Abstract
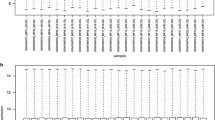
Spinal cord injury (SCI) leads to the loss of sensory, motor, and autonomic function. We aimed to identify the therapeutic targets of-SCI by bioinformatics analysis. The gene expression profile of GSE20907 was downloaded from gene expression omnibus database. By comparing gene expression profiles with control samples, we screened out several differentially expressed genes (DEGs) in 3 days, 2 weeks and 1 month post-SCI. The pathway enrichment and protein–protein interaction (PPI) network analysis for the identified DEGs were performed. Then, transcription factors and microRNAs for DEGs were predicted. We found that up-regulated DEGs mainly participated in cell cycle, oxidative phosphorylation and immune-related pathways; while down-regulated DEGs were mainly involved in oxidative phosphorylation and central nervous system disease signaling pathways. In the constructed PPI network, Bub1, Vascular endothelial growth factor, Topoisomerase IIα (TOP2a) and Cdc20 showed better correspondence with cell cycle, repair system and nerve system. Furthermore, the up-regulated genes (Arpc1b, CD74 and Brd2) significantly mapped to the target genes of transcription factors. The down-regulated genes of 3 days post-injury and the up-regulated genes of 2 weeks post-injury were significantly enriched as the target genes of microRNAs (miR-129 and miR-124). In conclusion, our results may provide guidelines to discuss the collaboration of PPI network in carcinogenesis of SCI.



Similar content being viewed by others
References
Baptiste DC, Fehlings MG (2006) Pharmacological approaches to repair the injured spinal cord. J Neurotrauma 23(3–4):318–334
Becker D, Sadowsky CL, McDonald JW (2003) Restoring function after spinal cord injury. Neurologist 9(1):1–15
Sekhon LH, Fehlings MG (2001) Epidemiology, demographics and pathophysiology of acute spinal cord injury. Spine 26(24S):S2–S12
Bracken MB, Shepard MJ, Collins WF, Holford TR, Young W, Baskin DS, Eisenberg HM, Flamm E, Leo-Summers L, Maroon J (1990) A randomized, controlled trial of methylprednisolone or naloxone in the treatment of acute spinal-cord injury: results of the Second National Acute Spinal Cord Injury Study. N Engl J Med 322(20):1405–1411
Bracken MB, Shepard MJ, Holford TR, Leo-Summers L, Aldrich EF, Fazl M, Fehlings M, Herr DL, Hitchon PW, Marshall LF (1997) Administration of methylprednisolone for 24 or 48 hours or tirilazad mesylate for 48 hours in the treatment of acute spinal cord injury results of the Third National Acute Spinal Cord Injury Randomized Controlled Trial. JAMA 277(20):1597–1604
Pereira JE, LsM Costa, AnM Cabrita, Couto PA, VtM Filipe, Magalhães LG, Fornaro M, Di Scipio F, Geuna S, Maurício AC, Varejão AS (2009) Methylprednisolone fails to improve functional and histological outcome following spinal cord injury in rats. Exp Neurol 220(1):71–81
Breslin K, Agrawal D (2012) The use of methylprednisolone in acute spinal cord injury: a review of the evidence, controversies, and recommendations. Pediatr Emerg Care 28(11):1238–1245
Belkina AC, Denis GV (2012) BET domain co-regulators in obesity, inflammation and cancer. Nat Rev Cancer 12(7):465–477
Zanzoni A, Soler-López M, Aloy P (2009) A network medicine approach to human disease. FEBS Lett 583(11):1759–1765
Barab¨¢si A-Ls, Gulbahce N, Loscalzo J (2011) Network medicine: a network-based approach to human disease. Nat Rev Genet 12(1):56–68
Ahn Y–Y, Bagrow JP, Lehmann S (2010) Link communities reveal multiscale complexity in networks. Nature 466(7307):761–764
Nepusz Ts YuH, Paccanaro A (2012) Detecting overlapping protein complexes in protein–protein interaction networks. Nat Methods 9(5):471–472
Palla G, Derényi I, Farkas Is, Vicsek Ts (2005) Uncovering the overlapping community structure of complex networks in nature and society. Nature 435(7043):814–818
Enright AJ, Van Dongen S, Ouzounis CA (2002) An efficient algorithm for large-scale detection of protein families. Nucleic Acids Res 30(7):1575–1584
Dai M, Wang P, Boyd AD, Kostov G, Athey B, Jones EG, Bunney WE, Myers RM, Speed TP, Akil H (2005) Evolving gene/transcript definitions significantly alter the interpretation of GeneChip data. Nucleic Acids Res 33(20):e175
Irizarry RA, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U, Speed TP (2003) Exploration, normalization and summaries of high density oligonucleotide array probe level data. Biostatistics 4(2):249–264
Da Wei Huang BTS, Lempicki RA (2008) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4(1):44–57
Kanehisa M, Goto S (2000) KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28(1):27–30
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B Stat Methodol (Methodological) 57(1):289–300
Szklarczyk D, Franceschini A, Kuhn M, Simonovic M, Roth A, Minguez P, Doerks T, Stark M, Muller J, Bork P (2011) The STRING database in 2011: functional interaction networks of proteins, globally integrated and scored. Nucleic Acids Res 39(suppl 1):D561–D568
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13(11):2498–2504
Maere S, Heymans K, Kuiper M (2005) BiNGO: a Cytoscape plugin to assess overrepresentation of gene ontology categories in biological networks. Bioinformatics 21(16):3448–3449
Zheng G, Tu K, Yang Q, Xiong Y, Wei C, Xie L, Zhu Y, Li Y (2008) ITFP: an integrated platform of mammalian transcription factors. Bioinformatics 24(20):2416–2417
Margolin AA, Nemenman I, Basso K, Wiggins C, Stolovitzky G, Favera RD, Califano A (2006) ARACNE: an algorithm for the reconstruction of gene regulatory networks in a mammalian cellular context. BMC Bioinform 7(Suppl 1):S7
Kelley BG, Mermelstein PG (2011) Progesterone blocks multiple routes of ion flux. Mol Cell Neurosci 48(2):137–141
Ando J, Sugimoto K, Tamayose K, Sasaki M, Ando M, Oshimi K (2008) Changes in cell morphology and cytoskeletal organization are induced by human mitotic checkpoint gene, Bub1. Biochem Biophys Res Commun 365(4):691–697
Kawashima SA, Yamagishi Y, Honda T, Ishiguro K-i, Watanabe Y (2010) Phosphorylation of H2A by Bub1 prevents chromosomal instability through localizing shugoshin. Science 327(5962):172–177
Eichten A, Adler AP, Cooper B, Griffith J, Wei Y, Yancopoulos GD, Lin HC, Thurston G (2012) Rapid decrease in tumor perfusion following VEGF blockade predicts long-term tumor growth inhibition in preclinical tumor models. Angiogenesis 16(2):429–441
Kim HM, Hwang DH, Lee JE, Kim SU, Kim BG (2009) Ex vivo VEGF delivery by neural stem cells enhances proliferation of glial progenitors, angiogenesis and tissue sparing after spinal cord injury. PLoS ONE 4(3):e4987
Chang Y-W, Goff LA, Li H, Kane-Goldsmith N, Tzatzalos E, Hart RP, Young W, Grumet M (2009) Rapid induction of genes associated with tissue protection and neural development in contused adult spinal cord after radial glial cell transplantation. J Neurotrauma 26(7):979–993
Hasegawa K, Chang Y-W, Li H, Berlin Y, Ikeda O, Kane-Goldsmith N, Grumet M (2005) Embryonic radial glia bridge spinal cord lesions and promote functional recovery following spinal cord injury. Exp Neurol 193(2):394–410
Kim AH, Puram SV, Bilimoria PM, Ikeuchi Y, Keough S, Wong M, Rowitch D, Bonni A (2009) A centrosomal Cdc20-APC pathway controls dendrite morphogenesis in postmitotic neurons. Cell 136(2):322–336
Molli PR, Li D-Q, Bagheri-Yarmand R, Pakala SB, Katayama H, Sen S, Iyer J, Chernoff J, Tsai M-Y, Nair SS (2010) Arpc1b, a centrosomal protein, is both an activator and substrate of Aurora A. J Cell Biol 190(1):101–114
Skinner M (2010) Cell cycle: ARPC1B—a regulator of regulators. Nat Rev Mol Cell Biol 11(8):542–543
Becker-Herman S, Arie G, Medvedovsky H, Kerem A, Shachar I (2005) CD74 is a member of the regulated intramembrane proteolysis-processed protein family. Mol Biol Cell 16(11):5061–5069
Watzka M, Beyenburg S, Blümcke I, Elger CE, Bidlingmaier F, Stoffel-Wagner B (2000) Expression of mineralocorticoid and glucocorticoid receptor mRNA in the human hippocampus. Neurosci Lett 290(2):121–124
Bryan KJ, Zhu X, Harris PL, Perry G, Castellani RJ, Smith MA, Casadesus G (2008) Expression of CD74 is increased in neurofibrillary tangles in Alzheimer’s disease. Mol Neurodegener 3:13
Hirata T, Cui YJ, Funakoshi T, Mizukami Y, Ishikawa Y-i, Shibasaki F, Matsumoto M, Sakabe T (2007) The temporal profile of genomic responses and protein synthesis in ischemic tolerance of the rat brain induced by repeated hyperbaric oxygen. Brain Res 1130:214–222
Hebbes TR, Thorne AW, Crane-Robinson C (1988) A direct link between core histone acetylation and transcriptionally active chromatin. EMBO J 7(5):1395
Tsume M, Kimura-Yoshida C, Mochida K, Shibukawa Y, Amazaki S, Wada Y, Hiramatsu R, Shimokawa K, Matsuo I (2012) Brd2 is required for cell cycle exit and neuronal differentiation through the E2F1 pathway in mouse neuroepithelial cells. Biochem Biophys Res Commun 425(4):762–768
Pal DK, Evgrafov OV, Tabares P, Zhang F, Durner M, Greenberg DA (2003) BRD2 (RING3) is a probable major susceptibility gene for common juvenile myoclonic epilepsy. AmJ Hum Genet 73(2):261–270
Velíšek L, Shang E, Velíšková J, Chachua T, Macchiarulo S, Maglakelidze G, Wolgemuth DJ, Greenberg DA (2011) GABAergic neuron deficit as an idiopathic generalized epilepsy mechanism: the role of BRD2 haploinsufficiency in juvenile myoclonic epilepsy. PLoS One 6(8):e23656
Mi S, Lu J, Sun M, Li Z, Zhang H, Neilly MB, Wang Y, Qian Z, Jin J, Zhang Y (2007) MicroRNA expression signatures accurately discriminate acute lymphoblastic leukemia from acute myeloid leukemia. Proc Natl Acad Sci 104(50):19971–19976
Katada T, Ishiguro H, Kuwabara Y, Kimura M, Mitui A, Mori Y, Ogawa R, Harata K, Fujii Y (2009) microRNA expression profile in undifferentiated gastric cancer. Int J Oncol 34(2):537
Gaur A, Jewell DA, Liang Y, Ridzon D, Moore JH, Chen C, Ambros VR, Israel MA (2007) Characterization of microRNA expression levels and their biological correlates in human cancer cell lines. Cancer Res 67(6):2456–2468
Seike M, Goto A, Okano T, Bowman ED, Schetter AJ, Horikawa I, Mathe EA, Jen J, Yang P, Sugimura H (2009) MiR-21 is an EGFR-regulated anti-apoptotic factor in lung cancer in never-smokers. Proc Natl Acad Sci 106(29):12085–12090
Wu J, Qian J, Li C, Kwok L, Cheng F, Liu P, Perdomo C, Kotton D, Vaziri C, Anderlind C (2010) miR-129 regulates cell proliferation by downregulating Cdk6 expression. Cell Cycle 9(9):1809–1818
Ponomarev ED, Veremeyko T, Barteneva N, Krichevsky AM, Weiner HL (2010) MicroRNA-124 promotes microglia quiescence and suppresses EAE by deactivating macrophages via the C/EBP-[alpha]-PU. 1 pathway. Nat Med 17(1):64–70
Acknowledgments
The article was sponsored by Program of Shanghai Subject Chief Scientist (2012XD1404400) and International Science & Technology Cooperation Program of China (2011DFB30010).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Lingjing Jin and Zhourui Wu should be the co-first author.
Rights and permissions
About this article
Cite this article
Jin, L., Wu, Z., Xu, W. et al. Identifying gene expression profile of spinal cord injury in rat by bioinformatics strategy. Mol Biol Rep 41, 3169–3177 (2014). https://doi.org/10.1007/s11033-014-3176-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-014-3176-8