Abstract
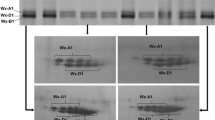
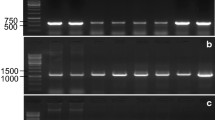
The Wx gene encodes the granule-bound starch synthase I or waxy protein, which is the sole enzyme responsible for amylose synthesis in wheat seeds. Triticum urartu and einkorn (T. monococcum L. ssp. monococcum), which are related to the A genome of bread wheat, could be important sources of variation for this gene. This study evaluated the Wx gene variability in 52 accessions of these species and compared their nucleotide sequences with the Wx-A1a allele of bread wheat. The level of polymorphism found was high, although not distributed equally between the two species. Five different alleles were found in T. urartu, of which four were novel (Wx-A u 1b, -A u 1c, -A u 1d and -A u 1e). All einkorn accessions had the same allele, which was also novel and was named Wx-A m 1a. A comparison between the proteins deduced from the novel alleles and the Wx-A1a protein showed that there were up to 33 amino acid changes in both the transit peptide and the mature protein. These results showed that these species, especially T. urartu, are a potential source of novel waxy variants.



Similar content being viewed by others
References
Ainsworth C, Clark J, Balsdon J (1993) Expression, organisation and structure of the genes encoding the waxy protein (granule-bound starch synthase) in wheat. Plant Mol Biol 22:67–82
Baldwin PM (2001) Starch granule-associated protein and polypeptides: a review. Starch/Stärcke 53:475–503
Brandolini A, Vaccino P, Boggini G, Özkan H, Kilian B, Salamini F (2006) Quantification of genetic relationships among A genomes of wheats. Genome 49:297–305
Caballero L, Bancel E, Debiton C, Branlard G (2008) Granule-bound starch synthase (GBSS) diversity of ancient wheat and related species. Plant Breed 127:548–553
Chao S, Sharp PJ, Worland AJ, Warham EJ, Koebner RMD, Gale MD (1989) RFLP-based genetic maps of wheat homoeologous group 7 chromosomes. Theor Appl Genet 78:495–504
Dvorak J, McGuire PE, Cassidy B (1988) Apparent source of the A genomes of wheats inferred from polymorphism in abundance and restriction fragment length of repeated nucleotide sequences. Genome 30:680–689
Dvorak J, Pd Terlizzi, Zhang H-B, Resta P (1993) The evolution of polyploid wheats: identification of the A genome donor species. Genome 36:21–31
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Feuillet C, Langridge P, Waugh R (2008) Cereal breeding takes a walk on the wild side. Trends Genet 24:24–32
Guzmán C, Alvarez JB (2012) Molecular characterization of a novel waxy allele (Wx-A u 1a) from Triticum urartu Thum. ex Gandil. Genet Resour Crop Evol 59:971–979
Guzmán C, Caballero L, Alvarez JB (2009) Variation in Spanish cultivated einkorn wheat (Triticum monococcum L. ssp. monococcum) as determined by morphological traits and waxy proteins. Genet Resour Crop Evol 56:601–604
Guzmán C, Caballero L, Moral A, Alvarez JB (2010) Genetic variation for waxy proteins and amylose content in Spanish spelt wheat (Triticum spelta L.). Genet Resour Crop Evol 57:721–725
Guzmán C, Caballero L, Alvarez JB (2011) Molecular characterisation of the Wx-B1 allelic variants identified in cultivated emmer wheat and comparison with those of durum wheat. Mol Breed 28:403–411
Guzmán C, Caballero L, Martín LM, Alvarez JB (2012) Waxy genes from spelt wheat: new alleles for modern wheat breeding and new phylogenetic inferences about the origin of this species. Ann Bot 110:1161–1171
Huang XQ, Brûlé-Babel A (2012) Sequence diversity, haplotype analysis, association mapping and functional marker development in the waxy and starch synthase IIa genes for grain-yield-related traits in hexaploid wheat (Triticum aestivum L.). Mol Breed 30:627–635
Kiribuchi-Otobe C, Nagamine T, Yanagisawa T, Ohnishi M, Yamaguchi I (1997) Production of hexaploid wheats with waxy endosperm character. Cereal Chem 74:72–74
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Leterrier M, Holappa LD, Broglie KE, Beckles DM (2008) Cloning, characterisation and comparative analysis of a starch synthase IV gene in wheat: functional and evolutionary implications. BMC Plant Biol 8:98
Li W, Gao Z, Xiao W, Wei YM, Liu YX, Chen GY, Pu ZE, Chen HP, Zheng YL (2012) Molecular diversity of restriction enzyme sites, Indels and upstream open reading frames (uORFs) of 5′ untranslated regions (UTRs) of Waxy genes in Triticum L. and Aegilops L. species. Genet Resour Crop Evol 59:1625–1647
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452
Liu YX, Li W, Wei YM, Chen GY, Zheng YL (2009) Molecular characterization of the waxy gene in einkorn wheat. J Plant Sci 4:114–121
Mason-Gamer RJ, Weil CF, Kellogg EA (1998) Granule-bound starch synthase: structure, function, and phylogenetic utility. Mol Biol Evol 15:1658–1673
McIntosh RA, Yamazaki Y, Dubcovsky J, Rogers WJ, Morris C, Appels R, Xia XC (2013) Catalogue of gene symbols for wheat. http://www.shigen.nig.ac.jp/wheat/komugi/genes/macgene/2013/GeneSymbol.pdf. Accessed 22 Apr 2014
Miller TE, Reader SM (1980) Variation in the meiotic chromosome pairing of hybrids between hexaploid and diploid wheats. Cereal Res Commun 8:477–483
Miura H, Sugawara A (1996) Dosage effects of the three Wx genes on amylose synthesis in wheat endosperm. Theor Appl Genet 93:1066–1070
Monari AM, Simeone MC, Urbano M, Margiotta B, Lafiandra D (2005) Molecular characterization of new waxy mutants identified in bread and durum wheat. Theor Appl Genet 110:1481–1489
Murai J, Taira T, Ohta D (1999) Isolation and characterization of the three Waxy genes encoding the granule-bound starch synthase in hexaploid wheat. Gene 234:71–79
Nakamura T, Yamamori M, Hirano H, Hidaka S, Nagamine T (1995) Production of waxy (amylose-free) wheats. Mol Gen Genet 248:253–259
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
Ortega R, Alvarez JB, Guzmán C (2014) Characterization of the Wx gene in diploid Aegilops species and its potential use in wheat breeding. Genet Resour Crop Evol 61:369–382
Rawat N, Sehgal SK, Joshi A, Rothe N, Wilson DL, McGraw N, Vadlani PV, Li W, Gill BS (2012) A diploid wheat TILLING resource for wheat functional genomics. BMC Plant Biol 12:205
Rice P, Longden I, Bleasby A (2000) EMBOSS: the European molecular biology open software suite. Trends Genet 16:276–277
Rodriguez-Quijano M, Vázquez JF, Carrillo JM (2004) Waxy proteins and amylose content in diploid Triticeae species with genomes A, S and D. Plant Breed 123:294–296
Sim NL, Kumar P, Hu J, Henikoff S, Schneider G, Ng PC (2012) SIFT web server: predicting effects of amino acid substitutions on proteins. Nucleic Acids Res 40(W1):W452–W457
Slade AJ, Fuerstenberg SI, Loeffler D, Steine MN, Facciotti D (2005) A reverse genetic, nontransgenic approach to wheat crop improvement by TILLING. Nat Biotechnol 23:75–81
Stacey J, Isaac P (1994) Isolation of DNA from plants. In: Isaac PG (ed) Methods in molecular biology: protocols for nucleic acid analysis by non-radiactive probes. Humana Press, Totawa, pp 9–15
Tamura K, Nei M, Kumar S (2004) Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc Natl Acad Sci USA 101:11030–11035
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Urbano M, Margiotta B, Colaprico G, Lafiandra D (2002) Waxy proteins in diploid, tetraploid and hexaploid wheats. Plant Breed 121:465–469
Watterson GA (1975) On the number of segregating sites in genetical models without recombination. Theor Popul Biol 7:256–276
Wu Y, Weber JL, Vladutiu GD, Tarnopolsky MA (2011) Six novel mutations in the myophosphorylase gene in patients with McArdle disease and a family with pseudo-dominant inheritance pattern. Mol Genet Metab 104:587–591
Yamamori M (2009) Amylose content and starch properties generated by five variant Wx alleles for granule-bound starch synthase in common wheat (Triticum aestivum L.). Euphytica 165:607–614
Yamamori M, Guzmán C (2013) SNPs and an insertion sequence in five Wx-A1 alleles as factors for variant Wx-A1 protein in wheat. Euphytica 192:325–338
Yamamori M, Yamamoto K (2011) Effects of two novel Wx-A1 alleles of common wheat (Triticum aestivum L.) on amylose and starch properties. J Cereal Sci 54:229–235
Yamamori M, Nakamura T, Endo TR, Nagamine T (1994) Waxy protein deficiency and chromosomal location of coding genes in common wheat. Theor Appl Genet 89:179–184
Yamamori M, Fujita S, Hayakawa K, Matsuki J, Yasui T (2000) Genetic elimination of a starch granule protein, SGP-1, of wheat generates an altered starch with apparent high amylose. Theor Appl Genet 101:21–29
Yan L, Bhave M (2001) Characterization of waxy proteins and waxy genes of Triticum timopheevii and T. zhukovskyi and implications for evolution of wheat. Genome 44:582–588
Yan L, Bhave M, Fairclough R, Konik C, Rahman S, Appels R (2000) The genes encoding granule-bound starch synthases at the waxy loci of the A, B, and D progenitors of common wheat. Genome 43:264–272
Yasui T, Sasaki T, Matsuki J, Yamamori M (1997) Waxy endosperm mutants of bread wheat (Triticum aestivum L.) and their starch properties. Breed Sci 47:161–163
Acknowledgments
This research was supported by grant AGL2010-19643-C02-01 from the Spanish Ministry of Economy and Competitiveness, co-financed with the European Regional Development Fund (FEDER) from the European Union. We thank the National Small Grain Collection (Aberdeen, USA) and the Institute for Plant Genetics and Crop Plant Research (Gatersleben, Germany) for supplying the material analyzed.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ortega, R., Guzmán, C. & Alvarez, J.B. Wx gene in diploid wheat: molecular characterization of five novel alleles from einkorn (Triticum monococcum L. ssp. monococcum) and T. urartu . Mol Breeding 34, 1137–1146 (2014). https://doi.org/10.1007/s11032-014-0105-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-014-0105-4