:517–523
Stritzelberger J, Distel L, Buslei R et al (2018) Acquired temozolomide resistance in human glioblastoma cell line U251 is caused by mismatch repair deficiency and can be overcome by lomustine. Clin Transl Oncol 20:508–516
Herrlinger U, Tzaridis T, Mack F et al (2019) Lomustine-temozolomide combination therapy versus standard temozolomide therapy in patients with newly diagnosed glioblastoma with methylated MGMT promoter (CeTeG/NOA-09): a randomised, open-label, phase 3 trial. Lancet 393:678–688
Blough MD, Westgate MR, Beauchamp D et al (2010) Sensitivity to temozolomide in brain tumor initiating cells. Neuro Oncology 12:756–760
Funding
The Terry Fox Research Institute and Foundation, Alberta Cancer Foundation, Genome Canada, Alberta Innovates Health Solutions, and the family of Clark H. Smith supported this work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11060_2020_3561_MOESM1_ESM.tif
Supplementary file1 (TIF 24678 kb) Figure S1. Western blot analysis of MLH1, MSH6, MSH2, PMS2 and MGMT protein expression in the parental U251N line and 18 TMZ- resistant sub-clones. Compared to parental sample levels, expression of MSH6, MSH2 and PMS2 was decreased in some sub-clones, while MLH1 expression remained similar. MGMT induction was observed in one resistant sub-line. “+” denotes the positive control. “P” denotes the parental U251N sample.
11060_2020_3561_MOESM2_ESM.tif
Supplementary file2 (TIF 24677 kb) Figure S2. Cropped chromatograms showing MMR mutations observed in U251N TMZ-resistant sub-clones as outlined in Table: (A) Colony 15 MLH1 mutation (B) Colony 6 MSH6 mutation #1 (C) Colony 6 MSH6 mutation #2 (D) Colony 6 MSH6 mutation #3 (E) Colony 7 MSH6 mutation (F) Colony 9 MSH6 mutation #1 (G) Colony 9 MSH6 mutation #2 (H) Colony 12 MSH6 mutation (I) Colony 12 MLH1 mutation and (J) Colony 1 MSH2 exon 1 deletion. All missense mutations presented with the wild-type and mutant nucleotide, suggesting allelic heterozygosity or heterogeneity among sampled cells.
11060_2020_3561_MOESM3_ESM.tif
Supplementary file3 (TIF 24677 kb) Figure S3. Uncropped versions of the Western blots Data presented in Figure S1 of (A) MLH1 (B) PMS2 (C) MSH6 (D) MGMT (E) MSH2 and (F) pADPr protein expression in TMZ-resistant U251N sub-clones. Molecular weight ladders could not be pictured on indicated (*) blots.
11060_2020_3561_MOESM4_ESM.tif
Supplementary file4 (TIF 24676 kb) Table S1. Mutational status of MMR repair genes in TMZ resistant colonies. Mutation status of MSH2, MSH6, MLH1 and PMS2 in the parental U251N line and a subset of TMZ-resistant sub-clones introduced above. Most sub-clones harbored one or more mutations in at least one MMR gene. One silent mutation and 2 single-nucleotide polymorphisms were observed in the PMS2 gene of the parental U251N line and all derivative sub-clones. WT = wild type. All amino acids and nucleotides are denoted by standard symbols.
Rights and permissions
About this article
Cite this article
Yuan, A.L., Meode, M., Tan, M. et al. PARP inhibition suppresses the emergence of temozolomide resistance in a model system. J Neurooncol 148, 463–472 (2020). https://doi.org/10.1007/s11060-020-03561-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-020-03561-1