Abstract
The small non-coding important regulatory molecules, microRNAs (miRNAs), have been widely and deeply studied especially combining high-throughput sequencing technologies. Here, we attempted to track detailed miRNA precursor metabolic products and gain further insight into pre-miRNA processing by completely analyzing high-throughput sequencing data. Highly expressed miRNA precursors could be entirely covered by various short RNAs and small RNA fragments with a hierarchical distribution. miRNAs and some miRNA* regions were detected quite abundant short RNAs as expected, while other regions of precursors were found shorter RNAs or small fragments with fewer sequence counts. Furthermore, we developed a method to analyze relative expression levels of special RNA classes according to divergence of 5′ and 3′ ends, respectively. Generally, there were several quite abundant RNA classes from a given miRNA locus, which suggested dominant cleavage sites of Drosha and Dicer during pre-miRNA processing. Compared with 3′ end, dominant cleavage site in 5′ end always focused on a specific position, which ensured conservation of the identity of miRNA (5′-seed sequence, nucleotides 2–8). Overall, a comprehensive analysis of sequencing data can be used to track pre-miRNA metabolic products and mechanism of pre-miRNA processing and metabolism.




Similar content being viewed by others
References
Bushati N, Cohen SM (2007) microRNA functions. Annu Rev Cell Dev Biol 23:175–205
He L, Hannon GJ (2004) Micrornas: small RNAs with a big role in gene regulation. Nat Rev Genet 5:522–531
Bartel DP (2004) MicroRNAs genomics, biogenesis, mechanism, and function. Cell 116:281–297
Lee Y, Jeon K, Lee JT, Kim S, Kim VN (2002) MicroRNA maturation: stepwise processing and subcellular localization. EMBO J 21:4663–4670
Lund E, Guttinger S, Calado A, Dahlberg JE, Kutay U (2004) Nuclear export of microRNA precursors. Science 303:95–98
Chendrimada TP, Gregory RI, Kumaraswamy E, Norman J, Cooch N et al (2005) TRBP recruits the Dicer complex to Ago2 for microRNA processing and gene silencing. Nature 436:740–744
O’Toole AS, Miller S, Haines N, Zink MC, Serra MJ (2006) Comprehensive thermodynamic analysis of 3′ double-nucleotide overhangs neighboring Watson-Crick terminal base pairs. Nucleic Acids Res 34:3338–3344
Filipowicz W, Bhattacharyya SN, Sonenberg N (2008) Mechanisms of post-transcriptional regulation by microRNAs: are the answers in sight? Nat Rev Genet 9:102–114
Okamura K, Phillips MD, Tyler DM, Duan H, Chou YT et al (2008) The regulatory activity of microRNA star species has substantial influence on microRNA and 3′ UTR evolution. Nat Struct Mol Biol 15:354–363
Jagadeeswaran G, Zheng Y, Sumathipala N, Jiang HB, Arrese EL et al (2010) Deep sequencing of small RNA libraries reveals dynamic regulation of conserved and novel microRNAs and microRNA-stars during silkworm development. BMC Genom 11:52
Czech B, Zhou R, Erlich Y, Brennecke J, Binari R et al (2009) Hierarchical rules for argonaute loading in drosophila. Mol Cell 36:445–456
Ghildiyal M, Xu J, Seitz H, Weng ZP, Zamore PD (2010) Sorting of Drosophila small silencing RNAs partitions microRNA* strands into the RNA interference pathway. RNA 16:43–56
Guo L, Lu ZH (2010) The fate of miRNA* strand through evolutionary analysis: implication for degradation as merely carrier strand or potential regulatory molecule? PLoS ONE 5:e11387
Borel C, Antonarakis SE (2008) Functional genetic variation of human miRNAs and phenotypic consequences. Mamm Genome 19:503–509
Cummins JM, He YP, Leary RJ, Pagliarini R, Diaz LA et al (2006) The colorectal microRNAome. Proc Natl Acad Sci USA 103:3687–3692
Kuchenbauer F, Morin RD, Argiropoulos B, Petriv OI, Griffith M et al (2008) In-depth characterization of the microRNA transcriptome in a leukemia progression model. Genome Res 18:1787–1797
Lagos-Quintana M, Rauhut R, Yalcin A, Meyer J, Lendeckel W et al (2002) Identification of tissue-specific microRNAs from mouse. Curr Biol 12:735–739
Morin RD, O’Connor MD, Griffith M, Kuchenbauer F, Delaney A et al (2008) Application of massively parallel sequencing to microRNA profiling and discovery in human embryonic stem cells. Genome Res 18:610–621
Rathjen T, Pais H, Sweetman D, Moulton V, Munsterberg A et al (2009) High throughput sequencing of microRNAs in chicken somites. Febs Lett 583:1422–1426
Ruby JG, Jan C, Player C, Axtell MJ, Lee W et al (2006) Large-scale sequencing reveals 21U-RNAs and additional microRNAs and endogenous siRNAs in C-elegans. Cell 127:1193–1207
Azuma-Mukai A, Oguri H, Mituyama T, Qian ZR, Asai K et al (2008) Characterization of endogenous human Argonautes and their miRNA partners in RNA silencing. Proc Natl Acad Sci USA 105:7964–7969
Underwood JG, Uzilov AV, Katzman S, Onodera CS, Mainzer JE et al (2010) FragSeq: transcriptome-wide RNA structure probing using high-throughput sequencing. Nat Methods 7:995–1001
Kertesz M, Wan Y, Mazor E, Rinn JL, Nutter RC et al (2010) Genome-wide measurement of RNA secondary structure in yeast. Nature 467:103–107
Westhof E, Romby P (2010) The RNA structurome: high-throughput probing. Nat Methods 7:965–967
Friedlander MR, Chen W, Adamidi C, Maaskola J, Einspanier R et al (2008) Discovering microRNAs from deep sequencing data using miRDeep. Nat Biotechnol 26:407–415
Griffiths-Jones S, Saini HK, van Dongen S, Enright AJ (2008) miRBase: tools for microRNA genomics. Nucleic Acids Res 36:D154–D158
Langmead B, Trapnell C, Pop M, Salzberg SL (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10:212
Guo L, Liang T, Lu Z (2011) A comprehensive study of multiple mapping and feature selection for correction strategy in the analysis of small RNAs from SOLiD sequencing. BioSystems 104:87–93
de Hoon MJL, Taft RJ, Hashimoto T, Kanamori-Katayama M, Kawaji H et al (2010) Cross-mapping and the identification of editing sites in mature microRNAs in high-throughput sequencing libraries. Genome Res 20:257–264
Burroughs AM, Ando Y, de Hoon MJ, Tomaru Y, Nishibu T et al (2010) A comprehensive survey of 3′ animal miRNA modification events and a possible role for 3′ adenylation in modulating miRNA targeting effectiveness. Genome Res 20:1398–1410
Ebhardt HA, Tsang HH, Dai DC, Liu YF, Bostan B et al (2009) Meta-analysis of small RNA-sequencing errors reveals ubiquitous post-transcriptional RNA modifications. Nucleic Acids Res 37:2461–2470
Lee LW, Zhang S, Etheridge A, Ma L, Martin D et al (2010) Complexity of the microRNA repertoire revealed by next generation sequencing. RNA 16:2170–2180
Li JJ, Yang ZY, Yu B, Liu J, Chen XM (2005) Methylation protects miRNAs and siRNAs from a 3′-end uridylation activity in Arabildopsis. Curr Biol 15:1501–1507
Lu SF, Sun YH, Chiang VL (2009) Adenylation of plant miRNAs. Nucleic Acids Res 37:1878–1885
Guo L, Lu Z (2010) Global expression analysis of miRNA gene cluster and family based on isomiRs from deep sequencing data. Comput Biol Chem 34:165–171
Bartel DP (2009) MicroRNAs: target recognition and regulatory functions. Cell 136:215–233
Acknowledgments
This work was supported by projects 30871393, 30900836 and 60971021 of the National Natural Science Foundation of China and funded by Tsinghua National Laboratory for Information Science and Technology (TNList) Cross-discipline Foundation. The work was also supported by a research grant from the Innovation Project for Graduate Students of Jiangsu Province (No. CX10B_081Z), the Scientific Research Foundation of Graduate School of Southeast University, science & technology project in Nanjing (201001095) and pre-research Project for National Natural Science Foundation Supported by Southeast University (KJ2010442). The authors have declared that no competing interests exist.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Li Guo and Hailing Li contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11033_2011_950_MOESM1_ESM.tif
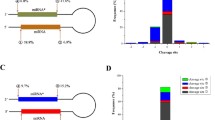
Fig. S1. An example of tracking miRNA precursor metabolic products based on high-throughput sequencing. Accurate mapping is performed using Bowtie, and short RNAs are not showed here if their sequence counts are less than 10. Two distinct regions that can yield hsa-miR-24 and hsa-miR-24-2* are accumulated with various short RNAs with various 5′ and/or 3′ ends, and these called multiple isomiRs always are detected with higher copy numbers. Other regions of hsa-mir-24 also can be detected all kinds of mutual overlapping shorter RNAs (especially shorter RNAs with 6 nt length). In fact, we can track detailed metabolic process with continuous lengths of short RNAs, but short RNAs more than 6 nt length always have fewer sequence counts. (TIFF 80 kb)
Rights and permissions
About this article
Cite this article
Guo, L., Li, H., Lu, J. et al. Tracking miRNA precursor metabolic products and processing sites through completely analyzing high-throughput sequencing data. Mol Biol Rep 39, 2031–2038 (2012). https://doi.org/10.1007/s11033-011-0950-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-011-0950-8