Abstract
Cancer is one of the leading causes of death worldwide and requires intense and growing research investments from the public and private sectors. This is expected to lead to the development of new medicines. A determining factor in this process is the structural understanding of molecules with potential anticancer properties. Since the major compounds used in cancer therapies fail to encompass every spectrum of this disease, there is a clear need to research new molecules for this purpose. As it follows, we have studied the class of quinolinones that seem effective for such therapy. This paper describes the structural elucidation of a novel dihydroquinoline by single-crystal X-ray diffraction and spectroscopy characterization. Topology studies were carried through Hirshfeld surfaces analysis and molecular electrostatic potential map; electronic stability was evaluated from the calculated energy of frontier molecular orbitals. Additionally, in silico studies by molecular docking indicated that this dihydroquinoline could act as an anticancer agent due to their higher binding affinity with human aldehyde dehydrogenase 1A1 (ALDH 1A1). Tests in vitro were performed for VERO (normal human skin keratinocytes), B16F10 (mouse melanoma), and MDA-MB-231 (metastatic breast adenocarcinoma), and the results certified that compound as a potential anticancer agent.
Graphic abstract
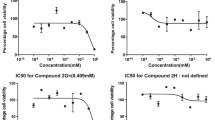
A Dihydroquinoline derivative was tested against three cancer cell lines and the results attest that compound as potential anticancer agent.








Similar content being viewed by others
References
Müller-Schiffmann A, Sticht H, Korth C (2012) Hybrid compounds. BioDrugs 26:21–31. https://doi.org/10.2165/11597630-000000000-00000
Lahtchev KL, Batovska DI, Parushev SP et al (2008) Antifungal activity of chalcones: a mechanistic study using various yeast strains. Eur J Med Chem 43:2220–2228. https://doi.org/10.1016/j.ejmech.2007.12.027
Custodio J, Faria E, Sallum L et al (2017) The influence of methoxy and ethoxy groups on supramolecular arrangement of two methoxy-chalcones. J Braz Chem Soc 28:2180–2191. https://doi.org/10.21577/0103-5053.20170067
Carvalho PS, Custodio JMF, Vaz WF et al (2017) Conformation analysis of a novel fluorinated chalcone. J Mol Model 23:97. https://doi.org/10.1007/s00894-017-3245-8
Silva WA, Andrade CKZ, Napolitano HB et al (2013) Biological and structure-activity evaluation of chalcone derivatives against bacteria and fungi. J Braz Chem Soc 24:133–144. https://doi.org/10.1590/S0103-50532013000100018
Ávila HP, de Smânia FAE, Monache FD, Smânia A (2008) Structure–activity relationship of antibacterial chalcones. Bioorg Med Chem 16:9790–9794. https://doi.org/10.1016/j.bmc.2008.09.064
Nowakowska Z (2007) A review of anti-infective and anti-inflammatory chalcones. Eur J Med Chem 42:125–137. https://doi.org/10.1016/j.ejmech.2006.09.019
Rosa GP, Seca AML, do Barreto MC et al (2019) Chalcones and flavanones bearing hydroxyl and/or methoxyl groups: synthesis and biological assessments. Appl Sci 9:2846. https://doi.org/10.3390/app9142846
Al-Karawi AJM, Hammood AJ, Awad AA et al (2018) Synthesis and mesomorphism behaviour of chalcones and pyrazoles type compounds as photo-luminescent materials. Liq Cryst 45:1603–1619. https://doi.org/10.1080/02678292.2018.1446553
Özaslan MS, Demir Y, Aslan HE et al (2018) Evaluation of chalcones as inhibitors of glutathione S-transferase. J Biochem Mol Toxicol 32:e22047. https://doi.org/10.1002/jbt.22047
Padhye S, Ahmad A, Oswal N et al (2010) Bioorganic & medicinal chemistry letters fluorinated 2 0-hydroxychalcones as garcinol analogs with enhanced antioxidant and anticancer activities. Bioorg Med Chem Lett 20:5818–5821. https://doi.org/10.1016/j.bmcl.2010.07.128
Scozzafava A, Owa T, Mastrolorenzo A, Supuran C (2003) Anticancer and antiviral sulfonamides. Curr Med Chem 10:925–953. https://doi.org/10.2174/0929867033457647
Supuran CT (2008) Carbonic anhydrases: novel therapeutic applications for inhibitors and activators. Nat Rev Drug Discov 7:168–181. https://doi.org/10.1038/nrd2467
Abdelli A, Gaucher A, Efrit ML et al (2015) Arylation of allylphosphonates and application to the preparation of phosphonomethyl-coumarin, -quinolinone and -benzoxepinone skeletons. Tetrahedron Lett 56:1679–1681. https://doi.org/10.1016/j.tetlet.2015.02.038
Chung HJ, Kamli MR, Lee HJ et al (2014) Discovery of quinolinone derivatives as potent FLT3 inhibitors. Biochem Biophys Res Commun 445:561–565. https://doi.org/10.1016/j.bbrc.2014.02.029
Ghorab MM, Ragab FA, Heiba HI et al (2015) Synthesis, anticancer and radiosensitizing evaluation of some novel sulfonamide derivatives. Eur J Med Chem 92:682–692. https://doi.org/10.1016/j.ejmech.2015.01.036
De Castro MRC, Aragão ÂQ, da Silva CC et al (2015) Conformational variability in sulfonamide chalcone hybrids: crystal structure and cytotoxicity. J Braz Chem Soc 27:884–898. https://doi.org/10.5935/0103-5053.20150341
Snejko N, Cascales C, Gomez-Lor B et al (2002) From rational octahedron design to reticulation serendipity. A thermally stable rare earth polymeric disulfonate family with CdI2-like structure, bifunctional catalysis and optical properties. Chem Commun. https://doi.org/10.1039/b202639b
Dolomanov OV, Bourhis LJ, Gildea RJ et al (2009) OLEX2: a complete structure solution, refinement and analysis program. J Appl Crystallogr 42:339–341. https://doi.org/10.1107/S0021889808042726
Sheldrick GM (2015) SHELXT: integrated space-group and crystal-structure determination. Acta Crystallogr A 71:3–8. https://doi.org/10.1107/S2053273314026370
Sheldrick GM (2015) Crystal structure refinement with SHELXL. Acta Crystallogr Sect C 71:3–8. https://doi.org/10.1107/S2053229614024218
Farrugia LJ (2012) WinGX and ORTEP for Windows: an update. J Appl Crystallogr 45:849–854. https://doi.org/10.1107/S0021889812029111
Macrae CF, Bruno IJ, Chisholm JA et al (2008) Mercury CSD 2.0—new features for the visualization and investigation of crystal structures. J Appl Crystallogr 41:466–470. https://doi.org/10.1107/S0021889807067908
Spek AL (2003) Single-crystal structure validation with the program PLATON. J Appl Crystallogr 36:7–13. https://doi.org/10.1107/S0021889802022112
McKinnon JJ, Spackman MA, Mitchell AS (2004) Novel tools for visualizing and exploring intermolecular interactions in molecular crystals. Acta Crystallogr Sect B Struct Sci 60:627–668. https://doi.org/10.1107/S0108768104020300
Allen FH (2002) The Cambridge structural database: a quarter of a million crystal structures and rising. Acta Crystallogr Sect B Struct Sci 58:380–388. https://doi.org/10.1107/S0108768102003890
Groom CR, Allen FH (2014) The Cambridge structural database in retrospect and prospect. Angew Chemie 53:662–671. https://doi.org/10.1002/anie.201306438
McKinnon JJ, Jayatilaka D, Spackman MA (2007) Towards quantitative analysis of intermolecular interactions with Hirshfeld surfaces. Chem Commun. https://doi.org/10.1039/b704980c
Spackman MA, McKinnon JJ (2002) Fingerprinting intermolecular interactions in molecular crystals. CrystEngComm 4:378–392. https://doi.org/10.1039/B203191B
Peón A, Naulaerts S, Ballester PJ (2017) Predicting the reliability of drug-target interaction predictions with maximum coverage of target space. Sci Rep 7:3820. https://doi.org/10.1038/s41598-017-04264-w
(2017) OMEGA v.2.5.1: OpenEye Scientific Software, Santa Fe, NM, USA. http://www.eyesopen.com
Hawkins PCD, Skillman GA, Warren GL et al (2010) Conformer generation with OMEGA: algorithm and validation using high quality structures from the protein databank and cambridge structural database. J Chem Inf Model 50:572–584. https://doi.org/10.1021/ci100031x
Jakalian A, Jack DB, Bayly CI (2002) Fast, efficient generation of high-quality atomic charges. AM1-BCC model: II, parameterization and validation. J Comput Chem 23:1623–1641. https://doi.org/10.1002/jcc.10128
OpenEye Scientific Software Inc. (2017) QUACPAC 1.6.3
Søndergaard CR, Olsson MHM, Rostkowski M, Jensen JH (2011) Improved treatment of ligands and coupling effects in empirical calculation and rationalization of p K a values. J Chem Theory Comput 7:2284–2295. https://doi.org/10.1021/ct200133y
Banks JL, Beard HS, Cao Y et al (2005) Integrated modeling program, applied chemical theory (IMPACT). J Comput Chem 26:1752–1780. https://doi.org/10.1002/jcc.20292
McGann M (2011) FRED pose prediction and virtual screening accuracy. J Chem Inf Model 51:578–596. https://doi.org/10.1021/ci100436p
McGann M (2012) FRED and HYBRID docking performance on standardized datasets. J Comput Aided Mol Des 26:897–906. https://doi.org/10.1007/s10822-012-9584-8
(2017) OEDocking v.3.2.0: OpenEye Scientific Software, Santa Fe, NM, USA. http://www.eyesopen.com
McGann MR, Almond HR, Nicholls A et al (2003) Gaussian docking functions. Biopolymers 68:76–90
Braga RC, Alves VM, Silva MFB et al (2014) Tuning HERG out: antitarget QSAR models for drug development. Curr Top Med Chem 14:1399–1415. https://doi.org/10.2174/1568026614666140506124442
Braga RC, Alves VM, Silva MFB et al (2015) Pred-hERG: a novel web-accessible computational tool for predicting cardiac toxicity. Mol Inform 34:698–701. https://doi.org/10.1002/minf.201500040
Cheng F, Li W, Zhou Y et al (2012) admetSAR: a comprehensive source and free tool for assessment of chemical ADMET properties. J Chem Inf Model 52:3099–3105. https://doi.org/10.1021/ci300367a
Shen J, Cheng F, Xu Y et al (2010) Estimation of ADME properties with substructure pattern recognition. J Chem Inf Model 50:1034–1041. https://doi.org/10.1021/ci100104j
Cheng F, Yu Y, Shen J et al (2011) Classification of cytochrome P450 inhibitors and noninhibitors using combined classifiers. J Chem Inf Model 51:996–1011. https://doi.org/10.1021/ci200028n
Hansen K, Mika S, Schroeter T et al (2009) Benchmark data set for in silico prediction of ames mutagenicity. J Chem Inf Model 49:2077–2081. https://doi.org/10.1021/ci900161g
Lagunin A, Filimonov D, Zakharov A et al (2009) Computer-aided prediction of rodent carcinogenicity by PASS and CISOC-PSCT. QSAR Comb Sci 28:806–810. https://doi.org/10.1002/qsar.200860192
Baell JB, Holloway GA (2010) New substructure filters for removal of pan assay interference compounds (PAINS) from screening libraries and for their exclusion in bioassays. J Med Chem 53:2719–2740. https://doi.org/10.1021/jm901137j
Baell J, Walters MA (2014) Chemistry: chemical con artists foil drug discovery. Nature 513:481–483
Spackman MA, Jayatilaka D (2009) Hirshfeld surface analysis. CrystEngComm 11:19–32. https://doi.org/10.1039/B818330A
Rognan D (2007) Chemogenomic approaches to rational drug design. Br J Pharmacol 152:38–52
Klabunde T (2007) Chemogenomic approaches to drug discovery: similar receptors bind similar ligands. Br J Pharmacol 152:5–7
Westermaier Y, Barril X, Scapozza L (2015) Virtual screening: an in silico tool for interlacing the chemical universe with the proteome. Methods 71:44–57. https://doi.org/10.1016/j.ymeth.2014.08.001
Buchman CD, Hurley TD (2017) Inhibition of the aldehyde dehydrogenase 1/2 family by Psoralen and Coumarin derivatives. J Med Chem 60:2439–2455. https://doi.org/10.1021/acs.jmedchem.6b01825
Gaulton A, Bellis LJ, Bento AP et al (2012) ChEMBL: a large-scale bioactivity database for drug discovery. Nucl Acids Res 40:D1100–D1107. https://doi.org/10.1093/nar/gkr777
Tomita H, Tanaka K, Tanaka T, Hara A (2016) Aldehyde dehydrogenase 1A1 in stem cells and cancer. Oncotarget 7:11018–11032. https://doi.org/10.18632/oncotarget.6920
van de Waterbeemd H, Gifford E (2003) ADMET in silico modelling: towards prediction paradise? Nat Rev Drug Discov 2:192–204. https://doi.org/10.1038/nrd1032
Alqahtani S (2017) In silico ADME-Tox modeling: progress and prospects. Expert Opin Drug Metab Toxicol 13:1147–1158. https://doi.org/10.1080/17425255.2017.1389897
Acknowledgements
The authors would like to acknowledge the Brazilian agencies Fundação de Amparo à Pesquisa do Estado de Goiás (FAPEG), Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), and Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) for financial support. We would like to thank the Universidade Federal de Juiz de Fora for the X-ray diffraction data collection, OpenEye Scientific Software Inc. (https://www.eyesopen.com/), and ChemAxon (https://chemaxon.com) for providing us with an academic license of their software. CHA and HBN are the researcher’s fellows of CNPq.
Author information
Authors and Affiliations
Contributions
WFV and HBN designed the project. JMFC, GDCO, and CNP performed the synthesis, crystallization, and spectroscopies studies of M-CNP. WFV, HBN, and PSCJ performed crystallography studies. JTMF, BJN, and CHA performed the in silico studies. EPSL performed the in vitro studies. All authors have written, critically reviewed, and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Vaz, W.F., Custodio, J.M.F., D’Oliveira, G.D.C. et al. Dihydroquinoline derivative as a potential anticancer agent: synthesis, crystal structure, and molecular modeling studies. Mol Divers 25, 55–66 (2021). https://doi.org/10.1007/s11030-019-10024-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11030-019-10024-x