Abstract
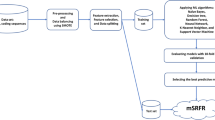
The Glutathione S-Transferases (GSTs) are detoxification enzymes which exist in variety of living organisms such as bacteria, fungi, plants and animals. These multifunctional enzymes play important roles in the biosynthesis of steroids, prostaglandins, apoptosis regulation, and stress signaling. In this study, we designed a method to independently predict the structures of animal, fungal and plant GSTs using Chou’s pseudo-amino acid composition concept. Support vector machine (SVM), Random Forests (RF), Covariance Discrimination (CD) and Optimized Evidence-Theoretic K-nearest Neighbor (OET-KNN) were used as powerful machine learnings algorithms. Based on our results, Random Forests demonstrated the best prediction for animal GSTs with 0.9339 accuracy and SVM showed the best results for fungal and plant GSTs with 0.8982 and 0.9655 accuracy, respectively. Our study provided an effective prediction for GSTs based on the concept of PseAAC and four different machine learning algorithms.
Similar content being viewed by others
References
Allocati N, Masulli M, Di Ilio C, Federici L (2018) Glutathione transferases: substrates, inihibitors and pro-drugs in cancer and neurodegenerative diseases. Oncogenesis 7:1–15
Behbahani M, Nosrati M, Moradi M, Mohabatkar H (2019) Using Chou’s general pseudo amino acid composition to classify laccases from bacterial and fungal sources via Chou’s five-step rule. Appl Biochem Biotechnol 190:1035–1048
Breiman L (2001) Random forests. Machine Learn 45:5–32
Cao D-S, Xu Q-S, Liang Y-Z (2013) propy: a tool to generate various modes of Chou’s PseAAC. Bioinformatics 29:960–962
Chakrabarti S, Ester M, Fayyad U, Gehrke J, Han J, Morishita S et al (2006) Data mining curriculum: A proposal (Version 1.0). Intensive Working Group of ACM SIGKDD Curriculum Committee
Chen H, Kihara D (2011) Effect of using suboptimal alignments in template-based protein structure prediction. Proteins Struct Funct Bioinf 79:315–334
Chen C, Chen L, Zou X, Cai P (2009) Prediction of protein secondary structure content by using the concept of Chou’s pseudo amino acid composition and support vector machine. Protein Pept Lett 16:27–31
Chen W, Lei T-Y, Jin D-C, Lin H, Chou K-C (2014) PseKNC: a flexible web server for generating pseudo K-tuple nucleotide composition. Anal Biochem 456:53–60
Chen W, Lin H, Chou K-C (2015) Pseudo nucleotide composition or PseKNC: an effective formulation for analyzing genomic sequences. Mol BioSyst 11:2620–2634
Chen W, Tang H, Ye J, Lin H, Chou K-C (2016) iRNA-PseU: identifying RNA pseudouridine sites. Mol Ther Nucleic Acids 5:e332
Chen Y, Banerjee D, Mukhopadhyay A, Petzold CJ (2020) Systems and synthetic biology tools for advanced bioproduction hosts. Curr Opin Biotechnol 64:101–109
Chou KC (2001) Prediction of protein cellular attributes using pseudo-amino acid composition. Proteins: Struct Funct Bioinf 43:246–255
Chou K-C (2005) Using amphiphilic pseudo amino acid composition to predict enzyme subfamily classes. Bioinformatics 21:10–19
Chou K-C (2009) Pseudo amino acid composition and its applications in bioinformatics, proteomics and system biology. Curr Proteomics 6:262–274
Chou K-C (2011) Some remarks on protein attribute prediction and pseudo amino acid composition. J Theor Biol 273:236–247
Chou K-C (2015) Impacts of bioinformatics to medicinal chemistry. Med Chem 11:218–234
Chou K-C (2017) An unprecedented revolution in medicinal chemistry driven by the progress of biological science. Curr Topics Med Chem 17:2337–2358
Chou K-C (2019) Advances in predicting subcellular localization of multi-label proteins and its implication for developing multi-target drugs. Curr Med Chem 26:4918–4943
Chou K-C (2020) Proposing 5-steps rule is a notable milestone for studying molecular biology. Nat Sci 12:74
Chou KC, Cai YD (2003) Predicting protein quaternary structure by pseudo amino acid composition. Proteins Struct Funct Bioinf 53:282–289
Chou K-C, Cheng X, Xiao X (2019) pLoc_bal-mEuk: predict subcellular localization of eukaryotic proteins by general PseAAC and quasi-balancing training dataset. Med Chem 15:472–485
Dasari S, Ganjayi MS, Yellanurkonda P, Basha S, Meriga B (2018) Role of glutathione S-transferases in detoxification of a polycyclic aromatic hydrocarbon, methylcholanthrene. Chemico-Biol Interact 294:81–90
Dehzangi A, Heffernan R, Sharma A, Lyons J, Paliwal K, Sattar A (2015) Gram-positive and Gram-negative protein subcellular localization by incorporating evolutionary-based descriptors into Chou׳ s general PseAAC. J Theor Biol 364:284–294
Di Matteo A, Federici L, Masulli M, Carletti E, Santorelli D, Cassidy J et al (2019) Structural characterization of the Xi Class glutathione transferase from the Haloalkaliphilic Archaeon Natrialba magadii. Front Microbiol 10:9
Du P, Wang X, Xu C, Gao Y (2012) PseAAC-Builder: a cross-platform stand-alone program for generating various special Chou’s pseudo-amino acid compositions. Anal Biochem 425:117–119
Du P, Gu S, Jiao Y (2014) PseAAC-General: fast building various modes of general form of Chou’s pseudo-amino acid composition for large-scale protein datasets. Int J Mol Sci 15:3495–3506
Esmaeili M, Mohabatkar H, Mohsenzadeh S (2010) Using the concept of Chou’s pseudo amino acid composition for risk type prediction of human papillomaviruses. J Theor Biol 263:203–209
Galetsi P, Katsaliaki K, Kumar S (2020) Big data analytics in health sector: theoretical framework, techniques and prospects. Int J Inf Manag 50:206–216
Ghosh C, Saha S, Saha S, Ghosh N, Singha K, Banerjee A et al (2020) Machine Learning Based Supplementary Prediction System Using K Nearest Neighbour Algorithm. Available at SSRN 3517197
Gupta CLP, Bihari A, Tripathi S (2019) Protein classification using machine learning and statistical techniques: a comparative analysis. arXiv preprint arXiv:190106152
Haghighi O, Davaeifar S, Zahiri HS, Maleki H, Noghabi KA (2019) Homology Modeling and Molecular Docking Studies of Glutamate Dehydrogenase (GDH) from Cyanobacterium Synechocystis sp. PCC 6803. Int J Pept Res Ther 26:783–793
Kam HT (1995) Random decision forest. In: Proceedings of the 3rd international conference on document analysis and recognition, Montreal, Canada, 14–16 August 1995. IEEE, p 278282
Kato T, Miyakawa H, Ishibashi M (2004) Frequency and significance of anti-glutathione S-transferase autoantibody (anti-GST A1-1) in autoimmune hepatitis. J Autoimmun 22:211–216
Kilty C, Doyle S, Hassett B, Manning F (1998) Glutathione S-transferases as biomarkers of organ damage: applications of rodent and canine GST enzyme immunoassays. Chemico-Biol Interact 111:123–135
Landi S (2000) Mammalian class theta GST and differential susceptibility to carcinogens: a review. Mutat Res/Rev Mutat Res 463:247–283
Lee S, Lee B, Kim D (2006) Prediction of protein secondary structure content using amino acid composition and evolutionary information. Proteins Struct Funct Bioinf 62:1107–1114
Li W, Godzik A (2006) Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22:1658–1659
Li Y, Wu F-X, Ngom A (2018) A review on machine learning principles for multi-view biological data integration. Brief Bioinform 19:325–340
Lin H (2008) The modified Mahalanobis discriminant for predicting outer membrane proteins by using Chou’s pseudo amino acid composition. J Theor Biol 252:350–356
Lin W, Xiao X, Qiu W, Chou K-C (2020) Use Chou’s 5-steps rule to predict remote homology proteins by merging grey incidence analysis and domain similarity analysis. Nat Sci 12:181
Liu B, Liu F, Wang X, Chen J, Fang L, Chou K-C (2015) Pse-in-One: a web server for generating various modes of pseudo components of DNA, RNA, and protein sequences. Nucleic Acids Res 43:W65–W71
Liu B, Fang L, Long R, Lan X, Chou K-C (2016) iEnhancer-2L: a two-layer predictor for identifying enhancers and their strength by pseudo k-tuple nucleotide composition. Bioinformatics 32:362–369
Liu B, Wu H, Chou K-C (2017) Pse-in-One 2.0: an improved package of web servers for generating various modes of pseudo components of DNA, RNA, and protein sequences. Nat Sci 9:67
Liu B, Yang F, Huang D-S, Chou K-C (2018) iPromoter-2L: a two-layer predictor for identifying promoters and their types by multi-window-based PseKNC. Bioinformatics 34:33–40
Mishra NK, Kumar M, Raghava G (2007) Support vector machine based prediction of glutathione S-transferase proteins. Protein Pept Lett 14:575–580
Mohabatkar H (2010) Prediction of cyclin proteins using Chou’s pseudo amino acid composition. Protein Pept Lett 17:1207–1214
Mohabatkar H, Beigi MM, Esmaeili A (2011) Prediction of GABAA receptor proteins using the concept of Chou’s pseudo-amino acid composition and support vector machine. J Theor Biol 281:18–23
Pal M (2005) Random forest classifier for remote sensing classification. Int J Remote Sens 26:217–222
Raza K (2012) Application of data mining in bioinformatics. arXiv preprint arXiv:12051125
Roberts E, Eargle J, Wright D, Luthey-Schulten Z (2006) MultiSeq: unifying sequence and structure data for evolutionary analysis. BMC Bioinform 7:382
Schultz IR, Sylvester SR (2001) Stereospecific toxicokinetics of bromochloro-and chlorofluoroacetate: Effect of GST-ζ depletion. Toxicol Appl Pharmcol 175:104–113
Schölkopf B, Smola AJ, Bach F (2002) Learning with kernels: support vector machines, regularization, optimization, and beyond. MIT press, Cambridge
Shen H, Chou K-C (2005) Using optimized evidence-theoretic K-nearest neighbor classifier and pseudo-amino acid composition to predict membrane protein types. Biochem Biophys Res Commun 334:288–292
Shen H-B, Chou K-C (2008) PseAAC: a flexible web server for generating various kinds of protein pseudo amino acid composition. Anal Biochem 373:386–388
Snoek J, Larochelle H, Adams RP (2012) Practical bayesian optimization of machine learning algorithms. Adv Neural Inf Process Syst 2:2951–2959
Suykens JA, Vandewalle J (1999) Least squares support vector machine classifiers. Neural Process Lett 9:293–300
Sylvestre-Gonon E, Law S, Schwartz M, Robe K, Keech O, Didierjean C et al (2019) Functional, structural and biochemical features of plant serinyl-glutathione transferases. Front Plant Sci 10:608
Tew KD, Ronai ZE (1999) GST function in drug and stress response. Drug Resist Updates 2:143–147
Tian B, Wu X, Chen C, Qiu W, Ma Q, Yu B (2019) Predicting protein–protein interactions by fusing various Chou’s pseudo components and using wavelet denoising approach. J Theor Biol 462:329–346
Xia J-F, Han K, Huang D-S (2010) Sequence-based prediction of protein-protein interactions by means of rotation forest and autocorrelation descriptor. Protein Pept Lett 17:137–145
Xiao X, Cheng X, Chen G, Mao Q, Chou K-C (2019) pLoc_bal-mVirus: predict subcellular localization of multi-label virus proteins by Chou’s general PseAAC and IHTS treatment to balance training dataset. Med Chem 15:496–509
Yadav SK, Tiwari AK (2015) Classification of enzymes using machine learning based approaches: a review. Machine Learn Appl 2:30–49
Yu B, Li S, Qiu W-Y, Chen C, Chen R-X, Wang L et al (2017) Accurate prediction of subcellular location of apoptosis proteins combining Chou’s PseAAC and PsePSSM based on wavelet denoising. Oncotarget 8:107640
Zou Q, Lin G, Jiang X, Liu X, Zeng X (2020) Sequence clustering in bioinformatics: an empirical study. Brief Bioinform 21:1–10
Acknowledgements
Support of this study by the University of Isfahan is acknowledged.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
There is no conflict to declare.
Informed Consent
There was no human participant and consent was not required.
Research involving Human and/or Animals Participants
No human or animal was participated in this study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Mohabatkar, H., Ebrahimi, S. & Moradi, M. Using Chou’s Five-steps Rule to Classify and Predict Glutathione S-transferases with Different Machine Learning Algorithms and Pseudo Amino Acid Composition. Int J Pept Res Ther 27, 309–316 (2021). https://doi.org/10.1007/s10989-020-10087-7
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10989-020-10087-7