Abstract
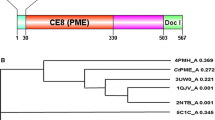
The gene encoding a thermostable pectinase was isolated from a soil metagenome sample. The gene sequence corresponded to an open reading frame of 1,311 bp encoding a translation product of 47.9 kDa. It showed maximum (93 %) identity to a Bacillus licheniformis glycoside hydrolase. Deduced amino acid analysis showed an absence of highly conserved cysteine residues in the N-terminal region at positions 24 and 42, and in the C-terminal region at positions 389, 394, 413 and 424. pQpecJKR01 (pQE30 expression vector containing the pectinase gene) was expressed in Escherichia coli strain M15 as a recombinant fusion protein containing an N-terminal 6× His tag. Biochemical properties of this pectinase were novel. The enzyme had temperature and pH optima of 70 °C and 7.0, respectively, but was active over a broad temperature and pH range. The enzyme was stable at 60 °C with a half-life of 5 h and the enzyme activity was inhibited by 0.1 % diethyl pyrocarbonate and 5 mM dicyclohexyl carbodiimide. The enzyme could be of great use in industrial processes due to its activity over a broad pH range and at high temperature.




Similar content being viewed by others
References
Yadav S, Yadav PK, Yadav D, Yadav KDS (2009) Pectin lyase: a review. Process Biochem 44:1–10
Henrissat B (1991) A classification of glycosyl hydrolases based on amino acid sequence similarities. Biochem J 280:309–316
Cantarel BL, Coutinho PM, Rancurel C, Bernard T, Lombard V, Henrissat B (2009) The carbohydrate-active enzymes database (CAZy), an expert resource for glycogenomics. Nucleic Acids Res 37:D233–D238
Brummell DA, Harpster MH (2001) Cell wall metabolism in fruit softening and quality and its manipulation in transgenic plants. Plant Mol Biol 47:311–340
Abbott DW, Boraston AB (2008) Structural biology of pectin degradation by Enterobacteriaceae. Microbiol Mol Biol Rev 72:301–316
Herron SR, Benen JAE, Scavetta RD, Visser J, Jurnak F (2000) Structure and function of pectic enzymes, virulence factors of plant pathogens. Proc Natl Acad Sci USA 97:8762–8769
Jyothi TC, Singh SA, Appu Rao AG (2005) The contribution of ionic interactions to the conformational stability and function of polygalacturonase from A. niger. Int J Biol Macromol 36:310–317
Asif MH, Nath P (2005) Expression of multiple forms of polygalacturonase gene during ripening in banana fruit. Plant Physiol Biochem 43:177–184
Levin L, Forchiassin F (1998) Culture conditions for the production of pectinolytic enzymes by the white-rot fungus Trametes trogii on a laboratory scale. Acta Biotechnol 18:157–166
Dosanjh NS, Hoondal GS (1996) Production of constitutive, thermostable, hyperactive exo-pectinase from Bacillus GK-8. Biotechnol Lett 18:1435–1438
Hadj-Taieb N, Ayadi M, Khlif M, Mrad K, Hassairi I, Gargouri A (2006) Fermentor production of pectinases on gruel, a local by-product and their use in olive oil extraction. Enzyme Microb Technol 39:1072–1076
Kashyap DR, Vohra PK, Chopra S, Tewari R (2001) Applications of pectinases in the commercial sector, a review. Bioresour Technol 77:215–227
Courtois S, Cappellano CM, Ball M, Francou FX, Normand P, Helynck G, Martinez A, Kolvek SJ, Hopke J, Osburne MS, August PR, Nalin R, Guerineau M, Jeannin P, Simonet P, Pernodet JL (2003) Recombinant environmental libraries provide access to microbial diversity for drug discovery from natural products. Appl Environ Microbiol 69:49–55
Diaz-Torres ML, McNab R, Spratt DA, Villedieu A, Hunt N, Wilson M, Mullany P (2003) Novel tetracycline resistance determinant from the oral metagenome. Antimicrob Agents Chemother 47:1430–1432
Majernik A, Gottschalk G, Daniel R (2001) Screening of environmental DNA libraries for the presence of genes conferring Na+(Li+)/H+ antiporter activity on Escherichia coli, characterization of the recovered genes and the corresponding gene products. J Bacteriol 183:6645–6653
Schipper C, Hornung C, Bijtenhoorn P, Quitschau M, Grond S, Streit WR (2009) Metagenome-derived clones encoding two novel lactonase family proteins involved in biofilm inhibition in Pseudomonas aeruginosa. Appl Environ Microbiol 75:224–233
Li G, Wang K, Liu YH (2008) Molecular cloning and characterization of a novel pyrethroid-hydrolyzing esterase originating from the metagenome. Microb Cell Factories 7:38
Lee MH, Lee CH, Oh TK, Song JK, Yoon JH (2006) Isolation and characterization of a novel lipase from a metagenomic library of tidal flat sediments, evidence for a new family of bacterial lipases. Appl Environ Microbiol 72:7406–7409
Zhou J, Bruns MA, Tiedje JM (1996) DNA recovery from soils of diverse composition. Appl Environ Microbiol 62:316–322
Sharma PK, Capalash N, Kaur J (2007) An improved method for single step purification of metagenomic DNA. Mol Biotechnol 36:61–63
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, New York
Higgins D, Thompson J, Gibson T, Thompson JD, Higgins DG, Gibson TJ (1994) Clustal W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Bendtsen Dyrlov J, Henrik N, von Gunnar H, Soren B (2004) Improved prediction of signal peptides—SignalP 3.0. J Mol Biol 340:783–795
Nielsen H, Krogh A (1998) Prediction of signal peptides and signal anchors by a hidden markov model. In: Proceedings of the sixth international conference on intelligent systems for molecular biology (ISMB6). AAAI Press, Menlo Park, pp 122–130
Lambert C, Leonard N, De Bolle X, Depiereux E (2002) ESyPred3D: prediction of proteins 3D structures. Bioinformatics 18:1250–1256
Singh K (2007) Cloning and characterization of gene(s) involved in catechin biosynthesis in tea (Camellia sinensis). PhD Thesis, Guru Nanak Dev University, Amritsar
Miller GL (1959) Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal Chem 31:426–428
Rey MW, Ramaiya P, Nelson BA, Brody-Karpin SD, Zaretsky EJ, Tang M, de Leon AL, Xiang H, Gusti V, Clausen IG, Olsen PB, Rasmussen MD, Andersen JT, Jorgensen PL, Larsen TS, Sorokin A, Bolotin A, Lapidus A, Galleron N, Ehrlich SD, Berka RM (2004) Complete genome sequence of the industrial bacterium Bacillus licheniformis and comparisons with closely related Bacillus species. Genome Biol 5(Suppl 10):R77
Veith B, Herzberg C, Steckel S, Feesche J, Maurer KH, Ehrenreich P, Baeumer S, Henne A, Liesegang H, Merkl R, Ehrenreich A, Gottschalk G (2004) The complete genome sequence of Bacillus licheniformis DSM13, an organism with great industrial potential. J Mol Microbiol Biotechnol 7:204–211
Gioia J, Yerrapragada S, Qin X, Jiang H, Igboeli OC, Muzny D, Dugan-Rocha S, Ding Y, Hawes A, Liu W, Perez L, Kovar C, Dinh H, Lee S, Nazareth L, Blyth P, Holder M, Buhay C, Tirumalai MR, Liu Y, Dasgupta I, Bokhetache L, Fujita M, Karouia F, Eswara PM, Siefert J, Uzman A, Buzumbo P, Verma A, Zwiya H, McWilliams BD, Olowu A, Clinkenbeard KD, Newcombe D, Golebiewski L, Petrosino JF, Nicholson WL, Fox GE, Venkateswaran K, Highlander SK, Weinstock GM (2007) Paradoxical DNA repair and peroxide resistance gene conservation in Bacillus pumilus SAFR-032. PLoS One 2:E928
Solbak AI, Richardson TH, McCann RT, Kline KA, Bartnek F, Tomlinson G, Tan X, Parra-Gessert L, Frey GJ, Podar M, Luginbuhl P, Gray KA, Mathur EJ, Robertson DE, Burk MJ, Hazlewood GP, Short JM, Kerovuo J (2005) Discovery of pectin-degrading enzymes and directed evolution of a novel pectate lyase for processing cotton fabric. J Biol Chem 280:9431–9438
Copeland A, Lucas S, Lapidus A, Barry K, Detter JC, Glavina del Rio T, Hammon N, Israni S, Dalin E, Tice H, Pitluck S, Thompson LS, Brettin T, Bruce D, Han C, Tapia R, Gilna P, Schmutz J, Larimer F, Land M, Hauser L, Kyrpides N, Mikhailova N, Janssen PH, Kuske CR, Richardson P (2006) Complete sequence of Solibacter usitatus Ellin6076, accession CP000473. US DOE Joint Genome Institute, Walnut Creek, p 1698
Esquivel JCC, Voget CE (2004) Purification and partial characterization of an acidic polygalacturonase from Aspergillus kawachii. J Biotechnol 110:21–28
Bonivento D, Pontiggia D, Di Matteo A, Fernandez-Recio J, Salvi G, Tsernoglou D, Cervone F, De Lorenzo G, Federici L (2008) Crystal structure of the endopolygalacturonase from the phytopathogenic fungus Colletotrichum lupini and its interaction with polygalacturonase-inhibiting proteins. Proteins 70:294–299
Cho SW, Lee S, Shin W (2001) The X-ray structure of Aspergillus aculeatus polygalacturonase and a modeled structure of the polygalacturonase-octagalacturonate complex. J Mol Biol 311:863–878
van Santen Y, Jacques AEB, Schroter KH, Kalk KH, Armand S, Visser J, Dijkstra BW (1999) 1.68-Å crystal structure of endopolygalacturonase II from Aspergillus niger and identification of active site residues by site-directed mutagenesis. J Biol Chem 274:30474–30480
Federici L, Caprari C, Mattei B, Savino C, Di Matteo A, De Lorenzo G, Cervone F, Tsernoglou D (2001) Structural requirements of endopolygalacturonase for the interaction with PGIP (polygalacturonase-inhibiting protein). Proc Natl Acad Sci USA 98:13425
van Pouderoyen G, Snijder HJ, Benen JA, Dijkstra BW (2003) Structural insights into the processivity of endopolygalacturonase I from Aspergillus niger. FEBS Lett 554:462–466
Martens-Uzunova ES, Zandleven JS, Benen JAE, Awad H, Kools HJ, Beldman G, Voragen AGJ, Van den Berg JA, Schaap PJ (2006) A new group of exo-acting family 28 glycoside hydrolases of Aspergillus niger that are involved in pectin degradation. Biochem J 400:43–52
Kluskens LD, Gert-Jan van Alebeek WM, Walther J, Voragen AGJ, de Vos WM, van der Oost J (2005) Characterization and mode of action of an exopolygalacturonase from the hyperthermophilic bacterium Thermotoga maritima. FEBS J 272:5464–5473
Markovic O, Stefan J (2001) Pectin degrading glycoside hydrolases of family 28, sequence—structural features, specificities and evolution. Protein Eng 14:615–631
Chang A, Scheer M, Grote A, Schomburg I, Schomburg D (2009) BRENDA, AMENDA and FRENDA the enzyme information system, new content and tools in 2009. Nucleic Acids Res 37:D588–D592. http://www.brenda-enzymes.info/index
Bernhard S, Zeikus JG (1983) Characterization of pectinolytic enzymes of Clostridium thermosulfurogenes. FEMS Microbiol Lett 17:295–298
Miyairi K, Okuno T, Sawai K (1985) Purification and properties of endopolygalacturonase I from Stereum purpureum, a factor inducing silver-leaf symptoms on apple trees. Agric Biol Chem 49(Suppl 4):1111–1118
Horikoshi K (1972) Production of alkaline enzymes by alkalophilic microorganisms. Agric Biol Chem 36:285–293
Martins ES, Silva D, da Silva R, Leite RS, Gomes E (2007) Purification and characterization of polygalacturonase produced by thermophilic Thermoascus aurantiacus CBMAI-756 in submerged fermentation. Antonie Van Leeuwenhoek 91(Suppl 3):291–299
Gillespie AM, Cook K, Coughlan MP (1990) Characterization of an endopolygalacturonase produced by solid–state cultures of the aerobic fungus Penicillium capsulatum. J Biotechnol 17:279–292
Hirose N, Kishida M, Kawasaki H, Sakai T (1999) Purification and characterization of an endo-polygalacturonase from a mutant of Saccharomyces cerevisiae. Biosci Biotechnol Biochem 63:1100–1103
Soriano M, Diaz P, Pastor FIJ (2005) Pectinolytic systems of two aerobic sporogenous bacterial strains with high activity on pectin. Curr Microbiol 50:114–118
Le Van T, Tuyen H, Helmke E, Le Tran B, Schweder T (2001) Cloning of two pectate lyase genes from the marine Antarctic bacterium Pseudoalteromonas haloplanktis strain ANT/505 and characterization of the enzymes. Extremophiles 5:35–44
Thangudu RR, Malini M, Srinivasan N, Cadet F, Sowdhamini R, Offmann B (2008) Analysis on conservation of disulphide bonds and their structural features in homologous protein domain families. BMC Struct Biol 8:55
Vullo A, Frasconi P (2004) Disulfide connectivity prediction using recursive neural networks and evolutionary information. Bioinformatics 20:653–659
Frasconi P, Passerini A, Vullo A (2002) A two-stage SVM architecture for predicting the disulfide bonding state of cysteines. In: Proceedings of IEEE workshop on neural networks for signal processing pp 25–34
Ceroni A, Frasconi P, Passerini A, Vullo A (2003) Predicting the disulfide bonding state of cysteines with combinations of kernel machines. J VLSI Signal Process 35:287–295
Ferre F, Clote P (2005) DiANNA: a web server for disulfide connectivity prediction. Nucleic Acids Res 33(web server issue):W230–W232
Kaur G, Kumar S, Satyanarayana T (2004) Production, characterization and application of a thermostable polygalacturonase of a thermophilic mould Sporotrichum thermophile Apinis. Bioresour Technol 94:239–243
You C, Huang Q, Xue H, Xu Y, Lu H (2010) Potential hydrophobic interaction between two cysteines in interior hydrophobic region improves thermostability of a family 11 xylanase from Neocallimastix patriciarum. Biotechnol Bioeng 105:861–870
Pijning T, van Pouderoyen G, Kluskens L, van der Oost J, Dijkstra BW (2009) The crystal structure of a hyperthermoactive exopolygalacturonase from Thermotoga maritima reveals a unique tetramer. FEBS Lett 583:3665–3670
Vieille C, Burdette SD, Zeikus GJ (1996) Thermozymes. Biotechnol Annu Rev 2:1–83
Watanabe K, Chishiro K, Kitamura K, Suzuki Y (1991) Proline residues responsible for thermostability occur with high frequency in the loop regions of an extremely thermostable oligo-1,6-glucosidase from Bacillus thermoglucosidasius KP1006. J Biol Chem 266:24287–24294
Nawani N, Kaur J (2007) Studies on lipolytic isoenzymes from a thermophilic Bacillus sp., production, purification and biochemical characterization. Enzyme Microb Technol 40:881–887
Acknowledgments
Fellowship granted to RS by University Grants Commission (UGC), India is acknowledged.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Singh, R., Dhawan, S., Singh, K. et al. Cloning, expression and characterization of a metagenome derived thermoactive/thermostable pectinase. Mol Biol Rep 39, 8353–8361 (2012). https://doi.org/10.1007/s11033-012-1685-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-012-1685-x