Abstract
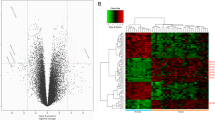
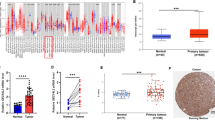
PPFIA family members and ALG3 play important roles in tumorigenesis and tumor progression. However, the exact roles of distinct PPFIA family members and ALG3 in head and neck squamous cell carcinoma (HNSCC) remain unclear. We studied the mRNA expressions of PPFIA family members and ALG3 in a variety of tumor types compared with the normal controls using the Oncomine database along with meta-analyses of their expressions in HNSCC cancer cell line. The mRNA expressions of PPFIA family members and ALG3 in laryngeal squamous cell carcinoma cell line and normal laryngeal cell line were detected by quantitative real-time polymerase chain reaction. Based on the cBioportal database, we further studied mRNA expression alterations and co-occurrence relationships of the PPFIA family members and ALG3 in HNSCC. The relationship between PPFIA1 and ALG3 mRNA expression alterations and prognoses in patients with HNSCC was explored. We found that PPFIA1 and ALG3 were distinctively overexpressed at the mRNA level in HNSCC tissues compared with normal tissues, they had a significant co-occurrence relationship, their mRNA expressions were significantly higher than other PPFIA family members in laryngeal squamous cell carcinoma cell line, and their mRNA expressions were also significantly higher in laryngeal carcinoma cell line than in normal laryngeal cell line. Patients without both PPFIA1 and ALG3 mRNA expression alterations had better overall survival and disease/progression-free survival compared with patients with both PPFIA1 and ALG3 alterations. Based on these findings, PPFIA1 and ALG3 may play roles in oncogene expression in HNSCC. Their combined overexpression is significantly associated with poor survival outcomes. The relationship between them and the mechanism of action in head and neck cancers deserve further investigation.






Similar content being viewed by others
References
Astro V, Asperti C et al (2011) Liprin-alpha1 regulates breast cancer cell invasion by affecting cell motility, invadopodia and extracellular matrix degradation. Oncogene 30(15):1841–1849. https://doi.org/10.1038/onc.2010.562
Astro V, Chiaretti S et al (2014) Liprin-alpha1, ERC1 and LL5 define polarized and dynamic structures that are implicated in cell migration. J Cell Sci 127(Pt 17):3862–3876. https://doi.org/10.1242/jcs.155663
Bhatia A, Burtness B (2017) Novel molecular targets for chemoprevention in malignancies of the head and neck. Cancers 9(9):113. https://doi.org/10.3390/cancers9090113
Blaszczak W, Barczak W et al (2017) Clinical value of monoclonal antibodies and tyrosine kinase inhibitors in the treatment of head and neck squamous cell carcinoma. Med Oncol 34(4):60. https://doi.org/10.1007/s12032-017-0918-1
Blessmann M, Al-Dam A et al (2013) Amplification of the PPFIA1 gene region on 11q13 in oral squamous cell carcinomas (OSCC). J Cranio Maxillo-fac Surg 41(8):845–849. https://doi.org/10.1016/j.jcms.2013.01.040
Cerami E, Gao J et al (2012) The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov 2(5):401–404. https://doi.org/10.1158/2159-8290.CD-12-0095
Chiaretti S, Astro V et al (2016) Effects of the scaffold proteins liprin-alpha1, beta1 and beta2 on invasion by breast cancer cells. Biol Cell 108(3):65–75. https://doi.org/10.1111/boc.201500063
Chiaretti S, de Curtis I (2016) Role of liprins in the regulation of tumor cell motility and invasion. Curr Cancer Drug Targets 16(3):238–248
Choi EJ, Yun JA et al (2014) Prognostic significance of TMEM16A, PPFIA1, and FADD expression in invasive ductal carcinoma of the breast. World J Surg Oncol 12:137. https://doi.org/10.1186/1477-7819-12-137
Choi YW, Bae SM et al (2007) Gene expression profiles in squamous cell cervical carcinoma using array-based comparative genomic hybridization analysis. Int J Gynecol Cancer 17(3):687–696. https://doi.org/10.1111/j.1525-1438.2007.00834.x
Cromer A, Carles A et al (2004) Identification of genes associated with tumorigenesis and metastatic potential of hypopharyngeal cancer by microarray analysis. Oncogene 23(14):2484–2498. https://doi.org/10.1038/sj.onc.1207345
Dai Z, Aryal UK et al (2013) Impact of alg3 gene deletion on growth, development, pigment production, protein secretion, and functions of recombinant Trichoderma reesei cellobiohydrolases in Aspergillus niger. Fungal Genet Biol 61:120–132. https://doi.org/10.1016/j.fgb.2013.09.004
Daraei P, Moore CE (2015) Racial disparity among the head and neck cancer population. J Cancer Educ 30(3):546–551. https://doi.org/10.1007/s13187-014-0753-4
Gao J, Aksoy BA et al (2013) Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal 6(269):pl1. https://doi.org/10.1126/scisignal.2004088
Ginos MA, Page GP et al (2004) Identification of a gene expression signature associated with recurrent disease in squamous cell carcinoma of the head and neck. Cancer Res 64(1):55–63
Glorieux M, Dok R et al (2017) Novel DNA targeted therapies for head and neck cancers: clinical potential and biomarkers. Oncotarget 8(46):81662–81678. https://doi.org/10.18632/oncotarget.20953
Hamoir M, Holvoet E et al (2017) Salvage surgery in recurrent head and neck squamous cell carcinoma: oncologic outcome and predictors of disease free survival. Oral Oncol 67:1–9. https://doi.org/10.1016/j.oraloncology.2017.01.008
Hamoir M, Schmitz S et al (2018) The current role of salvage surgery in recurrent head and neck squamous cell carcinoma. Cancers 10(8):267. https://doi.org/10.3390/cancers10080267
Hawthorne F, Feng S et al (2013) Association mapping of the high-grade myopia MYP3 locus reveals novel candidates UHRF1BP1L, PTPRR, and PPFIA2. Investig Ophthalmol Visual Sci 54(3):2076–2086. https://doi.org/10.1167/iovs.12-11102
Husain N, Neyaz A (2017) Human papillomavirus associated head and neck squamous cell carcinoma: controversies and new concepts. J Oral Biol Craniofacial Res 7(3):198–205. https://doi.org/10.1016/j.jobcr.2017.08.003
Jou A, Hess J (2017) Epidemiology and molecular biology of head and neck cancer. Oncol Res Treat 40(6):328–332. https://doi.org/10.1159/000477127
Katoh M, Katoh M (2003) Identification and characterization of human PPFIA4 gene in silico. Int J Mol Med 12(6):1009–1014
Lampri ES, Chondrogiannis G et al (2015) Biomarkers of head and neck cancer, tools or a gordian knot? Int J Clin Exp Med 8(7):10340–10357
Li WH, Zhou ZJ et al (2016) Detection of OSR2, VAV3, and PPFIA3 methylation in the serum of patients with gastric cancer. Dis Markers 2016:5780538. https://doi.org/10.1155/2016/5780538
Liu B, Ma X et al (2018) Aberrant mannosylation profile and FTX/miR-342/ALG3-axis contribute to development of drug resistance in acute myeloid leukemia. Cell Death Dis 9(6):688. https://doi.org/10.1038/s41419-018-0706-7
Lo Nigro C, Denaro N et al (2017) Head and neck cancer: improving outcomes with a multidisciplinary approach. Cancer Manag Res 9:363–371. https://doi.org/10.2147/CMAR.S115761
Mana G, Clapero F et al (2016) PPFIA1 drives active α5β1 integrin recycling and controls fibronectin fibrillogenesis and vascular morphogenesis. Nat Commun 7:13546. https://doi.org/10.1038/ncomms13546
Pehkonen H, von Nandelstadh P et al (2016) Liprin-alpha1 is a regulator of vimentin intermediate filament network in the cancer cell adhesion machinery. Sci Rep 6:24486. https://doi.org/10.1038/srep24486
Rhodes DR, Kalyana-Sundaram S et al (2007) Oncomine 3.0: genes, pathways, and networks in a collection of 18,000 cancer gene expression profiles. Neoplasia 9(2):166–180. https://doi.org/10.1593/neo.07112
Rhodes DR, Yu J et al (2004) ONCOMINE: a cancer microarray database and integrated data-mining platform. Neoplasia 6(1):1–6
Saleh K, Eid R et al (2018) New developments in the management of head and neck cancer—impact of pembrolizumab. Ther Clin Risk Manag 14:295–303. https://doi.org/10.2147/TCRM.S125059
Serra-Pages C, Kedersha NL et al (1995) The LAR transmembrane protein tyrosine phosphatase and a coiled-coil LAR-interacting protein co-localize at focal adhesions. EMBO J 14(12):2827–2838
Shi ZZ, Jiang YY et al (2014) Identification of putative target genes for amplification within 11q13.2 and 3q27.1 in esophageal squamous cell carcinoma. Clin Transl Oncol 16(7):606–615. https://doi.org/10.1007/s12094-013-1124-z
Stucky A, Sedghizadeh PP et al (2017) Single-cell genomic analysis of head and neck squamous cell carcinoma. Oncotarget 8(42):73208–73218. https://doi.org/10.18632/oncotarget.18021
Tan KD, Zhu Y et al (2008) Amplification and overexpression of PPFIA1, a putative 11q13 invasion suppressor gene, in head and neck squamous cell carcinoma. Genes Chromosomes Cancer 47(4):353–362. https://doi.org/10.1002/gcc.20539
Van Waes C, Musbahi O (2017) Genomics and advances towards precision medicine for head and neck squamous cell carcinoma. Laryngoscope Investig Otolaryngol 2(5):310–319. https://doi.org/10.1002/lio2.86
Yang J, Wu NN et al (2017) PPFIA1 is upregulated in liver metastasis of breast cancer and is a potential poor prognostic indicator of metastatic relapse. Tumour Biol 39(7):1–8. https://doi.org/10.1177/1010428317713492
Yang Y, Zhou Y et al (2018) ALG3 is activated by heat shock factor 2 and promotes breast cancer growth. Med Sci Monit 24:3479–3487. https://doi.org/10.12659/MSM.907461
Funding
This research was supported by Shanghai Pudong Hospital Talent Introduction Program (Grant No. 2015YJ-07).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Research involving human participants and/or animals
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zhou, H., Cao, T., Li, W.P. et al. Combined expression and prognostic significance of PPFIA1 and ALG3 in head and neck squamous cell carcinoma. Mol Biol Rep 46, 2693–2701 (2019). https://doi.org/10.1007/s11033-019-04712-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-019-04712-y