Abstract
Background
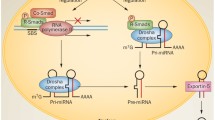
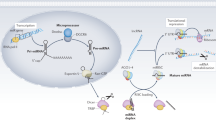
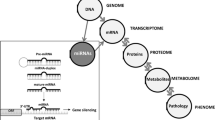
MicroRNAs (miRNAs) are non-coding, endogenous, single-stranded, small (21–25 nucleotides) RNAs. Various target genes at the post-transcriptional stage are modulated by miRNAs that are involved in the regulation of a variety of biological processes such as embryonic development, differentiation, proliferation, apoptosis, inflammation, and metabolic homeostasis. Abnormal miRNA expression is strongly associated with the pathogenesis of multiple common human diseases including cardiovascular diseases, cancer, hepatitis, and metabolic diseases.
Methods and results
Various signaling pathways including transforming growth factor-β, apoptosis, and Wnt signaling pathways have also been characterized to play an essential role in kidney diseases. Most importantly, miRNA-targeted pharmaceutical manipulation has represented a promising new therapeutic approach against kidney diseases. Furthermore, miRNAs such as miR-30e-5p, miR-98-5p, miR-30d-5p, miR-30a-5p, miR-194-5p, and miR-192-5p may be potentially employed as biomarkers for various human kidney diseases.
Conclusions
A significant correlation has also been found between some miRNAs and the clinical markers of renal function like baseline estimated glomerular filtration rate (eGFR). Classification of miRNAs in different genetic renal disorders may promote discoveries in developing innovative therapeutic interventions and treatment tools. Herein, the recent advances in miRNAs associated with renal pathogenesis, emphasizing genetic kidney diseases and development, have been summarized.




Similar content being viewed by others
Abbreviations
- DGCR8:
-
DiGeorge syndrome critical region 8
- AGO2:
-
Argonaute 2
- TRBP:
-
TAR RNA-binding protein 2
- C-P4H:
-
Collagen prolyl 4-hydroxylase
- pri-miRNA:
-
Primary miRNA
- pre-miRNA:
-
Precursor miRNA
- ERK1:
-
Extracellular signal-regulated kinase 1
- MAPKAPK2:
-
MAPK-activated protein kinase 2
- GSK3β:
-
Glycogen synthase kinase 3β
- Ud:
-
Uridylation
- CHD1L :
-
Chromodomain helicase/ATPase DNA binding protein 1-like gene
- CAKUT:
-
Congenital anomalies of the kidney and urinary tract
- Dgcr8:
-
DiGeorge syndrome critical region 8
- PKD:
-
Polycystic kidney disease
- eGFR:
-
Estimated glomerular filtration rate
- SRF:
-
Serum response factor
- SMAD:
-
Mothers against decapentaplegic homolog
- SOCS:
-
Suppressor of cytokine signaling
- CDKN1A:
-
Cyclin-dependent kinase inhibitor 1
- PAKs:
-
P21-activated kinases
- SOD1:
-
Superoxide dismutase type 1
- APOE:
-
Apolipoprotein E
- PPARα:
-
Peroxisome proliferator-activated receptor alpha
- TLR2:
-
Toll-like receptor 2
- KLF12:
-
Kruppel Like Factor 12
- HMGA2:
-
High Mobility Group AT-Hook 2
- CCN2:
-
Cellular Communication Network Factor 2
- PTEN:
-
Phosphatase and tensin homolog deleted on chromosome 10
- ACTA2:
-
Actin Alpha 2
- COL4:
-
Collagen IV
References
Lakhia R, Yheskel M, Flaten A, Ramalingam H, Aboudehen K, Ferrè S, Biggers L, Mishra A, Chaney C, Wallace DP, Carroll T, Igarashi P, Patel V (2020) Interstitial microRNA miR-214 attenuates inflammation and polycystic kidney disease progression. JCI Insight. https://doi.org/10.1172/jci.insight.133785
Wang Z, Bao W, Zou X, Tan P, Chen H, Lai C, Liu D, Luo Z, Huang M (2019) Co-expression analysis reveals dysregulated miRNAs and miRNA–mRNA interactions in the development of contrast-induced acute kidney injury. PLoS ONE 14(7):e0218574. https://doi.org/10.1371/journal.pone.0218574
Sakurai F, Hashimoto R, Inoue C, Wakabayashi K, Tsukamoto T, Imaizumi T, Andres TGM, Sakai E, Itsuki K, Sakamoto N, Wakita T, Mizuguchi H (2019) miR-27b-mediated suppression of aquaporin-11 expression in hepatocytes reduces HCV genomic RNA levels but not viral titers. Virol J 16(1):58. https://doi.org/10.1186/s12985-019-1160-6
Ciceri S, Montalvão-de-Azevedo R, Tajbakhsh A, Bertolotti A, Spagnuolo RD, Boschetti L, Capasso M, D’Angelo P, Serra A, Diomedi-Camassei F, Meli M, Nantron M, Quarello P, Buccoliero AM, Tamburini A, Ciniselli CM, Verderio P, Collini P, Radice P, Spreafico F, Perotti D (2020) Analysis of the mutational status of SIX1/2 and microRNA processing genes in paired primary and relapsed Wilms tumors and association with relapse. Cancer Gene Ther. https://doi.org/10.1038/s41417-020-00268-3
Seelan RS, Pisano MM, Greene RM (2022) MicroRNAs as biomarkers for birth defects. MicroRNA. https://doi.org/10.2174/2211536611666220215123423
Lee Y, Ahn C, Han J, Choi H, Kim J, Yim J, Lee J, Provost P, Rådmark O, Kim S, Kim VN (2003) The nuclear RNase III Drosha initiates microRNA processing. Nature 425(6956):415–419. https://doi.org/10.1038/nature01957
Zhiyanov A, Nersisyan S, Tonevitsky A (2021) Hairpin sequence and structure is associated with features of isomiR biogenesis. RNA Biol 18(sup1):430–438. https://doi.org/10.1080/15476286.2021.1952759
Tabatabaeian H, Lim SK, Chu T, Seah SH, Lim YP (2021) WBP2 inhibits microRNA biogenesis via interaction with the microprocessor complex. Life Sci Alliance. https://doi.org/10.26508/lsa.202101038
Macgregor-Das AM, Das S (2018) A microRNA’s journey to the center of the mitochondria. Am J Physiol Heart Circ Physiol 315(2):H206-h215. https://doi.org/10.1152/ajpheart.00714.2017
Li Z, Rana TM (2014) Therapeutic targeting of microRNAs: current status and future challenges. Nat Rev Drug Discov 13(8):622–638. https://doi.org/10.1038/nrd4359
Lund E, Güttinger S, Calado A, Dahlberg JE, Kutay U (2004) Nuclear export of microRNA precursors. Science (New York NY) 303(5654):95–98. https://doi.org/10.1126/science.1090599
Grishok A, Pasquinelli AE, Conte D, Li N, Parrish S, Ha I, Baillie DL, Fire A, Ruvkun G, Mello CC (2001) Genes and mechanisms related to RNA interference regulate expression of the small temporal RNAs that control C. elegans developmental timing. Cell 106(1):23–34. https://doi.org/10.1016/s0092-8674(01)00431-7
Tang X, Li M, Tucker L, Ramratnam B (2011) Glycogen synthase kinase 3 beta (GSK3β) phosphorylates the RNAase III enzyme Drosha at S300 and S302. PLoS ONE 6(6):e20391. https://doi.org/10.1371/journal.pone.0020391
Paroo Z, Ye X, Chen S, Liu Q (2009) Phosphorylation of the human microRNA-generating complex mediates MAPK/Erk signaling. Cell 139(1):112–122. https://doi.org/10.1016/j.cell.2009.06.044
McMahon RS, Penfold D, Bashir K (2021) Anatomy, abdomen and pelvis, kidney collecting ducts. StatPearls Publishing, Treasure Island
Davies JA, Bard JB (1998) The development of the kidney. Curr Top Dev Biol 39:245–301. https://doi.org/10.1016/s0070-2153(08)60458-5
Sequeira Lopez ML, Gomez RA (2004) The role of angiotensin II in kidney embryogenesis and kidney abnormalities. Curr Opin Nephrol Hypertens 13(1):117–122. https://doi.org/10.1097/00041552-200401000-00016
Wang Y, Lumbers ER, Arthurs AL, Corbisier de Meaultsart C, Mathe A, Avery-Kiejda KA, Roberts CT, Pipkin FB, Marques FZ, Morris BJ, Pringle KG (2018) Regulation of the human placental (pro)renin receptor-prorenin–angiotensin system by microRNAs. Mol Hum Reprod 24(9):453–464. https://doi.org/10.1093/molehr/gay031
Laganà AS, Giordano D, Loddo S, Zoccali G, Vitale SG, Santamaria A, Buemi M, D’Anna R (2017) Decreased Endothelial Progenitor Cells (EPCs) and increased Natural Killer (NK) cells in peripheral blood as possible early markers of preeclampsia: a case–control analysis. Arch Gynecol Obstet 295(4):867–872. https://doi.org/10.1007/s00404-017-4296-x
Tsochandaridis M, Nasca L, Toga C, Levy-Mozziconacci A (2015) Circulating microRNAs as clinical biomarkers in the predictions of pregnancy complications. Biomed Res Int 2015:294954. https://doi.org/10.1155/2015/294954
Chevalier RL (2018) Evolution and kidney development: a Rosetta stone for nephrology. J Am Soc Nephrol 29(3):706–709. https://doi.org/10.1681/ASN.2018010013
Kömhoff M, Wang JL, Cheng HF, Langenbach R, McKanna JA, Harris RC, Breyer MD (2000) Cyclooxygenase-2-selective inhibitors impair glomerulogenesis and renal cortical development. Kidney Int 57(2):414–422. https://doi.org/10.1046/j.1523-1755.2000.00861.x
Klein T, Klaus G, Kömhoff M (2015) Prostacyclin synthase: upregulation during renal development and in glomerular disease as well as its constitutive expression in cultured human mesangial cells. Mediat Inflamm 2015:654151. https://doi.org/10.1155/2015/654151
Wu YJ, Liu Y, Hu YQ, Wang L, Bai FR, Xu C, Wu JW (2021) Control of multiciliogenesis by miR-34/449 in the male reproductive tract through enforcing cell cycle exit. J Cell Sci. https://doi.org/10.1242/jcs.253450
Marra AN, Adeeb BD, Chambers BE, Drummond BE, Ulrich M, Addiego A, Springer M, Poureetezadi SJ, Chambers JM, Ronshaugen M, Wingert RA (2019) Prostaglandin signaling regulates renal multiciliated cell specification and maturation. Proc Natl Acad Sci USA 116(17):8409–8418. https://doi.org/10.1073/pnas.1813492116
Song M-S, Rossi JJ (2017) Molecular mechanisms of Dicer: endonuclease and enzymatic activity. Biochem J 474(10):1603–1618. https://doi.org/10.1042/BCJ20160759
Chu JY, Sims-Lucas S, Bushnell DS, Bodnar AJ, Kreidberg JA, Ho J (2014) Dicer function is required in the metanephric mesenchyme for early kidney development. Am J Physiol Ren Physiol 306(7):F764–F772. https://doi.org/10.1152/ajprenal.00426.2013
Nagalakshmi VK, Ren Q, Pugh MM, Valerius MT, McMahon AP, Yu J (2011) Dicer regulates the development of nephrogenic and ureteric compartments in the mammalian kidney. Kidney Int 79(3):317–330. https://doi.org/10.1038/ki.2010.385
Puri P, Bushnell D, Schaefer CM, Bates CM (2016) Six2creFrs2α knockout mice are a novel model of renal cystogenesis. Sci Rep 6:36736. https://doi.org/10.1038/srep36736
Yadav RP, Mäkelä JA, Hyssälä H, Cisneros-Montalvo S, Kotaja N (2020) DICER regulates the expression of major satellite repeat transcripts and meiotic chromosome segregation during spermatogenesis. Nucleic Acids Res 48(13):7135–7153. https://doi.org/10.1093/nar/gkaa460
Bartram MP, Dafinger C, Habbig S, Benzing T, Schermer B, Müller RU (2015) Loss of Dgcr8-mediated microRNA expression in the kidney results in hydronephrosis and renal malformation. BMC Nephrol 16:55. https://doi.org/10.1186/s12882-015-0053-1
Xie M, Steitz JA (2014) Versatile microRNA biogenesis in animals and their viruses. RNA Biol 11(6):673–681. https://doi.org/10.4161/rna.28985
Cerqueira DM, Bodnar AJ, Phua YL, Freer R, Hemker SL, Walensky LD, Hukriede NA, Ho J (2017) Bim gene dosage is critical in modulating nephron progenitor survival in the absence of microRNAs during kidney development. FASEB J Off Publ Fed Am Soc Exp Biol 31(8):3540–3554. https://doi.org/10.1096/fj.201700010R
Fluitt MB, Shivapurkar N, Kumari M, Singh S, Li L, Tiwari S, Ecelbarger CM (2020) Systemic inhibition of miR-451 increases fibrotic signaling and diminishes autophagic response to exacerbate renal damage in Tallyho/Jng mice. Am J Physiol Ren Physiol 319(3):F476–F486. https://doi.org/10.1152/ajprenal.00594.2019
Beizavi Z, Gheibihayat SM, Moghadasian H, Zare H, Yeganeh BS, Askari H, Vakili S, Tajbakhsh A, Savardashtaki A (2021) The regulation of CD47-SIRPα signaling axis by microRNAs in combination with conventional cytotoxic drugs together with the help of nano-delivery: a choice for therapy? Mol Biol Rep 48(7):5707–5722. https://doi.org/10.1007/s11033-021-06547-y
Tan Z, Rak-Raszewska A, Skovorodkin I, Vainio SJ (2020) Mouse embryonic stem cell-derived ureteric bud progenitors induce nephrogenesis. Cells. https://doi.org/10.3390/cells9020329
Davidson AJ (2008) Mouse kidney development. In: StemBook. Harvard Stem Cell Institute, Cambridge
Cwiek A, Suzuki M, DeRonde K, Conaway M, Bennett KM, El Dahr S, Reidy KJ, Charlton JR (2021) Premature differentiation of nephron progenitor cell and dysregulation of gene pathways critical to kidney development in a model of preterm birth. Sci Rep 11(1):21667. https://doi.org/10.1038/s41598-021-00489-y
Ezuruike U, Blenkinsop A, Pansari A, Abduljalil K (2022) Quantification of fetal renal function using fetal urine production rate and its reflection on the amniotic and fetal creatinine levels during pregnancy. Front Pediatr 10:841495. https://doi.org/10.3389/fped.2022.841495
Hinchliffe SA, Sargent PH, Howard CV, Chan YF, van Velzen D (1991) Human intrauterine renal growth expressed in absolute number of glomeruli assessed by the dissector method and Cavalieri principle. Lab Investig 64(6):777–784
Awazu M (2022) Structural and functional changes in the kidney caused by adverse fetal and neonatal environments. Mol Biol Rep 49(3):2335–2344. https://doi.org/10.1007/s11033-021-06967-w
Nam Y, Chen C, Gregory RI, Chou JJ, Sliz P (2011) Molecular basis for interaction of let-7 microRNAs with Lin28. Cell 147(5):1080–1091. https://doi.org/10.1016/j.cell.2011.10.020
Clugston A, Bodnar A, Cerqueira DM, Phua YL, Lawler A, Boggs K, Pfenning AR, Ho J, Kostka D (2022) Chromatin accessibility and microRNA expression in nephron progenitor cells during kidney development. Genomics 114(1):278–291. https://doi.org/10.1016/j.ygeno.2021.12.017
Yokoyama S, Hashimoto M, Shimizu H, Ueno-Kudoh H, Uchibe K, Kimura I, Asahara H (2008) Dynamic gene expression of Lin-28 during embryonic development in mouse and chicken. Gene Expr Patterns 8(3):155–160. https://doi.org/10.1016/j.gep.2007.11.001
Wilbert ML, Huelga SC, Kapeli K, Stark TJ, Liang TY, Chen SX, Yan BY, Nathanson JL, Hutt KR, Lovci MT, Kazan H, Vu AQ, Massirer KB, Morris Q, Hoon S, Yeo GW (2012) LIN28 binds messenger RNAs at GGAGA motifs and regulates splicing factor abundance. Mol Cell 48(2):195–206. https://doi.org/10.1016/j.molcel.2012.08.004
Yermalovich AV, Osborne JK, Sousa P, Han A, Kinney MA, Chen MJ, Robinton DA, Montie H, Pearson DS, Wilson SB, Combes AN, Little MH, Daley GQ (2019) Lin28 and let-7 regulate the timing of cessation of murine nephrogenesis. Nat Commun 10(1):168–168. https://doi.org/10.1038/s41467-018-08127-4
Urbach A, Yermalovich A, Zhang J, Spina CS, Zhu H, Perez-Atayde AR, Shukrun R, Charlton J, Sebire N, Mifsud W, Dekel B, Pritchard-Jones K, Daley GQ (2014) Lin28 sustains early renal progenitors and induces Wilms tumor. Genes Dev 28(9):971–982. https://doi.org/10.1101/gad.237149.113
Nguyen LH, Robinton DA, Seligson MT, Wu L, Li L, Rakheja D, Comerford SA, Ramezani S, Sun X, Parikh MS, Yang EH, Powers JT, Shinoda G, Shah SP, Hammer RE, Daley GQ, Zhu H (2014) Lin28b is sufficient to drive liver cancer and necessary for its maintenance in murine models. Cancer Cell 26(2):248–261. https://doi.org/10.1016/j.ccr.2014.06.018
Hunter RW, Liu Y, Manjunath H, Acharya A, Jones BT, Zhang H, Chen B, Ramalingam H, Hammer RE, Xie Y, Richardson JA, Rakheja D, Carroll TJ, Mendell JT (2018) Loss of Dis3l2 partially phenocopies Perlman syndrome in mice and results in up-regulation of Igf2 in nephron progenitor cells. Genes Dev 32(13–14):903–908. https://doi.org/10.1101/gad.315804.118
Nozu K, Nakanishi K, Abe Y, Udagawa T, Okada S, Okamoto T, Kaito H, Kanemoto K, Kobayashi A, Tanaka E, Tanaka K, Hama T, Fujimaru R, Miwa S, Yamamura T, Yamamura N, Horinouchi T, Minamikawa S, Nagata M, Iijima K (2019) A review of clinical characteristics and genetic backgrounds in Alport syndrome. Clin Exp Nephrol 23(2):158–168. https://doi.org/10.1007/s10157-018-1629-4
Clark SD, Song W, Cianciolo R, Lees G, Nabity M, Liu S (2019) Abnormal expression of miR-21 in kidney tissue of dogs with X-linked hereditary nephropathy: a canine model of chronic kidney disease. Vet Pathol 56(1):93–105. https://doi.org/10.1177/0300985818806050
Chen W, Tang D, Dai Y, Diao H (2019) Establishment of microRNA, transcript and protein regulatory networks in Alport syndrome induced pluripotent stem cells. Mol Biol Rep 19(1):238–250. https://doi.org/10.3892/mmr.2018.9672
Lautrette A, Li S, Alili R, Sunnarborg SW, Burtin M, Lee DC, Friedlander G, Terzi F (2005) Angiotensin II and EGF receptor cross-talk in chronic kidney diseases: a new therapeutic approach. Nat Med 11(8):867–874. https://doi.org/10.1038/nm1275
Dawson NA, Guo C, Zak R, Dorsey B, Smoot J, Wong J, Hussain A (2004) A phase II trial of gefitinib (Iressa, ZD1839) in stage IV and recurrent renal cell carcinoma. Clin Cancer Res 10(23):7812–7819. https://doi.org/10.1158/1078-0432.ccr-04-0310
Wang IK, Palanisamy K, Sun KT, Yu SH, Yu TM, Li CH, Lin FY, Chou AK, Wang GJ, Chen KB, Li CY (2020) The functional interplay of lncRNA EGOT and HuR regulates hypoxia-induced autophagy in renal tubular cells. J Cell Biochem. https://doi.org/10.1002/jcb.29669
Shuai C, Guo W, Wu P, Yang W, Hu S, Xia Y, Feng P (2018) A graphene oxide-Ag co-dispersing nanosystem: dual synergistic effects on antibacterial activities and mechanical properties of polymer scaffolds. Chem Eng J 347:322–333. https://doi.org/10.1016/j.cej.2018.04.092
Guo J, Song W, Boulanger J, Xu EY, Wang F, Zhang Y, He Q, Wang S, Yang L, Pryce C, Phillips L, MacKenna D, Leberer E, Ibraghimov-Beskrovnaya O, Ding J, Liu S (2019) Dysregulated expression of microRNA-21 and disease-related genes in human patients and in a mouse model of Alport syndrome. Hum Gene Ther 30(7):865–881. https://doi.org/10.1089/hum.2018.205
Mezei G, Chang ET, Mowat FS, Moolgavkar SH (2021) Comments on a recent case–control study of malignant mesothelioma of the pericardium and the tunica vaginalis testis. Scand J Work Environ Health 47(1):85–86. https://doi.org/10.5271/sjweh.3909
Lopes RI, Lorenzo A (2017) Recent advances in the management of Wilms’ tumor. F1000Res 6:670. https://doi.org/10.12688/f1000research.10760.1
Ramezani A, Devaney JM, Cohen S, Wing MR, Scott R, Knoblach S, Singhal R, Howard L, Kopp JB, Raj DS (2015) Circulating and urinary microRNA profile in focal segmental glomerulosclerosis: a pilot study. Eur J Clin Investig 45(4):394–404. https://doi.org/10.1111/eci.12420
Szeto CC, Li PK (2014) MicroRNAs in IgA nephropathy. Nat Rev Nephrol 10(5):249–256. https://doi.org/10.1038/nrneph.2014.50
Chau BN, Xin C, Hartner J, Ren S, Castano AP, Linn G, Li J, Tran PT, Kaimal V, Huang X, Chang AN, Li S, Kalra A, Grafals M, Portilla D, MacKenna DA, Orkin SH, Duffield JS (2012) MicroRNA-21 promotes fibrosis of the kidney by silencing metabolic pathways. Sci Transl Med 4(121):121ra18. https://doi.org/10.1126/scitranslmed.3003205
Patel V, Williams D, Hajarnis S, Hunter R, Pontoglio M, Somlo S, Igarashi P (2013) miR-17~92 miRNA cluster promotes kidney cyst growth in polycystic kidney disease. Proc Natl Acad Sci USA 110(26):10765–10770. https://doi.org/10.1073/pnas.1301693110
Gomez IG, MacKenna DA, Johnson BG, Kaimal V, Roach AM, Ren S, Nakagawa N, Xin C, Newitt R, Pandya S, Xia TH, Liu X, Borza DB, Grafals M, Shankland SJ, Himmelfarb J, Portilla D, Liu S, Chau BN, Duffield JS (2015) Anti-microRNA-21 oligonucleotides prevent Alport nephropathy progression by stimulating metabolic pathways. J Clin Investig 125(1):141–156. https://doi.org/10.1172/jci75852
Patel V, Hajarnis S, Williams D, Hunter R, Huynh D, Igarashi P (2012) MicroRNAs regulate renal tubule maturation through modulation of Pkd1. J Am Soc Nephrol 23(12):1941–1948. https://doi.org/10.1681/asn.2012030321
Sun L, Zhu J, Wu M, Sun H, Zhou C, Fu L, Xu C, Mei C (2015) Inhibition of miR-199a-5p reduced cell proliferation in autosomal dominant polycystic kidney disease through targeting CDKN1C. Med Sci Monit 21:195–200. https://doi.org/10.12659/msm.892141
Shin Y, Kim DY, Ko JY, Woo YM, Park JH (2018) Regulation of KLF12 by microRNA-20b and microRNA-106a in cystogenesis. FASEB J 32(7):3574–3582. https://doi.org/10.1096/fj.201700923R
Sun L, Hu C, Wang Z, Zhang X (2020) miR-182 inhibits kidney fibrosis by regulating transforming growth factor β1/Smad3 pathway in autosomal dominant polycystic kidney disease. IUBMB Life 72(7):1340–1348. https://doi.org/10.1002/iub.2255
Usui T, Morito N, Shawki HH, Sato Y, Tsukaguchi H, Hamada M, Jeon H, Yadav MK, Kuno A, Tsunakawa Y, Okada R, Ojima T, Kanai M, Asano K, Imamura Y, Koshida R, Yoh K, Usui J, Yokoi H, Kasahara M, Yoshimura A, Muratani M, Kudo T, Oishi H, Yamagata K, Takahashi S (2020) Transcription factor MafB in podocytes protects against the development of focal segmental glomerulosclerosis. Kidney Int 98(2):391–403. https://doi.org/10.1016/j.kint.2020.02.038
Rosenberg AZ, Kopp JB (2017) Focal segmental glomerulosclerosis. Clin J Am Soc Nephrol 12(3):502–517. https://doi.org/10.2215/cjn.05960616
Leslie NR, Downes CP (2002) PTEN: The down side of PI3-kinase signalling. Cell Signal 14(4):285–295. https://doi.org/10.1016/s0898-6568(01)00234-0
Bennett MR, Czech KA, Arend LJ, Witte DP, Devarajan P, Potter SS (2007) Laser capture microdissection-microarray analysis of focal segmental glomerulosclerosis glomeruli. Nephron Exp Nephrol 107(1):e30-40. https://doi.org/10.1159/000106775
Marrone AK, Stolz DB, Bastacky SI, Kostka D, Bodnar AJ, Ho J (2014) MicroRNA-17~92 is required for nephrogenesis and renal function. J Am Soc Nephrol 25(7):1440–1452. https://doi.org/10.1681/asn.2013040390
Bian LH, Duan JL, Zhou C, Shen GW, Wang XY, Yang Y, Zhang XL, Xiao SJ (2020) MicroRNA-19b inhibitors can attenuate the STAT3 signaling pathway in NPC C666–1 cells. Mol Med Rep 22(1):51–56. https://doi.org/10.3892/mmr.2020.11112
Shuai C, Liu G, Yang Y, Qi F, Peng S, Yang W, He C, Wang G, Qian G (2020) A strawberry-like Ag-decorated barium titanate enhances piezoelectric and antibacterial activities of polymer scaffold. Nano Energy 74:104825. https://doi.org/10.1016/j.nanoen.2020.104825
Huang Z, Zhang Y, Zhou J, Zhang Y (2017) Urinary exosomal miR-193a can be a potential biomarker for the diagnosis of primary focal segmental glomerulosclerosis in children. Biomed Res Int 2017:7298160. https://doi.org/10.1155/2017/7298160
Capuano I, Buonanno P, Riccio E, Amicone M, Pisani A (2022) Therapeutic advances in ADPKD: the future awaits. J Nephrol 35(2):397–415. https://doi.org/10.1007/s40620-021-01062-6
Su W, Cao R, Zhang XY, Guan Y (2020) Aquaporins in the kidney: physiology and pathophysiology. Am J Physiol Ren Physiol 318(1):F193–F203. https://doi.org/10.1152/ajprenal.00304.2019
Lee EC, Valencia T, Allerson C, Schairer A, Flaten A, Yheskel M, Kersjes K, Li J, Gatto S, Takhar M, Lockton S, Pavlicek A, Kim M, Chu T, Soriano R, Davis S, Androsavich JR, Sarwary S, Owen T, Kaplan J, Liu K, Jang G, Neben S, Bentley P, Wright T, Patel V (2019) Discovery and preclinical evaluation of anti-miR-17 oligonucleotide RGLS4326 for the treatment of polycystic kidney disease. Nat Commun 10(1):4148. https://doi.org/10.1038/s41467-019-11918-y
Ramalingam H, Yheskel M, Patel V (2020) Modulation of polycystic kidney disease by non-coding RNAs. Cell Signal 71:109548. https://doi.org/10.1016/j.cellsig.2020.109548
Yheskel M, Patel V (2017) Therapeutic microRNAs in polycystic kidney disease. Curr Opin Nephrol Hypertens 26(4):282–289. https://doi.org/10.1097/mnh.0000000000000333
Feng P, Wu P, Gao C, Yang Y, Guo W, Yang W, Shuai C (2018) A multimaterial scaffold with tunable properties: toward bone tissue repair. Adv Sci (Weinh) 5(6):1700817. https://doi.org/10.1002/advs.201700817
Kocyigit I, Taheri S, Sener EF, Eroglu E, Ozturk F, Unal A, Korkmaz K, Zararsiz G, Sipahioglu MH, Ozkul Y, Tokgoz B, Oymak O, Ecder T, Axelsson J (2017) Serum micro-RNA profiles in patients with autosomal dominant polycystic kidney disease according to hypertension and renal function. BMC Nephrol 18(1):179–179. https://doi.org/10.1186/s12882-017-0600-z
Ben-Dov IZ, Tan Y-C, Morozov P, Wilson PD, Rennert H, Blumenfeld JD, Tuschl T (2014) Urine microRNA as potential biomarkers of autosomal dominant polycystic kidney disease progression: description of miRNA profiles at baseline. PLoS ONE 9(1):e86856. https://doi.org/10.1371/journal.pone.0086856
Pawluczyk IZA, Didangelos A, Barbour SJ, Er L, Becker JU, Martin R, Taylor S, Bhachu JS, Lyons EG, Jenkins RH, Fraser D, Molyneux K, Perales-Patón J, Saez-Rodriguez J, Barratt J (2021) Differential expression of microRNA miR-150-5p in IgA nephropathy as a potential mediator and marker of disease progression. Kidney Int 99(5):1127–1139. https://doi.org/10.1016/j.kint.2020.12.028
Szeto C-C, Ching-Ha KB, Ka-Bik L, Mac-Moune LF, Cheung-Lung CP, Gang W, Kai-Ming C, Kam-Tao LP (2012) Micro-RNA expression in the urinary sediment of patients with chronic kidney diseases. Dis Markers 33(3):137–144. https://doi.org/10.1155/2012/842764
Wang G, Kwan BC, Lai FM, Chow KM, Li PK, Szeto CC (2012) Urinary miR-21, miR-29, and miR-93: novel biomarkers of fibrosis. Am J Nephrol 36(5):412–418. https://doi.org/10.1159/000343452
Fan Q, Lu R, Zhu M, Yan Y, Guo X, Qian Y, Zhang L, Dai H, Ni Z, Gu L (2019) Serum miR-192 is related to tubulointerstitial lesion and short-term disease progression in iga nephropathy. Nephron 142(3):195–207. https://doi.org/10.1159/000497488
Ichii O, Otsuka-Kanazawa S, Horino T, Kimura J, Nakamura T, Matsumoto M, Toi M, Kon Y (2014) Decreased miR-26a expression correlates with the progression of podocyte injury in autoimmune glomerulonephritis. PLoS ONE 9(10):e110383. https://doi.org/10.1371/journal.pone.0110383
Su B, Han H, Ji C, Hu W, Yao J, Yang J, Fan Y, Li J (2020) miR-21 promotes calcium oxalate-induced renal tubular cell injury by targeting PPARA. Am J Physiol Ren Physiol 319(2):F202–F214. https://doi.org/10.1152/ajprenal.00132.2020
Perez-Hernandez J, Forner MJ, Pinto C, Chaves FJ, Cortes R, Redon J (2015) Increased urinary exosomal microRNAs in patients with systemic lupus erythematosus. PLoS ONE 10(9):e0138618. https://doi.org/10.1371/journal.pone.0138618
Zhang L, Li LX, Zhou JX, Harris PC, Calvet JP, Li X (2020) RNA helicase p68 inhibits the transcription and post-transcription of Pkd1 in ADPKD. Theranostics 10(18):8281–8297. https://doi.org/10.7150/thno.47315
Gurtner A, Falcone E, Garibaldi F, Piaggio G (2016) Dysregulation of microRNA biogenesis in cancer: the impact of mutant p53 on Drosha complex activity. J Exp Clin Cancer Res 35:45. https://doi.org/10.1186/s13046-016-0319-x
Hong S, Noh H, Chen H, Padia R, Pan ZK, Su SB, Jing Q, Ding HF, Huang S (2013) Signaling by p38 MAPK stimulates nuclear localization of the microprocessor component p68 for processing of selected primary microRNAs. Sci Signal 6(266):ra16. https://doi.org/10.1126/scisignal.2003706
Salzman DW, Shubert-Coleman J, Furneaux H (2007) P68 RNA helicase unwinds the human let-7 microRNA precursor duplex and is required for let-7-directed silencing of gene expression. J Biol Chem 282(45):32773–32779. https://doi.org/10.1074/jbc.M705054200
Li H, Lai P, Jia J, Song Y, Xia Q, Huang K, He N, Ping W, Chen J, Yang Z, Li J, Yao M, Dong X, Zhao J, Hou C, Esteban MA, Gao S, Pei D, Hutchins AP, Yao H (2017) RNA helicase DDX5 inhibits reprogramming to pluripotency by miRNA-based repression of RYBP and its PRC1-dependent and -independent functions. Cell Stem Cell 20(4):571. https://doi.org/10.1016/j.stem.2017.03.014
Taira N, Yamaguchi T, Kimura J, Lu ZG, Fukuda S, Higashiyama S, Ono M, Yoshida K (2014) Induction of amphiregulin by p53 promotes apoptosis via control of microRNA biogenesis in response to DNA damage. Proc Natl Acad Sci USA 111(2):717–722. https://doi.org/10.1073/pnas.1313675111
Tee JB, Dnyanmote AV, Lorenzo MK, Lee OR, Grisaru S, Suk M, Acott PD (2020) Pkd1-targeted mutation reveals a role for the Wolffian duct in autosomal dominant polycystic kidney disease. J Dev Orig Health Dis 11(1):78–85. https://doi.org/10.1017/s2040174419000436
Magayr TA, Song X, Streets AJ, Vergoz L, Chang L, Valluru MK, Yap HL, Lannoy M, Haghighi A, Simms RJ, Tam FWK, Pei Y, Ong ACM (2020) Global microRNA profiling in human urinary exosomes reveals novel disease biomarkers and cellular pathways for autosomal dominant polycystic kidney disease. Kidney Int 98(2):420–435. https://doi.org/10.1016/j.kint.2020.02.008
Liu D, Xia M, Liu Y, Tan X, He L, Liu Y, Chen G, Liu H (2020) The upregulation of miR-98-5p affects the glycosylation of IgA1 through cytokines in IgA nephropathy. Int Immunopharmacol 82:106362. https://doi.org/10.1016/j.intimp.2020.106362
Serino G, Sallustio F, Cox SN, Pesce F, Schena FP (2012) Abnormal miR-148b expression promotes aberrant glycosylation of IgA1 in IgA nephropathy. J Am Soc Nephrol 23(5):814–824. https://doi.org/10.1681/asn.2011060567
Serino G, Sallustio F, Curci C, Cox SN, Pesce F, De Palma G, Schena FP (2015) Role of let-7b in the regulation of N-acetylgalactosaminyltransferase 2 in IgA nephropathy. Nephrol Dial Transplant 30(7):1132–1139. https://doi.org/10.1093/ndt/gfv032
Li C, Shi J, Zhao Y (2018) miR-320 promotes B cell proliferation and the production of aberrant glycosylated IgA1 in IgA nephropathy. J Cell Biochem 119(6):4607–4614. https://doi.org/10.1002/jcb.26628
Hu S, Bao H, Xu X, Zhou X, Qin W, Zeng C, Liu Z (2015) Increased miR-374b promotes cell proliferation and the production of aberrant glycosylated IgA1 in B cells of IgA nephropathy. FEBS Lett 589(24 Pt B):4019–4025. https://doi.org/10.1016/j.febslet.2015.10.033
Acknowledgements
This research received no specific grants from any funding agency in the public, commercial, or not-for-profit sectors. Hereby, the authors would like to thank Ms. A. Keivanshekouh at the Research Consultation Center (RCC) of Shiraz University of Medical Sciences for her invaluable assistance in editing the manuscript.
Funding
The authors declare that no funds, grants, or other support have been received during the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
HA, AT, SZK, ER-A, AS, PRA, LR, AA, and MFA drafted the manuscript. HA and NS presented the idea. SV and NS performed the literature search and prepared the figures. MFA, HA, and ARS edited the manuscript. HA supervised the whole study. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Ethical approval
Not applicable.
Informed consent
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Askari, H., Raeis-Abdollahi, E., Abazari, M.F. et al. Recent findings on the role of microRNAs in genetic kidney diseases. Mol Biol Rep 49, 7039–7056 (2022). https://doi.org/10.1007/s11033-022-07620-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-022-07620-w