Abstract
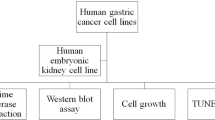
The upregulation or mutation of C-MYC has been observed in gastric, colon, breast, and lung tumors and in Burkitt’s lymphoma. However, little is known about the role C-MYC plays in gastric adenocarcinoma. In the present study, we intended to investigate the influence of C-MYC on the growth, proliferation, apoptosis, invasion, and cell cycle of the gastric cancer cell line SGC7901 and the gastric cell line HFE145. C-MYC cDNA was subcloned into a constitutive vector PCDNA3.1 followed by transfection in normal gastric cell line HFE145 by using liposome. Then stable transfectants were selected and appraised. Specific inhibition of C-MYC was achieved using a vector-based siRNA system which was transfected in gastric cancer cell line SGC7901. The apoptosis and cell cycles of these clones were analyzed by using flow cytometric assay. The growth and proliferation were analyzed by cell growth curves and colony-forming assay, respectively. The invasion of these clones was analyzed by using cell migration assay. The C-MYC stable expression clones (HFE-Myc) and C-MYC RNAi cells (SGC-MR) were detected and compared with their control groups, respectively. HFE-Myc grew faster than HFE145 and HFE-PC (HFE145 transfected with PCDNA3.1 vector). SGC-MR1, 2 grew slower than SGC7901 and SGC-MS1, 2 (SGC7901 transfected with scrambled control duplexes). The cell counts of HFE-Myc in the third, fourth, fifth, sixth, and seventh days were significantly more than those of control groups (P < 0.05). Those of SGC-MR1, 2 in the fourth, fifth, sixth, and seventh days were significantly fewer than those of control groups (P < 0.05). Cell cycle analysis showed that proportions of HFE-Myc and SGC-MR cells in G0–G1 and G2–M were different significantly with their control groups, respectively (P < 0.05). The apoptosis rate of HFE-Myc was significantly higher than those of control groups (P < 0.05). Results of colony-forming assay showed that the colony formation rate of HFE-Myc was higher than those of control groups; otherwise, the rate of SGC-MR was lower than those of their control groups (P < 0.05). The results of cell migration assay showed that there were no significant differences between experimental groups and control groups (P > 0.05). In conclusion, C-MYC can promote the growth and proliferation of normal gastric cells, and knockdown of C-MYC can restrain the growth and proliferation of gastric cancer cells. It can induce cell apoptosis and help tumor cell maintain malignant phenotype. But it can have not a detectable influence on the ability of invasion of gastric cancer cells.










Similar content being viewed by others
References
Parkin DM, Poisani P, Ferlay (1999) Global cancer statistics. CA Cancer J Clin 49(1):33–64
Correa P (1992) Human gastric carcinogenesis: a multistep and multifactorial process—First American Cancer society award lecture on cancer epidemiology and prevention. Cancer Res 52(24):6735–6740
Gonzalez CA, Sala N, Capella G (2002) Genetic susceptibility and gastric cancer risk. Int J Cancer 100(3):249–260
Xue FB, Xu YY, Wan Y et al (2001) Association of H. pylori infection with gastric carcinoma: a meta analysis. World J Gastroenterol 7(6):801–804
Nesbit CE, Tersak JM, Prochownik EV (1999) MYC oncogenes and human neoplastic disease. Oncogene 18(19):3004–3016
Dang CV (1999) C-MYC target genes involved in cell growth, apoptosis, an metabolism. Mol Cell Biol 19(1):1–11
Cole MD, McMahon SB (1999) The Myc oncoprotein: a critical evaluation of transactivation and target gene regulation. Oncogene 18(19):2916–2924
Obaya AJ, Mateyak MK, Sedivy JM (1999) Mysterious liaisons: the relationship between C-MYC and the cell cycle. Oncogene 18(19):2934–2941
Hoffman B, Liebermann DA (1998) The proto-oncogene c-myc and apoptosis. Oncogene 17(25):3351–3357
Prendergast GC (1999) Mechanisms of apoptosis by C-Myc. Oncogene 18(19):2967–2987
Dang CV, Resar LM, Emison E et al (1999) Function of the C-MYC oncogenic transcription factor. Exp Cell Res 253(1):63–77
Milne AN, Sitarz R, Carvalho R et al (2007) Early onset gastric cancer: on the road to unraveling gastric carcinogenesis. Curr Mol Med 7(1):15–28
Calcagno DQ, Leal MF, Seabra AD et al (2006) Interrelationship between chromosome 8 aneuploidy, C-MYC amplification and increased expression in individuals from northern Brazil with gastric adenocarcinoma. World J Gastroenterol 12(38):6207–6211
Kozma L, Kiss I, Hajdu J et al (2001) C-myc amplification and cluster analysis in human gastric carcinoma. Anticancer Res 21(1B):707–710
Yang GF, Deng CS, Xiong YY et al (2004) Expression of nuclear factor-kappa B and target genes in gastric precancerous lesions and adenocarcinoma: association with Helicobactor pylori cagA (+) infection. World J Gastroenterol 10(4):491–496
Onoda N, Maeda K, Chung YS et al (1996) Overexpression of c-myc messenger RNA in primary and metastatic lesions of carcinoma of the stomach. J Am Coll Surg 182(1):55–59
Han S, Kim HY, Park K et al (1999) c-Myc expression is related with cell proliferation and associated with poor clinical outcome in human gastric cancer. J Korean Med Sci 14(5):526–530
Nakata B, Onoda N, Chung YS et al (1995) Correlation between malignancy of gastric cancer and c-myc DNA amplification or overexpression of c-myc protein. Gan To Kagaku Ryoho 22(Suppl 2):176–179
Sanz-Ortega J, Steinberg SM, Moro E et al (2000) Comparative study of tumor angiogenesis and immunohistochemistry for p53, c-ErbB2, c-myc and EGFr as prognostic factors in gastric cancer. Histol Histopathol 15(2):455–462
Ishii HH, Gobe GC, Pan W et al (2002) Apoptosis and cell proliferation in the development of gastric carcinomas: associations with c-myc and p53 protein expression. J Gastroenterol Hepatol 17(9):966–972
Costa Raiol LC, Figueira Silva EC, Mendes da Fonseca D et al (2008) Interrelationship between MYC gene numerical aberrations and protein expression in individuals from northern Brazil with early gastric adenocarcinoma. Cancer Genet Cytogenet 181(1):31–35
Xu AG, Li SG, Liu JH et al (2001) Function of apoptosis and expression of the proteins Bcl-2, p53 and C-myc in the development of gastric cancer. World J Gastroenterol 7(3):403–406
Lan J, Xiong YY, Lin YX et al (2003) Helicobacter pylori infection generated gastric cancer through p53-Rb tumor-suppressor system mutation and telomerase reactivation. World J Gastroenterol 9(1):54–58
Hashimoto K, Nakagawa Y, Morikawa H et al (2001) Co-overexpression of DEAD box protein rck/p54 and c-myc protein in human colorectal adenomas and the relevance of their expression in cultured cell lines. Carcinogenesis 22(12):1965–1970
Henriksson M, Luscher B (1996) Proteins of the Myc network: essential regulators of cell growth and differentiation. Adv Cancer Res 68:109–182
Oster SK, Ho CS, Soucie EL et al (2002) The myc oncogene: MarvelouslY Complex. Adv Cancer Res 84:81–154
Evan GI, Wyllie AH, Gilbert CS et al (1992) Induction of apoptosis in fibroblasts by c-myc protein. Cell 69(1):119–128
Langlois NE, Lamb J, Eremin O et al (1997) Apoptosis in colorectal carcinoma occurring in patients aged 45 years and under: relationship to prognosis, mitosis, and immunohistochemical demonstration of p53, c-myc and bcl-2 protein products. J Pathol 182(4):392–397
Xiangming C, Natsugoe S, Takao S et al (2001) Preserved Smad4 expression in the transforming growth factor beta signaling pathway is a favorable prognostic factor in patients with advanced gastric cancer. Clin Cancer Res 7(2):277–282
Vaux DL, Cory S, Adams JM (1988) Bcl-2 gene promotes haemopoietic cell survival and cooperates with c-myc to immortalize pre-B cells. Nature 335(6189):440–442
Fernandez PC, Frank SR, Wang L et al (2003) Genomic targets of the human c-Myc protein. Genes Dev 17(9):1115–1129
Robinson K, Asawachaicharn N, Denise A (2009) c-Myc accelerates S-phase and requires WRN to avoid replication stress. PLoS One 4(6):e5951
Bouchard C, Dittrich O, Kiermaier A et al (2001) Regulation of cyclin D2 gene expression by the Myc/Max/Mad network: Myc-dependent TRRAP recruitment and histone acetylation at the cyclin D2 promoter. Genes Dev 15(16):2042–2047
Menssen A, Hermeking H (2002) Characterization of the c-MYC-regulated transcriptome by SAGE: identification and analysis of c-MYC target genes. Proc Natl Acad Sci USA 99(9):6274–6279
Hermeking H, Rago C, Schuhmacher M et al (2000) Identification of CDK4 as a target of c-MYC. Proc Natl Acad Sci USA 97(5):2229–2234
Orian A, van Steensel B, Delrow J et al (2003) Genomic binding by the Drosophila Myc, Max, Mad/Mnt transcription factor network. Genes Dev 17(9):1101–1114
Miliani de Marval PL, Macias E, Rounbehler R et al (2004) Lack of cyclin-dependent kinase 4 inhibits c-myc tumorigenic activities in epithelial tissues. Mol Cell Biol 24(17):7538–7547
Kozar K, Ciemerych MA, Rebel VI et al (2004) Mouse development and cell proliferation in the absence of d-cyclins. Cell 118(4):477–491
Wu S, Cetinkaya C, Munoz-Alonso MJ, von der Lehr N et al (2003) Myc represses differentiation -induced p21CIP1 expression via Miz-1-dependent interaction with the p21 core promoter. Oncogene 22(3):351–360
Seoane J, Pouponnot C, Staller P et al (2001) TGFbeta influences Myc, Miz-1 and Smad to control the CDK inhibitor p15INK4b. Nat Cell Biol 3(4):400–408
Claassen GF, Hann SR (2000) A role for transcriptional repression of p21CIP1 by c-Myc in overcoming transforming growth factor beta induced cell-cycle arrest. Proc Natl Acad Sci USA 97(17):9498–9503
Peltenburg LT, de Bruin EC, Meersma D et al (2004) c-Myc is able to sensitize human melanoma cells to diverse apoptotic triggers. Melanoma Res 14(1):3–12
Pelengaris S, Khan M, Evan G (2002) c-MYC: more than just a matter of life and death. Nat Rev Cancer 2(10):764–776
Nilsson JA, Cleveland JL (2003) Myc pathways provoking cell suicide and cancer. Oncogene 22(56):9007–9021
Yang BS, Geddes TJ, Pogulis RJ et al (1991) Transcriptional suppression of cellular gene expression by c-Myc. Mol Cell Biol 11(4):2291–2295
Yang BS, Gilbert JD, Freytag SO et al (1993) Overexpression of Myc suppresses CCAAT transcription factor/nuclear factor 1-dependent promoters in vivo. Mol Cell Biol 13(5):3093–3102
Shiio Y, Donohoe S, Yi EC et al (2002) Quantitative proteomic analysis of Myc oncoprotein function. EMBO J 21(19):5088–5096
Geisler JP, Geisler HE, Manahan KJ et al (2004) Nuclear and cytoplasmic c-myc staining in endometrial carcinoma and their relationship to survival. Int J Gynecol Cancer 14(1):133–137
Ruzinova MB, Caron T, Rodig SJ (2010) Altered Subcellular Localization of c-Myc Protein Identifies Aggressive B-cell Lymphomas Harboring a c-MYC Translocation. Am J Surg Pathol 34(6):882–891
Acknowledgments
The authors wish to thank Drs. Haili Huang and Gangshi Wang, and Nurse Weidi You, Weihua Wang et al., for handling patient contacts. We wish to thank the Forth Military Medical University of PLA for providing means for the current investigation.
Conflict of interest statement
The authors declare no competing interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhang, L., Hou, Y., Ashktorab, H. et al. The impact of C-MYC gene expression on gastric cancer cell. Mol Cell Biochem 344, 125–135 (2010). https://doi.org/10.1007/s11010-010-0536-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-010-0536-0