Abstract
Initial phase of COVID-19 infection is associated with the binding of viral spike protein S1 receptor binding domain (RBD) with the host cell surface receptor, ACE2. Peptide inhibitors typically interact with spike proteins in order to block its interaction with ACE2, and this knowledge would promote the use of such peptides as therapeutic scaffolds. The present study examined the competitive inhibitor activity of a broad spectrum antimicrobial peptide, Dermaseptin-S4 (S4) and its analogues. Three structural S4 analogues viz., S4 (K4), S4 (K20) and S4 (K4K20) were modelled by substituting charged lysine for non-polar residues in S4 and subsequently, docked with S1. Further, the comparative analysis of inter-residue contacts and non-covalent intermolecular interactions among S1–S4 (K4), S1–S4 (K4K20) and S1–ACE2 complexes were carried out to explore their mode of binding with S1. Interestingly, S1–S4 (K4) established more inter-molecular interactions compared to S4 (K4K20) and S1–ACE2. In order to substantiate this study, the normal mode analysis (NMA) was conducted to show how the structural stability of the flexible loop region in S1 is affected by atomic displacements in unbound S1 and docked complexes. Markedly, the strong interactions consistently maintained by S1–S4 (K4) complex revealed their conformational transition over the harmonic motion period. Moreover, S1–S4 (K4) peptide complex showed a higher energy deformation profile compared to S1–S4 (K4K20), where the higher energy deformation suggests the rigidity of the docked complex and thus it’s harder deformability, which is also substantiated by molecular dynamics simulation. In conclusion, S1–S4 (K4) complex has definitely exhibited a functionally significant dynamics compared to S1–ACE2 complex; this peptide inhibitor, S4 (K4) will need to be considered as the best therapeutic scaffold to block SARS-CoV-2 infection.
Similar content being viewed by others
Introduction
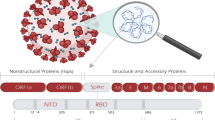
Vaccine discovery for current pandemic Coronavirus disease 2019 (COVID-19) persist as one of the major challenges in the medical arena. More than 9.2 million confirmed deaths and 28 million active cases as of 14th September 2020, since the outbreak at Wuhan, China from December 2019 have been reported by World Health Organisation. Apart from casualties, billions of people worldwide have lost their livelihood due to quarantine and isolations. On the grounds, the causative agent of this novel beta-coronavirus COVID-19 is found to be severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2). As like SARS-CoV, SARS-CoV-2 also contains type I transmembrane glycoprotein called Spike protein (S) consisting of S1 and S2 subunits, where S1 holds a receptor binding domain (RBD) between 334th and 528th position (comprising a total of 194 amino acids). Notably, there exists a loop region comprising 56 residues (from 446th position to 502th position) between β1 and β2 strands of RBD, the most flexible region imparting several interactions with binding partners (Lan et al. 2020; Wang et al. 2020; Wrapp et al. 2020; Wu et al. 2020). And that loop region is found to bind ACE2 receptor, which is the crucial step of viral infection (Li 2015; Shang et al. 2020; Yan et al. 2020). Therefore, S1 emerge as an attractive drug target of SARS-CoV-2, for which, the development of small molecule inhibitors (Abdelmageed et al. 2020; Barh et al. 2020; Xia et al. 2020) and the peptide based drugs (Baig et al. 2020; Elnagdy and AlKhazindar 2020; Maiti 2020) are in progress. Recently, the 3D structural complex of S1 and the neutralising convalescent antibody B38 have been identified, which anticipates the rational design of vaccines (Wu et al. 2020).
Likewise, the alpha helical anti-microbial peptides (AMPs) are indeed the effective alternate therapeutic option for combating SARS-CoV-2 virus. AMPs are generally amphipathic in nature which have a broad spectrum of activities against bacteria, fungi, viruses and protozoans. Similar studies have reported that modifying AMPs with positively charged lysine might increase the cationicity and suppress the enveloped viruses like HIV-1 (Hancock and Diamond 2000). One such potential AMPs, Dermaseptin-S4 (S4) have demonstrated a wide antiviral activity against Human immunodeficiency virus1 [HIV-1] (Lorin et al. 2005), Herpes simplex virus-1 [HSV-1] (Belaid et al. 2002), Herpes simplex virus-2 [HSV-2] (Bergaoui et al. 2013) and Rabies virus [RV] (Mechlia et al. 2019). S4 extracted from the skin secretion of South American tree frogs, Phyllomedusa sauvagei (Mor and Nicolas 1994), composed of 27 amino acids acquires alpha-helical conformation when interacting with membrane lipid layers. When modified with lysine residues at 4th, 20th and at both, 4th and 20th positions of S4 generated the analogues viz., S4 (K4), S4 (K20) and S4 (K4K20) of enhanced antiviral activity and low haemolytic activity (Krugliak et al. 2000; Lorin et al. 2005; Bergaoui et al. 2013). As reported earlier, mutating the native S4 with lysine residues improves the net positive charge, thereby increasing the antimicrobial activity and decreasing the haemolytic activity of S4 analogues (Kustanovich et al. 2002; Lorin et al. 2005; Shalev et al. 2006; Zaïri et al. 2014). The presence of positively charged residues will not disrupt the zwitterion lipids in the eukaryotic membrane (Jiang et al. 2008), which elucidates the role of cationic residues in Dermaseptin for preserving low haemolytic activity and high antimicrobial activity. Moreover, the peptide-protein interaction studies of AMPs targeting Middle East Respiratory Syndrome (MERS) coronavirus spike protein established Dermaseptin-S4 and Dermaseptin-S9 as best docked AMPs with high binding affinity (Mustafa et al. 2019). With these perspectives, we intended to study the efficacy of S4 and its analogues mutated with lysine residues, as a possible competitive inhibitor for ACE2 to prevent the binding of S1 to ACE2, and thereby arresting the SARS-CoV-2 infection.
Materials and Methods
Modelling the Dermaseptin S4 and Its Analogues
Using the sequence of Dermaseptin (S4) (ID: P80280) from UniProtKB database and the 3D structural template (PDB ID: 2DD6), the 3D structure of S4 and its analogues viz., S4 (K4), S4 (K20) and S4 (K4K20) were modelled, using PEP-FOLD3 program. PEP-FOLD3 is a de-novo approach aimed at predicting peptide structures from amino acid sequences. This method is based on the structural alphabet (SA) letters to describe the conformations of four consecutive residues. It couples the predicted series of SA letters to a greedy algorithm and a coarse-grained force field which is expressed as a sum of local, non-bonded, and hydrogen bond (Shen et al. 2014). Specifically, this method provides the profile which exhibits the structural alphabet (SA) probability values corresponding to the residue position, determined on the basis of the Hidden Markov Model (HMM). All the models were energy minimized, using YASARA package (Krieger and Vriend 2014). The steepest descent energy minimization of peptides was carried out, using YASARA2 force field with 10.5 Å cut off and the peptides were geometrically optimized, using explicit solvent. The experiment was converged as soon as the energy improves by < 0.05 kJ/mol per atom during 200 steps. Energy minimized models were validated for steric stability through Ramachandran plot by Structure Analysis and Verification Server (SAVES) (Laskowski et al. 1993). Further, the secondary structural information of S4 and its analogues were annotated using POLYVIEW-2D (Porollo et al. 2004). Further, the physiochemical properties were predicted using HeliQuest program (Gautier et al. 2008).
Computation of Binding Affinity, Binding Interface and Intermolecular Interactions
To compute the binding affinity among spike glycoprotein (S1) of SARS-CoV-2 with Dermaseptin (S4) and its analogues, the 3D Structure of SARS-CoV-2 RBD of spike glycoprotein (PDB ID: 6M0J:E chain) with a resolution of 2.45Å was retrieved from PDB database. Initially, the blind docking was carried out using HPEPDOCK program which is based on hierarchical algorithm by fast conformational modelling and global sampling of binding orientations (Zhou et al. 2018). To optimize the pose and orientation between protein-peptide complex, the site specific docking was carried out, using ZDOCK version 3.0.2, where the best binding solution of all complexes viz., S1–S4, S1–S4 (K4), S1–S4 (K20), S1–S4 (K4K20) were categorized based on statistical potential, shape complementarity and electrostatics (Pierce et al. 2014). In addition, PDBePISA (Proteins, Interfaces, Structures and Assemblies) tool (Krissinel and Henrick 2005) was used to identify the interface area and the solvation free energy of docked complex, in the form of free energy gain probability value indicated as ΔiG (P value). Moreover, the intermolecular interactions between the docked complex such as hydrophobic interactions, hydrogen bonds, aromatic-aromatic and cation-π interactions were analysed through Protein Interaction Calculator (PIC) program (Tina et al. 2007). In addition, structure-based multiple sequence alignment (MSA) of S1 key binding site of ACE2 receptor protein and S4 and its analogues was performed through UCSF Chimera tool (Pettersen et al. 2004) and visualized via ESPRIPT tool (Robert and Gouet 2014). Finally, all types of interactions were visualized using PyMol visualization tool (Rigsby and Parker 2016).
Analysing the Protein–Peptide Complex by Elastic Network Model
To characterize the intrinsic flexibility and to validate the low frequency vibration modes of S1 and docked complexes, Normal Mode Analysis (NMA) was performed (Tiwari et al. 2014). Most proteins carry out their functions depending upon the conformational changes of their structure at various circumstances (Yang et al. 2009). NMA is based upon Elastic network model (ENM) which uses a single parameter harmonic potential, usually coarse-grained, has been used for studying protein dynamics (Skjaerven et al. 2009). In our present study, we used WEBnm@ v2.0 program which functions on the theory of ENMs by representing a protein as a collection of spring-connected particles and provide the collective motion of the protein (Skjaerven et al. 2009). Using this program, we calculated the deformation energies and normalized square fluctuations of S1 and docked complexes viz., S1–S4, S1–S4 (K4), S1–S4 (K20) and S1–S4 (K4K20), using the spectrum of low frequency vibration modes (Tiwari et al. 2014; Fuglebakk et al. 2015). Moreover, the deformation energy is calculated for each atom in the protein structure. It depends on the changes in the distance between the atom in question and each of its close neighbours. Higher deformation energies indicate relatively rigid regions, whereas lower deformation energies indicate flexible regions. In the present study, the deformation energies of the 50 lowest-frequency non-trivial modes are calculated. Besides, the normalized square fluctuations signify the sum of the atomic displacements in each mode, weighted by the inverse of their corresponding eigenvalues. Also, the square of the fluctuation of each Cα atom was calculated from 200 lowest non-trivial modes and normalized, so that the sum over all residues is equal to 100.
Analysing the Protein–Peptide Complex by Structural Dynamics
In order to further substantiate the stability of complex formation, S1–S4 (K4) and S1–S4 (K4K20) complexes were simulated to a period of 1 × 105 time units via discrete molecular dynamics (DMD) that corresponds towards 25 ns in relating with the classical MD (Ding et al. 2008; Srinivasan and Rajasekaran 2017). DMD is a unique method that uses the atomistic Medusa force field and discrete energy potential modelled with discontinuous functions for pair-wise interaction. DMD simulations were performed for complex systems with constant volume and periodic boundary conditions. In addition, Anderson’s thermostat was used during DMD simulation to maintain a constant temperature (Shirvanyants et al. 2012). From trajectories obtained during the DMD simulation, the different geometrical observables viz., Root Mean Square Deviation (RMSD), Root Mean Square Fluctuation (RMSF) and Radius of Gyration (Rg) that describe the structural variations of S1–S4 (K4) and S1–S4 (K4K20) complexes were analysed via GROMACS tools (Singh et al. 2020; Bhardwaj and Purohit 2020a, b). Further, the RMSD, RMSF and Rg graphs were plotted using Xmgrace tool to visualize the structural differences between the complex trajectories (Bhardwaj et al. 2020a, b; Muralidharan et al. 2020).
Statistical Analysis
Statistical analysis of the Wilcoxon Matched Pair test was performed, using StatPlus software version 7 (AnalystSoft, www.analystsoft.com/en)for the conformational sampling data of NMA. Wilcoxon Matched Pair test is considered to be equivalent to the Student’s t-Test for Matched Pairs, which is used for evaluating any type of normal distribution data (Scheff 2016).This specific test would be a very appropriate non-parametric test for evaluating the statistical significance of deformation energies of S1–S4 (K4) and S1–S4 (K4K20) complexes evaluated in NMA analysis. Similarly, the nonparametric values obtained through DMD were subjected to basic univariate and multivariate data test, using StatPlus program. The same nonparametric test was performed for all the parameters viz., RMSD, RMSF, Rg, conformational free energies and membrane-free energies of peptides. Both S1–S4 (K4) and S1–S4 (K4K20) complexes determine their significant difference and the probability value (P value) less than 0.05 were considered to be statistically significant.
Result and Discussion
Structural Characterization and Validation of Modelled Structures of S4 and Its Analogues
In order to understand the binding effects of S4 and its analogues with S1 spike protein of SARS-CoV-2, the 3D structures were modelled using PEP-FOLD3. The structural profiles predicted by PEP-FOLD3 showed that the majority of the amino acids in both, S4 and its analogues have contributed for the formation of alpha helix as compared to extended sheet and coil in terms of structural alphabet (SA) probability values (Fig. 1). Overall, the modelled structures of S4 and its analogues exhibited 84.61% of alpha helical content and 15.39% of coil content as confirmed from Polyview-2D and further substantiated with their 3D structures using PyMol (Fig. 1).
3D structure prediction and the annotation of a S4, b S4 (K4), c S4 (K20) and d S4 (K4K20) peptides, where the colours- red, green and blue represents helical, extended sheet and coil, respectively. In cartoon representations of peptides, brown, red, orange and pink represents S4, S4 (K4), S4 (K20) and S4 (K4K20) peptides, respectively (Color figure online)
The modelled 3D structures were also validated by Ramachandran plot (Fig. 2), wherein, 100% of amino acids in S4 (K4) resided only in the favoured regions, whereas, in both S4 and S4 (K4K20) 95.8% of amino acids resided in favoured regions and the remaining 4.2% amino acids resided in additionally allowed regions . In the case of S4 (K20), 87.5% of amino acids resided in favoured regions, 8.3% of amino acids resided in additionally allowed region and 4.2% amino acids resided in generously allowed region. Since, none of the amino acids of all the modelled 3D structures resided in the disallowed regions of Ramachandran plot, we claim that the modelled 3D structures are considered to be sterically stable and hence subjected into molecular docking studies, as in silico studies reported the antiviral activity of Dermaseptin through its significant binding effects on MERS-CoV spike protein (Mustafa et al. 2019). For Dermaseptin and its analogues, the physiochemical properties such as hydrophobicity (H) and hydrophobic moment (μH) were measured, using HeliQuest tool to demonstrate their cationic properties. Hydrophobicity influences the interaction between the microbial membrane and the cationic peptide; the hydrophobic moment plays a significant role in interfacial binding of peptide with the membrane (Eisenberg et al. 1982). Hydrophobic moment is defined as the vector sum of hydrophobicity of individual amino acids (Eisenberg et al. 1984; Gautier et al. 2008). The predicted values of Dermaseptin analogues were tabulated corresponding to their sequences (Supplementary Table 1), in which in which S4 (K4) showed the highest range of hydrophobic moment (µH) compared to other peptides.
Rational Assessment of Protein–Peptide Complex Formation
It is well known that the human ACE2 receptor is bound to the loop region (residue position between 446 and 502) of S1-RBD, as revealed from the solved 3D structure of S1–ACE2 complex (PDB ID: 6M0J) (Lan et al. 2020) (Fig. 3a). In this context, the blind docking of S4 and its analogues viz., S4 (K4), S4 (K20) and S4 (K4K20) with S1 spike protein was carried out by HPEPDOCK. The blind docking enables rapid conformational modelling, based on a hierarchical algorithm, in which S4 and its analogue peptides with S1 spike protein binding orientations are sampled, globally. Further, to optimize the precise pose and orientation of S4 and its analogues into the loop region of S1-RBD, ZDOCK program was used to compute dock score of all the complexes. Through site-specific docking, ZDOCK enhances the position and orientation between protein-peptide complexes, where the best binding solution of all complexes were classified based on statistical potential, complementarity of shape and electrostatics. Surprisingly, most of the binding prediction models indicated that the loop region of S1 is the binding site for S4 and its analogues, as that of ACE2 (Fig. 3b, c, d and e). From this analysis, it was found that all the docked complexes obtained the similar ZDOCK score marginally in a range between 1202.35 and 1281.06 (Table 1).
a PDB structure of S1–ACE2 (PDB ID: 6M0J); Representative illustration of blind docking poses for b S1–S4; c S1–S4 (K4); d S1–S4 (K20) and e S1–S4 (K4K20) highlighted with different colours such as brown, red, orange and pink, respectively. Additionally the key interacting residues represented as stick and loop is highlighted in yellow colour (Color figure online)
In addition, to check the binding interface conservation among ACE2 receptor, S4 and its analogues, S1 binding site of ACE2 receptor (PDB ID: 6M0J) comprising of nearly 34 residues, present in the N terminal region was separated (Fig. 4a, b). Further, it is compared with S4 and its analogues by structure-based sequence alignment, where the significant conservation was observed (Fig. 4c). Hence, the significant conservation of binding interface resemblance inferred that S4 and its analogues could likely be a competitive inhibitor for ACE2 receptor.
Besides, the docked complex prominently indicated that S1 and its analogues captured the similar pose and orientation on to the loop region of S1-RBD (Fig. 5).
Site specific docking of a S1–S4, b S1–S4 (K4), c S1–S4 (K20) and d S1–S4 (K4K20) sharing common binding cleft, where S1 is represented in surface model and the peptides, S4, S4 (K4), S4 (K20) and S4 (K4K20) represented as cartoon highlighted with different colours such as brown, red, orange and pink, respectively (Color figure online)
Furthermore, the binding effects of the docked complex were explored in terms of interface area, and the free energy gain probability value (ΔiG (P)) other than docked score. It is well known that, proteins or peptide interacts with other proteins through their interfaces generally accomplishing their functions (May and Zacharias 2005; London et al. 2010). Moreover, the interfaces are formed by residues whose properties determine binding specificity and affinity (Reichen et al. 2014). In addition, the polarity of interfaces increased with decreasing interface size (Ansari and Helms 2005). In this context, it was observed that the docked complex of S1–S4 (K4K20) had lesser interface area followed by S1–K4S4 as compared to other docked complexes (Table 1). Further, ΔiG (P value) was computed for all the docked complexes, where ΔiG measures the probability interface of docked complexes denoting P > 0.5 that the interface is less hydrophobic and the interaction will stay intact (Patil et al. 2010).Thermodynamically desirable parameters are exhibited by ΔiG (P value), such as the possibility of achieving a lower solvation-free energy gain upon complex formation by docking (Sowmya and Ranganathan 2015). Besides, we observed that, all the docked complexes possess ΔiG in the range of 0.334 to 0.543. S1–K4S4 and S1–S4 (K4K20) obtained the value of P > 0.5, expressing that the complex structures were likely to stay intact. Eventually, this directed us to intensively correlate the range of interface area and the binding affinity among the docked complex with respect to their inter-residue contacts (binding residues) by computing the inter-molecular interactions of all docked complexes through PIC program. Of note, this analysis would not only correlate the interface area and binding affinity, but also provide the information on the mode of binding inhibition of S4 and its analogues with S1. Total number of inter-molecular interactions was concerned where; S1–S4 (K4) complex obtained maximum number of interaction as compared to other complexes (Table 2, Supplementary Tables 2, 3 and 4). Notably, the hydrogen bonds are significantly more in S1–S4 (K4), where many studies reported that hydrogen bond was considered to be a major bond to influence the binding strength of protein–protein or protein–peptide interactions (Chen et al. 2016). Moreover, the contribution of more number of interactions of S1–S4 (K4) was due to the significant number and the nature of its binding amino acids (inter-residues) (Table 2).
From Table 2, it was very interesting to observe that, about nine binding amino acids viz., L2, W3, L6, L7, V10, A14, A17, L18 and V21 are commonly found in S4 and its analogues; more specifically, L11 is commonly found in S4, S4 (K4) and S4 (K20) A13 in S4, S4 (K4) and S4 (K4K20) and A27 in S4, S4 (K4) and S4 (K20). The presence of hydrophobic residues such as alanine, valine, and leucine will influence a peptide’s antiviral activity, as several studies claimed that therapeutic potential of an AMP would be great if its hydrophobicity is high (Dathe et al. 1997; Shai and Oren 2001). Besides, residue K4 is commonly found in both S4 (K4) and S4 (K4K20). Ultimately, the binding amino acids viz., A20, K16, M4 and K20 are found to be unique in each of the peptides viz., S4, S4 (K4), S4 (K20) and S4 (K4K20) respectively. From this analysis, we could infer that more positive charges are imposed on S4 (K4) and S4 (K4K20), which accounted for the enhanced and intact binding affinity with S1 spike protein as compared to other peptides. To explore the mode of binding inhibition of S4 (K4) and S4 (K4K20) analogues with S1, a comparative analysis of inter-residue contacts and inter-molecular interactions among S1–ACE2 complex, S1–S4 (K4) and S1–S4 (K4K20) was done. In which, a total of 12 binding amino acids were observed in S1 and about 14 binding amino acids were observed in ACE2 of S1–ACE2 complex (PDB ID:6M0J) (Table 2).
As we already noticed that, a total of 14 binding amino acids in S1 and a total of 13 binding amino acids in S4 (K4), on the other hand, a total of 16 binding amino acids in S1 and a total of 13 binding amino acids in S4 (K4K20) were observed in S1–S4 (K4) and S1–S4 (K4K20) complexes respectively (Table 2). As inter-molecular interactions were concerned, a total of 43 interactions were found in S1–ACE2 complex, 45 inter-molecular interactions in S1–S4 (K4) complex and 36 inter-molecular interactions in S1–S4 (K4K20) complex (Table 3). Hence, it was inferred that both S4 (K4) and S4 (K4K20) were bound with S1 in the similar region as like ACE2, since about 7 binding amino acids of S1 are commonly shared with ACE2, S4 (K4) and S4 (K4K20) (Table 3).
Therefore, both S4 (K4) and S4 (K4K20) could act competitive inhibitors for S1. It is to be noted that, when a cationic side chain is in close proximity with an aromatic side chain, its structural geometry influences and endures a desirable cation-π interaction (Gallivan and Dougherty 1999). Most importantly, cation-π interaction plays a crucial role in AMP activity (Mishra et al. 2018). In accordance, cation-π interactions were observed in both S4 (K4) and S4 (K4K20) with S1 (Table 3) through the cationic amino acid, lysine in both S4–K4 and S4–K4K20 with the aromatic amino acids, tryptophan and phenylalanine in S1 (Fig. 6). Besides, the number of binding amino acids in S1 was significantly more when bound with S4 (K4) and S4 (K4K20) as compared to ACE2 (Table 3). Of note, S1–S4 (K4) complex established more number of inter-molecular interactions as compared to S1–ACE2 complex and S1–S4 (K4K20) (Table 3). Hence our finding anticipated that S4 (K4) could be the best therapeutic scaffold against S1. In order to substantiate this statement, normal mode analysis was performed to exhibit how the structural stability of S1 protein change to affected, by comparing S1–S4 (K4) complex and S1–S4 (K4K20) complex.
Normal Mode Analysis on S1–S4 (K4) and S1–S4 (K4K20) Complexes
NMA provides the information about the native conformational changes of protein in order to infer its function. Upon the binding of ligands, the conformational transition of protein will occur and gets deviated from its native conformations. In practice, this strategy is being taken into account for rational based drug design (Dobbins et al. 2008). In this context, unbound and bound forms of S1 spike protein were subjected into NMA to explore the conformational transition between them. From NMA, the normalized square fluctuations signified the sum of atomic displacements in S1 and docked complexes, where the flexible loop region between β1 and β2 strands (G446 to G502) of S1 significantly reduced its peak in S1–S4 (K4) and S1–S4 (K4K20) complexes as compared to unbound S1, which clearly indicated that the strong interactions are consistently retained by both S4 (K4) and S4 (K4K20) with S1 and also revealed their conformational transition over the period of harmonic motion (Fig. 7).
a Super-imposition of normal modes of S1–S4 (K4) and S1–S4 (K4K20), where the loop region of S1 is highlighted in yellow, S1–S4 (K4) highlighted in red and S1–S4 (K4K20) highlighted in pink. b Normalized square fluctuation of S1 spike protein .Yellow colour shade in the fluctuation plot represents the loop region of S1, red and pink colour shades represent S1–S4 (K4) and S1–S4 (K4K20), respectively (Color figure online)
In a competitive inhibitory manner, it was evident that S4 (K4) and S4 (K4K20) bind within the loop region of S1, thereby blocking the interaction of S1 with ACE2. Further, the stability of docked complexes was validated by its deformation energy. Based on the deformation energies obtained from the normal modes (Table 4), it was observed that S1–S4 (K4) peptide complex had the higher deformation energy profile as compared to S1–S4 (K4K20) (Fig. 8), where the higher deformation energy indicates the rigidity of the docked complex that is hard to deform (Hollup et al. 2005).
On the other hand, lesser deformation energy indicates the less rigidity of peptide complex which easily deform. From this study, it was found that S1–S4 (K4) sustained its stability evidently as it acquired more energy to deform when compared to S1–S4 (K4K20). Furthermore, the statistical analysis determines both the peptides to have significant difference and the probability value (P value) less than 0.05 were considered to be statistically significant.
DMD Analysis on S1–S4 (K4) and S1–S4 (K4K20) Complexes
RMSD is a typical measure of the average distance of any two superimposed sets of coordinates between Cα backbones (Martínez 2015). The calculated RMSD average values of S1–S4 (K4) and S1–S4 (K4K20) complexes ranged between 0.5 and 0.8 nm, respectively (P value < 0.0001) (Fig. 9). As a result, S1–S4 (K4) exhibited the intact RMSD value when compared to S1–S4 (K4K20), which deviate more than 0.8 Å till 25 ns.
On the other hand, the mean RMSF of complexes ranged between 0.2 and 1.32 nm. RMSF is indeed a measurement of the distance between atom clusters relative to the well-defined average position coordinate set (Martínez 2015). The fluctuations of S1–S4 (K4) showed lower average RMSF values compared to S1–S4 (K4K20) complexes (P value < 0.0001) (Fig. 9). Radius of Gyration measures the compactness and stability of peptide complexes through molecular spatial packaging of amino acid residues, an important parameter for stability analysis. It is obvious that S1–S4 (K4) displayed excellent compactness and proved to be a stable peptide complex compared to S1–S4 (K4K20) (Fig. 9).
Conclusion
Modifying Dermaseptin S4 peptides with positively charged lysine has increased cationicity and improved antiviral properties. Spike viral protein S1–Dermaseptin S4 (K4) docked complex holds reduced interface area in blind docking and further, its position and orientation on the S1-RBD loop region was exposed through site-specific docking. Distinctly, S1–S4 (K4) exhibited strong intermolecular interactions; its NMA studies demonstrated the additional perspective essential to explain the strong interfaces in the functional movements of protein-peptide complexes with functionally significant dynamics. Markedly, the S1–S4 (K4) complex was indeed more stable, as it acquires more energy to deform. Further, the MD simulation revealed that the S1–S4 (K4) complex is structurally stable and retains all its interactions. Hence, we anticipate that S4 (K4) might be used as a potential therapeutic peptide scaffold and also, routed towards the designing of peptidomimetics in the treatment of COVID-19 viral infection.
References
Abdelmageed MI et al (2020) Design of a multiepitope-based peptide vaccine against the E protein of human COVID-19: an immunoinformatics approach. Biomed Res Int 2020:1–12. https://doi.org/10.1155/2020/2683286
Ansari S, Helms V (2005) Statistical analysis of predominantly transient protein-protein interfaces. Proteins: Struct Funct Bioinform 61(2):344–355. https://doi.org/10.1002/prot.20593
Baig MS et al (2020) Identification of a potential peptide inhibitor of SARS-CoV-2 targeting its entry into the host cells. Drugs R&D 20(3):161–169. https://doi.org/10.1007/s40268-020-00312-5
Barh D et al (2020) Potential chimeric peptides to block the SARS-CoV-2 spike RBD. Preprint. Life Sci. https://doi.org/10.20944/preprints202004.0347.v1
Belaid A et al (2002) In vitro antiviral activity of dermaseptins against herpes simplex virus type 1. J Med Virol 66(2):229–234. https://doi.org/10.1002/jmv.2134
Bergaoui I et al (2013) In vitro antiviral activity of dermaseptin S4 and derivatives from amphibian skin against herpes simplex virus type 2. J Med Virol 85(2):272–281. https://doi.org/10.1002/jmv.23450
Bhardwaj VK, Purohit R (2020a) A new insight into protein-protein interactions and the effect of conformational alterations in PCNA. Int J Biol Macromol 148:999–1009. https://doi.org/10.1016/j.ijbiomac.2020.01.212
Bhardwaj V, Purohit R (2020b) Computational investigation on effect of mutations in PCNA resulting in structural perturbations and inhibition of mismatch repair pathway. J Biomol Struct Dyn 38(7):1963–1974. https://doi.org/10.1080/07391102.2019.1621210
Bhardwaj VK, Singh R, Sharma J, Rajendran V et al (2020a) Identification of bioactive molecules from tea plant as SARS-CoV-2 main protease inhibitors. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2020.1766572
Bhardwaj VK, Singh R, Sharma J, Das P et al (2020b) Structural based study to identify new potential inhibitors for dual specificity tyrosine-phosphorylation- regulated kinase. Comput Methods Progr Biomed 194:105494. https://doi.org/10.1016/j.cmpb.2020.105494
Chen D et al (2016) Regulation of protein-ligand binding affinity by hydrogen bond pairing. Sci Adv 2(3):e1501240. https://doi.org/10.1126/sciadv.1501240
Dathe M et al (1997) Hydrophobicity, hydrophobic moment and angle subtended by charged residues modulate antibacterial and haemolytic activity of amphipathic helical peptides. FEBS Lett 403(2):208–212. https://doi.org/10.1016/S0014-5793(97)00055-0
Ding F et al (2008) Ab initio folding of proteins with all-atom discrete molecular dynamics. Structure 16(7):1010–1018. https://doi.org/10.1016/j.str.2008.03.013
Dobbins SE, Lesk VI, Sternberg MJE (2008) Insights into protein flexibility: the relationship between normal modes and conformational change upon protein-protein docking. Proc Natl Acad Sci 105(30):10390–10395. https://doi.org/10.1073/pnas.0802496105
Eisenberg D, Weiss RM, Terwilliger TC (1982) The helical hydrophobic moment: a measure of the amphiphilicity of a helix. Nature 299(5881):371–374. https://doi.org/10.1038/299371a0
Eisenberg D, Weiss RM, Terwilliger TC (1984) The hydrophobic moment detects periodicity in protein hydrophobicity. Proc Natl Acad Sci 81(1):140–144. https://doi.org/10.1073/pnas.81.1.140
Elnagdy S, AlKhazindar M (2020) The potential of antimicrobial peptides as an antiviral therapy against COVID-19. ACS Pharmacol Transl Sci 3(4):780–782. https://doi.org/10.1021/acsptsci.0c00059
Fuglebakk E, Tiwari SP, Reuter N (2015) Comparing the intrinsic dynamics of multiple protein structures using elastic network models. Biochim Biophys Acta 1850(5):911–922. https://doi.org/10.1016/j.bbagen.2014.09.021
Gallivan JP, Dougherty DA (1999) Cation-π interactions in structural biology. Natl Acad Sci USA, Proc, p 6
Gautier R et al (2008) HELIQUEST: a web server to screen sequences with specific -helical properties. Bioinformatics 24(18):2101–2102. https://doi.org/10.1093/bioinformatics/btn392
Hancock REW, Diamond G (2000) The role of cationic antimicrobial peptides in innate host defences. Trends Microbiol 8(9):402–410. https://doi.org/10.1016/S0966-842X(00)01823-0
Hollup S, Salensminde G, Reuter N (2005) WEBnm@: a web application for normal mode analyses of proteins. BMC Bioinform 6(1):52. https://doi.org/10.1186/1471-2105-6-52
Jiang Z et al (2008) Effects of net charge and the number of positively charged residues on the biological activity of amphipathic α-helical cationic antimicrobial peptides. Biopolymers 90(3):369–383. https://doi.org/10.1002/bip.20911
Krieger E, Vriend G (2014) YASARA view—molecular graphics for all devices—from smartphones to workstations. Bioinformatics 30(20):2981–2982. https://doi.org/10.1093/bioinformatics/btu426
Krissinel E, Henrick K (2005) Detection of protein assemblies in crystals. In: Berthold RM et al (eds) Computational life sciences. Lecture Notes in Computer Science. Springer, Berlin, pp 163–174
Krugliak M et al (2000) Antimalarial activities of dermaseptin S4 derivatives. Antimicrob Agents Chemother 44(9):2442–2451. https://doi.org/10.1128/AAC.44.9.2442-2451.2000
Kustanovich I et al (2002) Structural requirements for potent versus selective cytotoxicity for antimicrobial dermaseptin S4 derivatives. J Biol Chem 277(19):16941–16951. https://doi.org/10.1074/jbc.M111071200
Lan J et al (2020) Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 581(7807):215–220. https://doi.org/10.1038/s41586-020-2180-5
Laskowski RA et al (1993) PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Crystallogr 26(2):283–291. https://doi.org/10.1107/S0021889892009944
Li F (2015) Receptor recognition mechanisms of coronaviruses: a decade of structural studies. J Virol 89(4):1954–1964. https://doi.org/10.1128/jvi.02615-14
London N, Movshovitz-Attias D, Schueler-Furman O (2010) The structural basis of peptide-protein binding strategies. Structure 18(2):188–199. https://doi.org/10.1016/j.str.2009.11.012
Lorin C et al (2005) The antimicrobial peptide dermaseptin S4 inhibits HIV-1 infectivity in vitro. Virology 334(2):264–275. https://doi.org/10.1016/j.virol.2005.02.002
Maiti BK (2020) Potential role of peptide-based antiviral therapy against SARS-CoV-2 infection. ACS Pharmacol Transl Sci 3(4):783–785. https://doi.org/10.1021/acsptsci.0c00081
Martínez L (2015) Automatic identification of mobile and rigid substructures in molecular dynamics simulations and fractional structural fluctuation analysis. PLoS ONE. https://doi.org/10.1371/journal.pone.0119264
May A, Zacharias M (2005) Accounting for global protein deformability during protein–protein and protein–ligand docking. Biochim Biophys Acta 1754(1–2):225–231. https://doi.org/10.1016/j.bbapap.2005.07.045
Mechlia MB et al (2019) Dermaseptins as potential antirabies compounds. Vaccine 37(33):4694–4700. https://doi.org/10.1016/j.vaccine.2018.01.066
Mishra AK, Choi J, Moon E, Baek KH (2018) Tryptophan-rich and proline-rich antimicrobial peptides. Molecules 23(4):815. https://doi.org/10.3390/molecules23040815
Mor A, Nicolas P (1994) Isolation and structure of novel defensive peptides from frog skin. Eur J Biochem 219(1–2):145–154. https://doi.org/10.1111/j.1432-1033.1994.tb19924.x
Muralidharan N et al (2020) Computational studies of drug repurposing and synergism of lopinavir, oseltamivir and ritonavir binding with SARS-CoV-2 protease against COVID-19. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2020.1752802
Mustafa S, Balkhy H, Gabere M (2019) Peptide-protein interaction studies of antimicrobial peptides targeting middle east respiratory syndrome coronavirus spike protein: an in silico approach. Adv Bioinform 2019:1–16. https://doi.org/10.1155/2019/6815105
Patil R et al (2010) Optimized hydrophobic interactions and hydrogen bonding at the target-ligand interface leads the pathways of drug-designing. PLoS ONE 5(8):e12029. https://doi.org/10.1371/journal.pone.0012029
Pettersen EF et al (2004) UCSF Chimera—a visualization system for exploratory research and analysis. J Comput Chem 25(13):1605–1612. https://doi.org/10.1002/jcc.20084
Pierce BG et al (2014) ZDOCK server: interactive docking prediction of protein-protein complexes and symmetric multimers. Bioinformatics 30(12):1771–1773. https://doi.org/10.1093/bioinformatics/btu097
Porollo AA, Adamczak R, Meller J (2004) POLYVIEW: a flexible visualization tool for structural and functional annotations of proteins. Bioinformatics 20(15):2460–2462. https://doi.org/10.1093/bioinformatics/bth248
Reichen C, Hansen S, Plückthun A (2014) Modular peptide binding: from a comparison of natural binders to designed armadillo repeat proteins. J Struct Biol 185(2):147–162. https://doi.org/10.1016/j.jsb.2013.07.012
Rigsby RE, Parker AB (2016) Using the PyMOL application to reinforce visual understanding of protein structure. Biochem Mol Biol Educ 44(5):433–437. https://doi.org/10.1002/bmb.20966
Robert X, Gouet P (2014) Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res 42(W1):W320–W324. https://doi.org/10.1093/nar/gku316
Scheff SW (2016) Nonparametric statistics. In: Fundamental statistical principles for the neurobiologist. Elsevier, Amsterdam, pp 157–182. https://doi.org/10.1016/b978-0-12-804753-8.00008-7
Shai Y, Oren Z (2001) From “carpet” mechanism to de-novo designed diastereomeric cell-selective antimicrobial peptides. Peptides 22(10):1629–1641. https://doi.org/10.1016/S0196-9781(01)00498-3
Shalev DE et al (2006) Consequences of N-acylation on structure and membrane binding properties of dermaseptin derivative K 4–S4-(1–13). J Biol Chem 281(14):9432–9438. https://doi.org/10.1074/jbc.M513051200
Shang J et al (2020) Cell entry mechanisms of SARS-CoV-2. Proc Natl Acad Sci 117(21):11727–11734. https://doi.org/10.1073/pnas.2003138117
Shen Y et al (2014) Improved PEP-FOLD approach for peptide and miniprotein structure prediction. J Chem Theory Comput 10(10):4745–4758. https://doi.org/10.1021/ct500592m
Shirvanyants D et al (2012) Discrete molecular dynamics: an efficient and versatile simulation method for fine protein characterization. J Phys Chem B 116(29):8375–8382. https://doi.org/10.1021/jp2114576
Singh R et al (2020) Natural analogues inhibiting selective cyclin-dependent kinase protein isoforms: a computational perspective. J Biomol Struct Dyn 38(17):5126–5135. https://doi.org/10.1080/07391102.2019.1696709
Skjaerven L, Hollup SM, Reuter N (2009) Normal mode analysis for proteins. J Mol Struct (Thoechem) 898(1–3):42–48. https://doi.org/10.1016/j.theochem.2008.09.024
Sowmya G, Ranganathan S (2015) Discrete structural features among interface residue-level classes. BMC Bioinform 16(S18):S8. https://doi.org/10.1186/1471-2105-16-S18-S8
Srinivasan E, Rajasekaran R (2017) Exploring the cause of aggregation and reduced Zn binding affinity by G85R mutation in SOD1 rendering amyotrophic lateral sclerosis: in silico study on SOD1 mutant G85R. Proteins: Struct Funct Bioinform 85(7):1276–1286. https://doi.org/10.1002/prot.25288
Tina KG, Bhadra R, Srinivasan N (2007) PIC: protein interactions calculator. Nucleic Acids Res 35(Web Server issue):W473–W476. https://doi.org/10.1093/nar/gkm423
Tiwari SP et al (2014) WEBnm@ v2.0: web server and services for comparing protein flexibility. BMC Bioinform 15(1):427. https://doi.org/10.1186/s12859-014-0427-6
Wang Q et al (2020) Structural and functional basis of SARS-CoV-2 entry by using human ACE2. Cell 181(4):894–904. https://doi.org/10.1016/j.cell.2020.03.045
Wrapp D et al (2020) Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 367(6483):1260–1263. https://doi.org/10.1126/science.abb2507
Wu Y et al (2020) A noncompeting pair of human neutralizing antibodies block COVID-19 virus binding to its receptor ACE2. Science 368(6496):1274–1278. https://doi.org/10.1126/science.abc2241
Xia S et al (2020) Inhibition of SARS-CoV-2 (previously 2019-nCoV) infection by a highly potent pan-coronavirus fusion inhibitor targeting its spike protein that harbors a high capacity to mediate membrane fusion. Cell Res 30(4):343–355. https://doi.org/10.1038/s41422-020-0305-x
Yan R et al (2020) Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2. Science 367(6485):1444–1448. https://doi.org/10.1126/science.abb2762
Yang L, Song G, Jernigan RL (2009) Comparisons of experimental and computed protein anisotropic temperature factors: protein anisotropic temperature factors. Proteins: Struct Funct Bioinform 76(1):164–175. https://doi.org/10.1002/prot.22328
Zaïri A et al (2014) In vitro activities of dermaseptins K4 S4 and K4 K20 S4 against Escherichia coli, Staphylococcus aureus, and Pseudomonas aeruginosa planktonic growth and biofilm formation. Antimicrob Agents Chemother 58(4):2221–2228. https://doi.org/10.1128/AAC.02142-13
Zhou P et al (2018) HPEPDOCK: a web server for blind peptide–protein docking based on a hierarchical algorithm. Nucleic Acids Res 46(W1):W443–W450. https://doi.org/10.1093/nar/gky357
Acknowledgements
The authors thank VIT for providing ‘VIT SEED GRANT (VIT/SG/2020-21/43)’ for carrying out this research work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there are no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sekar, P.C., Rajasekaran, R. Could Dermaseptin Analogue be a Competitive Inhibitor for ACE2 Towards Binding with Viral Spike Protein Causing COVID19?: Computational Investigation. Int J Pept Res Ther 27, 1043–1056 (2021). https://doi.org/10.1007/s10989-020-10149-w
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10989-020-10149-w