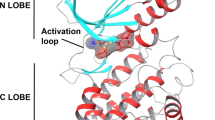
Abstract
High expression of Nek2 has been detected in several types of cancer and it represents a novel target for human cancer. In the current study, structure-based pharmacophore modeling combined with multiple linear regression (MLR)-based QSAR analyses was applied to disclose the structural requirements for NEK2 inhibition. Generated pharmacophoric models were initially validated with receiver operating characteristic (ROC) curve, and optimum models were subsequently implemented in QSAR modeling with other physiochemical descriptors. QSAR-selected models were implied as 3D search filters to mine the National Cancer Institute (NCI) database for novel NEK2 inhibitors, whereas the associated QSAR model prioritized the bioactivities of captured hits for in vitro evaluation. Experimental validation identified several potent NEK2 inhibitors of novel structural scaffolds. The most potent captured hit exhibited an \(\hbox {IC}_{50}\) value of 237 nM.








Similar content being viewed by others
Abbreviations
- AML:
-
Acute myeloid leukemia
- GFA:
-
Genetic function algorithm
- MLR:
-
Multiple linear regression
- NCI:
-
National Cancer Institute
- NEK2:
-
Never in mitosis (NIMA)-related kinase 2
- NIMA:
-
Never in mitosis
- ROC:
-
Receiver operating characteristic
References
O’Regan L, Blot J, Fry AM (2007) Mitotic regulation by NIMA-related kinases. Cell Div 2:25. doi:10.1186/1747-1028-2-25
Fry AM, Meraldi P, Nigg EA (1998) A centrosomal function for the human Nek2 protein kinase, a member of the NIMA family of cell cycle regulators. EMBO J 17:470–481. doi:10.1093/emboj/17.2.470
Bahe S, Stierhof YD, Wilkinson CJ, Leiss F, Nigg EA (2005) Rootletin forms centriole-associated filaments and functions in centrosome cohesion. J Cell Biol 171:27–33. doi:10.1083/jcb.200504107
Chen Y, Riley DJ, Zheng L, Chen PL, Lee WH (2002) Phosphorylation of the mitotic regulator protein Hec1 by Nek2 kinase is essential for faithful chromosome segregation. J Biol Chem 277:49408–49416. doi:10.1074/jbc.M207069200
Di Agostino S, Fedele M, Chieffi P, Fusco A, Rossi P, Geremia R, Sette C (2004) Phosphorylation of high-mobility group protein A2 by Nek2 kinase during the first meiotic division in mouse spermatocytes. Mol Biol Cell 15:1224–1232. doi:10.1091/mbc.E03-09-0638
Xia J, Franqui Machin R, Gu Z, Zhan F (2015) Role of NEK2A in human cancer and its therapeutic potentials. Biomed Res Int 2015:862461. doi:10.1155/2015/862461
Kokuryo T, Senga T, Yokoyama Y, Nagino M, Nimura Y, Hamaguchi M (2007) Nek2 as an effective target for inhibition of tumorigenic growth and peritoneal dissemination of cholangiocarcinoma. Cancer Res 67:9637–9642. doi:10.1158/0008-5472.CAN-07-1489
Tsunoda N, Kokuryo T, Oda K, Senga T, Yokoyama Y, Nagino M, Nimura Y, Hamaguchi M (2009) Nek2 as a novel molecular target for the treatment of breast carcinoma. Cancer Sci 100:111–116. doi:10.1111/j.1349-7006.2008.01007.x
Fletcher L, Cerniglia GJ, Nigg EA, Yend TJ, Muschel RJ (2004) Inhibition of centrosome separation after DNA damage: a role for Nek2. Radiat Res 162:128–135
Suzuki K, Kokuryo T, Senga T, Yokoyama Y, Nagino M, Hamaguchi M (2010) Novel combination treatment for colorectal cancer using Nek2 siRNA and cisplatin. Cancer Sci 101:1163–1169. doi:10.1111/j.1349-7006.2010.01504.x
Whelligan DK, Solanki S, Taylor D, Thomson DW, Cheung KM, Boxall K, Mas-Droux C, Barillari C, Burns S, Grummitt CG, Collins I, van Montfort RL, Aherne GW, Bayliss R, Hoelder S (2010) Aminopyrazine inhibitors binding to an unusual inactive conformation of the mitotic kinase Nek2: SAR and structural characterization. J Med Chem 53:7682–7698. doi:10.1021/jm1008727
Solanki S, Innocenti P, Mas-Droux C, Boxall K, Barillari C, van Montfort RL, Aherne GW, Bayliss R, Hoelder S (2011) Benzimidazole inhibitors induce a DFG-out conformation of never in mitosis gene A-related kinase 2 (Nek2) without binding to the back pocket and reveal a nonlinear structure–activity relationship. J Med Chem 54:1626–1639. doi:10.1021/jm1011726
Innocenti P, Cheung KM, Solanki S, Mas-Droux C, Rowan F, Yeoh S, Boxall K, Westlake M, Pickard L, Hardy T, Baxter JE, Aherne GW, Bayliss R, Fry AM, Hoelder S (2012) Design of potent and selective hybrid inhibitors of the mitotic kinase Nek2: structure–activity relationship, structural biology, and cellular activity. J Med Chem 55:3228–3241. doi:10.1021/jm201683b
Khanfar MA, Taha MO (2013) Elaborate ligand-based modeling coupled with multiple linear regression and k nearest neighbor QSAR analyses unveiled new nanomolar mTOR inhibitors. J Chem Inf Model 53:2587–2612. doi:10.1021/ci4003798
ChemDraw Professional 15.0 for Windows, PerkinElmer Informatics, Waltham Massachusetts USA. http://www.cambridgesoft.com
Discovery Studio 2.5 for Windows, Dassault Systèmes BIOVIA, Discovery Studio Modeling Environment, Release 2009, San Diego: Dassault Systèmes. http://www.accelrys.com/products/collaborative-science/biovia-discovery-studio
Verdonk ML, Berdini V, Hartshorn MJ, Mooij WT, Murray CW, Taylor RD, Watson P (2004) Virtual screening using protein-ligand docking: avoiding artificial enrichment. J Chem Inf Comput Sci 44:793–806. doi:10.1021/ci034289q
Rogers D, Hopfinger AJ (1994) Application of genetic function approximation to quantitative structure–activity relationships and quantitative structure-property relationships. J Chem Inf Comput Sci 34:854–866. doi:10.1021/Ci00020a020
Nonlinear Regression was performed using GraphPad Prism version 5.01 for Windows, GraphPad Software, San Diego California USA. http://www.graphpad.com
Irwin JJ, Sterling T, Mysinger MM, Bolstad ES, Coleman RG (2012) ZINC: a free tool to discover chemistry for biology. J Chem Inf Model 52:1757–1768. doi:10.1021/ci3001277
Gaulton A, Bellis LJ, Bento AP, Chambers J, Davies M, Hersey A, Light Y, McGlinchey S, Michalovich D, Al-Lazikani B, Overington JP (2012) ChEMBL: a large-scale bioactivity database for drug discovery. Nucl Acids Res 40:D1100–1107. doi:10.1093/nar/gkr777
Roy K, Mitra I, Kar S, Ojha PK, Das RN, Kabir H (2012) Comparative studies on some metrics for external validation of QSPR models. J Chem Inf Model 52:396–408. doi:10.1021/ci200520g
Ojha PK, Mitra I, Das RN, Roy K (2011) Further exploring rm2 metrics for validation of QSPR models dataset. Chemom Intell Lab Syst 107:194–205. doi:10.1016/j.chemolab.2011.03.011
Veber DF, Johnson SR, Cheng HY, Smith BR, Ward KW, Kopple KD (2002) Molecular properties that influence the oral bioavailability of drug candidates. J Med Chem 45:2615–2623. doi:10.1021/jm020017n
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (2001) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 46:3–26. doi:10.1016/S0169-409X(00)00129-0
Shoichet BK (2006) Interpreting steep dose–response curves in early inhibitor discovery. J Med Chem 49:7274–7277. doi:10.1021/jm061103g
Acknowledgments
The authors thank the Deanship of Scientific Research at the University of Jordan for their generous funds. We are also thankful for NCI institution for supporting us with free samples. We are thankful for Prof. Mutasem Taha at Faculty of Pharmacy, University of Jordan for fruitful discussions and insight ideas.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
11030_2016_9696_MOESM1_ESM.docx
Supplementary Materials The detailed theoretical and experimental procedures of pharmacophoric and QSAR modeling, and analytical data of active hits discovered in this study. (doc 715 KB)
Rights and permissions
About this article
Cite this article
Khanfar, M.A., Banat, F., Alabed, S. et al. Discovery of potent NEK2 inhibitors as potential anticancer agents using structure-based exploration of NEK2 pharmacophoric space coupled with QSAR analyses. Mol Divers 21, 187–200 (2017). https://doi.org/10.1007/s11030-016-9696-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11030-016-9696-5