Abstract
Introduction
Machine learning methods have been introduced as a computer aided diagnostic tool, with applications to glioma characterisation on MRI. Such an algorithmic approach may provide a useful adjunct for a rapid and accurate diagnosis of a glioma. The aim of this study is to devise a machine learning algorithm that may be used by radiologists in routine practice to aid diagnosis of both: WHO grade and IDH mutation status in de novo gliomas.
Methods
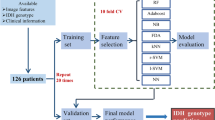
To evaluate the status quo, we interrogated the accuracy of neuroradiology reports in relation to WHO grade: grade II 96.49% (95% confidence intervals [CI] 0.88, 0.99); III 36.51% (95% CI 0.24, 0.50); IV 72.9% (95% CI 0.67, 0.78). We derived five MRI parameters from the same diagnostic brain scans, in under two minutes per case, and then supplied these data to a random forest algorithm.
Results
Machine learning resulted in a high level of accuracy in prediction of tumour grade: grade II/III; area under the receiver operating characteristic curve (AUC) = 98%, sensitivity = 0.82, specificity = 0.94; grade II/IV; AUC = 100%, sensitivity = 1.0, specificity = 1.0; grade III/IV; AUC = 97%, sensitivity = 0.83, specificity = 0.97. Furthermore, machine learning also facilitated the discrimination of IDH status: AUC of 88%, sensitivity = 0.81, specificity = 0.77.
Conclusions
These data demonstrate the ability of machine learning to accurately classify diffuse gliomas by both WHO grade and IDH status from routine MRI alone—without significant image processing, which may facilitate usage as a diagnostic adjunct in clinical practice.


Similar content being viewed by others
References
Ostrom QT, Gittleman H, Fulop J et al (2015) CBTRUS statistical report: primary brain and central nervous system tumors diagnosed in the United States in 2008–2012. Neuro Oncol 17:iv1–iv62. https://doi.org/10.1093/neuonc/nov189
Gittleman H, Kromer C, Ostrom QT et al (2017) Is mortality due to primary malignant brain and other central nervous system tumors decreasing? J Neurooncol 133:265–275. https://doi.org/10.1007/s11060-017-2449-1
Louis DN, Perry A, Reifenberger G et al (2016) The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathol 131:803–820. https://doi.org/10.1007/s00401-016-1545-1
Yan H, Parsons DW, Jin G et al (2009) IDH1 and IDH2 Mutations in Gliomas. N Engl J Med 360:765–773. https://doi.org/10.1056/NEJMoa0808710
Metellus P, Coulibaly B, Colin C et al (2010) Absence of IDH mutation identifies a novel radiologic and molecular subtype of WHO grade II gliomas with dismal prognosis. Acta Neuropathol 120:719–729. https://doi.org/10.1007/s00401-010-0777-8
Thrall JH, Li X, Li Q et al (2018) Artificial intelligence and machine learning in radiology: opportunities, challenges, pitfalls, and criteria for success. J Am Coll Radiol 15:504–508. https://doi.org/10.1016/j.jacr.2017.12.026
Kohli M, Prevedello LM, Filice RW, Geis JR (2017) Implementing machine learning in radiology practice and research. Am J Roentgenol 208:754–760
Zacharaki EI, Wang S, Chawla S, Soo D (2010) Classification of brain tumor type and grade using MRI texture and shape in a machine learning scheme. Magn Reson Med 62:1609–1618. https://doi.org/10.1002/mrm.22147.Classification
Zacharaki EI, Morita N, Bhatt P et al (2012) Survival analysis of patients with high-grade gliomas based on data mining of imaging variables. AJNR Am J Neuroradiol 33:1065–1071. https://doi.org/10.3174/ajnr.A2939
Svolos P, Tsolaki E, Kapsalaki E et al (2013) Investigating brain tumor differentiation with diffusion and perfusion metrics at 3T MRI using pattern recognition techniques. Magn Reson Imaging 31:1567–1577. https://doi.org/10.1016/j.mri.2013.06.010
Wen PY, Reardon DA (2016) Neuro-oncology in 2015: progress in glioma diagnosis, classification and treatment. Nat Rev Neurol 12:2015–2016. https://doi.org/10.1038/nrneurol.2015.242
Kondziolka D, Lunsford LD, Martinez a J (1993) Unreliability of contemporary neurodiagnostic imaging in evaluating suspected adult supratentorial (low-grade) astrocytoma. J Neurosurg 79:533–536. https://doi.org/10.3171/jns.1993.79.4.0533
Aronen HJ, Gazit IE, Louis DN et al (1994) Cerebral blood volume maps of gliomas: comparison with tumor grade and histologic findings. Radiology 191:41–51. https://doi.org/10.1148/radiology.191.1.8134596
Knopp EA, Cha S, Johnson G et al (1999) Glial neoplasms: dynamic contrast-enhanced T2*-weighted MR imaging. Radiology 211:791–798. https://doi.org/10.1148/radiology.211.3.r99jn46791
Guzman-De-Villoria JA, Mateos-Perez JM, Fernandez-Garcia P et al (2014) Added value of advanced over conventional magnetic resonance imaging in grading gliomas and other primary brain tumors. Cancer Imaging 14:1–10. https://doi.org/10.1186/s40644-014-0035-8
Miloushev VZ, Chow DS, Filippi CG (2015) Meta-analysis of diffusion metrics for the prediction of tumor grade in gliomas. AJNR Am J Neuroradiol 36:302–308. https://doi.org/10.3174/ajnr.A4097
Ranjith G, Parvathy R, Vikas V et al (2015) Machine learning methods for the classification of gliomas: initial results using features extracted from MR spectroscopy. Neuroradiol J 28:106–111. https://doi.org/10.1177/1971400915576637
Wiestler B, Kluge A, Lukas M et al (2016) Multiparametric MRI-based differentiation of WHO grade II/III glioma and WHO grade IV glioblastoma. Sci Rep 6:35142. https://doi.org/10.1038/srep35142
Flavahan WA, Drier Y, Liau BB et al (2016) Insulator dysfunction and oncogene activation in IDH mutant gliomas. Nature 529:110–114. https://doi.org/10.1038/nature16490
Sanson M, Marie Y, Paris S et al (2009) Isocitrate dehydrogenase 1 codon 132 mutation is an important prognostic biomarker in gliomas. J Clin Oncol 27:4150–4154. https://doi.org/10.1200/JCO.2009.21.9832
Beiko J, Suki D, Hess KR et al (2014) IDH1 mutant malignant astrocytomas are more amenable to surgical resection and have a survival benefit associated with maximal surgical resection. Neuro Oncol 16:81–91. https://doi.org/10.1093/neuonc/not159
Qi S, Yu L, Li H et al (2014) Isocitrate dehydrogenase mutation is associated with tumor location and magnetic resonance imaging characteristics in astrocytic neoplasms. Oncol Lett 7:1895–1902. https://doi.org/10.3892/ol.2014.2013
Delfanti RL, Piccioni DE, Handwerker J et al (2017) Imaging correlates for the 2016 update on WHO classification of grade II/III gliomas: implications for IDH, 1p/19q and ATRX status. J Neurooncol 135:601–609. https://doi.org/10.1007/s11060-017-2613-7
Branzoli F, Di Stefano AL, Capelle L et al (2017) Highly specific determination of IDH status using edited in vivo magnetic resonance spectroscopy. Neuro Oncol. https://doi.org/10.1093/neuonc/nox214
Zhou H, Vallières M, Bai HX et al (2017) MRI features predict survival and molecular markers in diffuse lower-grade gliomas. Neuro Oncol 19:862–870. https://doi.org/10.1093/neuonc/now256
Zhang B, Chang K, Ramkissoon S et al (2017) Multimodal MRI features predict isocitrate dehydrogenase genotype in high-grade gliomas. Neuro Oncol 19:109–117. https://doi.org/10.1093/neuonc/now121
Elkhaled A, Jalbert LE, Phillips JJ et al (2012) Magnetic resonance of 2-hydroxyglutarate in IDH1-mutated low-grade gliomas. Sci Transl Med 4:116ra5–116ra5. https://doi.org/10.1126/scitranslmed.3002796
Reuss DE, Sahm F, Schrimpf D et al (2015) ATRX and IDH1-R132H immunohistochemistry with subsequent copy number analysis and IDH sequencing as a basis for an “integrated” diagnostic approach for adult astrocytoma, oligodendroglioma and glioblastoma. Acta Neuropathol 129:133–146. https://doi.org/10.1007/s00401-014-1370-3
Arita H, Narita Y, Matsushita Y et al (2015) Development of a robust and sensitive pyrosequencing assay for the detection of IDH1/2 mutations in gliomas. Brain Tumor Pathol 32:22–30. https://doi.org/10.1007/s10014-014-0186-0
Bakas S, Akbari H, Sotiras A et al (2017) Advancing the cancer genome atlas glioma MRI collections with expert segmentation labels and radiomic features. Sci data 4:170117. https://doi.org/10.1038/sdata.2017.117
Clark K, Vendt B, Smith K et al (2013) The Cancer imaging archive (TCIA): maintaining and operating a public information repository. J Digit Imaging 26:1045–1057. https://doi.org/10.1007/s10278-013-9622-7
Smits M, van den Bent MJ (2017) Imaging correlates of adult glioma genotypes. Radiology 284:316–331. https://doi.org/10.1148/radiol.2017151930
Lee EJ, terBrugge K, Mikulis D et al (2011) Diagnostic value of peritumoral minimum apparent diffusion coefficient for differentiation of glioblastoma multiforme from solitary metastatic lesions. Am J Roentgenol 196:71–76. https://doi.org/10.2214/AJR.10.4752
Breiman L (2001) Statistical modeling: the two cultures (with comments and a rejoinder by the author). Stat Sci 16:199–231. https://doi.org/10.1214/ss/1009213726
Zikic D, Glocker B, Konukoglu E et al (2012) Decision forests for tissue-specific segmentation of high-grade gliomas in multi-channel MR. Med Image Comput Comput Assist Interv 15:369–376
Fernández-Delgado M, Cernadas E, Barro S et al (2014) Do we need hundreds of classifiers to solve real world classification problems? J Mach Learn Res 15:3133–3181
Bland JM, Altman DG (1986) Statistical methods for assessing agreement between two methods of clinical measurement. Lancet 1:307–310
Srivastava N, Hinton G, Krizhevsky A et al (2014) Dropout: a simple way to prevent neural networks from overfitting. J Mach Learn Res 15:1929–1958
Chawla NV, Bowyer KW, Hall LO, Kegelmeyer WP (2002) SMOTE: synthetic minority over-sampling technique. J Artif Intell Res 16:321–357. https://doi.org/10.1613/jair.953
Williams LH, Drew T (2017) Distraction in diagnostic radiology: how is search through volumetric medical images affected by interruptions? Cogn Res Princ Implic 2:12. https://doi.org/10.1186/s41235-017-0050-y
Waite S, Kolla S, Jeudy J et al (2017) Tired in the reading room: the influence of fatigue in radiology. J Am Coll Radiol 14:191–197. https://doi.org/10.1016/j.jacr.2016.10.009
Lee CS, Nagy PG, Weaver SJ, Newman-Toker DE (2013) Cognitive and system factors contributing to diagnostic errors in radiology. Am J Roentgenol 201:611–617
Emblem KE, Pinho MC, Zöllner FG et al (2015) A generic support vector machine model for preoperative glioma survival associations. Radiology 275:228–234. https://doi.org/10.1148/radiol.14140770
Kickingereder P, Bonekamp D, Nowosielski M et al (2016) Radiogenomics of Glioblastoma: Machine Learning-based Classification of Molecular Characteristics by Using Multiparametric and Multiregional MR Imaging Features. Radiology 281:907–918. https://doi.org/10.1148/radiol.2016161382
Zacharaki EI, Wang S, Chawla S et al (2009) Classification of brain tumor type and grade using MRI texture and shape in a machine learning scheme. Magn Reson Med 62:1609–1618. https://doi.org/10.1002/mrm.22147
Halevy A, Norvig P, Pereira F (2009) The Unreasonable effectiveness of data. IEEE Intell Syst 24:8–12. https://doi.org/10.1109/MIS.2009.36
Wang Y, Zhang T, Li S et al (2015) Anatomical localization of isocitrate dehydrogenase 1 mutation: a voxel-based radiographic study of 146 low-grade gliomas. Eur J Neurol 22:348–354. https://doi.org/10.1111/ene.12578
Horská A, Barker PB (2010) Imaging of brain tumors: MR spectroscopy and metabolic imaging. Neuroimaging Clin N Am 20:293–310. https://doi.org/10.1016/j.nic.2010.04.003
Jain R, Poisson LM, Gutman D et al (2014) Outcome prediction in patients with glioblastoma by using imaging, clinical, and genomic biomarkers: focus on the nonenhancing component of the tumor. Radiology 272:484–493. https://doi.org/10.1148/radiol.14131691
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
None of the authors of this study have any conflict of interest in relation to this work.
Human and animal participants
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all participants in this study.
Rights and permissions
About this article
Cite this article
De Looze, C., Beausang, A., Cryan, J. et al. Machine learning: a useful radiological adjunct in determination of a newly diagnosed glioma’s grade and IDH status. J Neurooncol 139, 491–499 (2018). https://doi.org/10.1007/s11060-018-2895-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-018-2895-4