Abstract
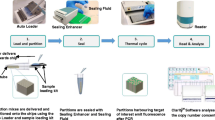
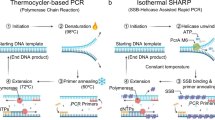
A new readout approach for the Hamiltonian Path Problem (HPP) in DNA computing based on the real-time polymerase chain reaction (PCR) is experimentally implemented and analyzed. Several types of fluorescent probes and detection mechanisms are currently employed in real-time PCR, including SYBR Green, molecular beacons, and hybridization probes. In this study, real-time amplification performed using the TaqMan probes is adopted, as the TaqMan detection mechanism can be exploited for the design and development of the proposed readout approach. Double-stranded DNA molecules of length 140 base-pairs are selected as the input molecules, which represent the solving path for an HPP instance. These input molecules are prepared via the self-assembly of 20-mer and 30-mer single-stranded DNAs, by parallel overlap assembly. The proposed readout approach consists of two steps: real-time amplification in vitro using TaqMan-based real-time PCR, followed by information processing in silico to assess the results of real-time amplification, which in turn, enables extraction of the Hamiltonian path. The performance of the proposed approach is compared with that of conventional graduated PCR. Experimental results establish the superior performance of the proposed approach, relative to graduated PCR, in terms of implementation time.







Similar content being viewed by others
Abbreviations
- bp:
-
Base-pairs
- dsDNA:
-
Double-stranded DNA
- FRET:
-
Fluorescence Resonance Energy Transfer
- HPP:
-
Hamiltonian Path Problem
- PCR:
-
Polymerase Chain Reaction
- ssDNA:
-
Single-stranded DNA
- TSP:
-
Traveling Salesman Problem
References
Adleman L (1994) Molecular computation of solutions to combinatorial problems. Science 266:1021–1024
Lakowicz JR (1999) Principles of fluorescence spectroscopy, 2nd edn. Kluwer Academic/Plenum Publishers, New York
Heid CA et al (1996) Real-time quantitative PCR. Genome Res 6:986–994
Holland PM et al (1991) Detection of specific polymerase chain reaction product by utilizing the 5′ → 3′ exonuclease activity of termus aquaticus DNA polymerase. Proceedings of the National Academy of Sciences of the United States of America 88:7276–7280
Mullis K et al (1986) Specific enzymatic amplification of DNA in vitro: the polymerase chain reaction. Cold Spring Harb Sym Quant Biol 51:263–273
Overbergh L (2003) The use of real-time reverse transcriptase PCR for the quantification of cytokine gene expression. J Biomol Tech 14:33-43
Rose JA et al (1997) The effect of uniform melting temperatures on the efficiency of DNA computing, DIMACS Workshop on DNA Based Computers III, pp 35–42
Walker NJ (2002) A technique whose time has come. Science 296:557–559
Wood DH (1998) A DNA computing algorithm for directed Hamiltonian Paths, Proceedings of the Third Annual Conference on Genetic Programming, pp 731–734
Wood DH et al (1999) Universal biochip readout of directed Hamiltonian Path problems. Lecture Notes in Computer Science 2568, 8th International Workshop on DNA Based Computers, DNA8, Sapporo, Japan, Hagiya, Masami; Ohuchi, Azuma (Eds.): pp 168–181
Acknowledgments
Zuwairie Ibrahim is very grateful to Ritsumeikan Asia Pacific University (APU) for kindly granting a research subsidy to support his expenses during laboratory experimental work at APU. This work was also partially supported by a eScienceFund Research Funding (Vot No. 79033; Z. Ibrahim, N.H. Sarmin, M. Khalid), from the Ministry of Science, Technology, and Innovation (MOSTI), Malaysia, a Grant-in-Aid for Scientific Research B (18300100; J. Rose, A. Suyama), from the Japan Society for the Promotion of Science (JSPS), and JST-CREST (A. Suyama, J. Rose).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ibrahim, Z., Rose, J.A., Suyama, A. et al. Experimental implementation and analysis of a DNA computing readout method based on real-time PCR with TaqMan probes. Nat Comput 7, 277–286 (2008). https://doi.org/10.1007/s11047-007-9047-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11047-007-9047-7