Abstract
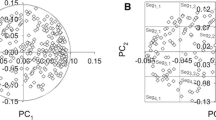
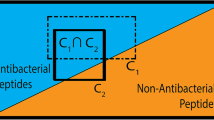
Antimicrobial peptides are a promising class of substances for overcoming multidrug resistant bacteria. In a previous study, high-throughput screening of short peptides for antimicrobial activity against Pseudomonas aeruginosa was performed and the resulting data with 1609 peptides was analyzed using quantitative structure–activity relationship models (QSAR) with excellent prediction power. To avoid non-interpretable black-box behavior of the QSAR models, new features based on fuzzy logic and molecular descriptors were introduced. They were used for comprehensive analysis and visualization. The new features provide good interpretability and are able to differentiate between active and inactive peptides. The statistical relevance of this differentiation was shown using a Wilcoxon rank sum test. The best compromise between activity prediction and interpretability was found for fuzzy terms of the Hopp-Woods scale and the Isoelectric point. A visualization of two of these terms enables an in-depth understanding of regions with active and inactive peptides and the identification of outliers. In addition, we generated rules to explain typical amino acid distributions in active peptides. These rules can be used to increase the probability of finding active peptides in new peptide libraries, which can improve the speed of finding leading substances for drug development against resistant bacteria.




Similar content being viewed by others
References
Bezdek JC (1981) Pattern recognition with fuzzy objective function algorithms. Plenum Press, New York
Brogden K (2005) Antimicrobial peptides pore formers or metabolic inhibitors in bacteria? Nat Rev Microbiol 3(3):238–250
Cherkasov A, Hilpert K, Jenssen H, Fjell CD, Waldbrook M, Mullaly SC, Volkmer R, Hancock RE (2008) Use of artificial intelligence in the design of small peptide antibiotics effective against a broad spectrum of highly antibiotic-resistant superbugs. ACS Chem Biol (in press)
Fabry-Asztalos L, Andonie R, Collar C, Abdul-Wahid S, Salim N (2008) A genetic algorithm optimized fuzzy neural network analysis of the affinity of inhibitors for HIV-1 protease. Bioorg Med Chem 11:2903–2911
Finking R, Marahiel M (2004) Biosynthesis of nonribosomal peptides. Ann Rev Microbiol 58(1):453–488
Frecer V (2006) QSAR analysis of antimicrobial and haemolytic effects of cyclic cationic antimicrobial peptides derived from protegrin-1. Bioorg Med Chem 14:6065–6074
Hamill P, Brown K, Jenssen H, Hancock R (2008) Novel anti-infectives: is host defence the answer? Curr Opin Biotechnol 19(6):628–636
Hancock R (2001) Cationic peptides effectors in innate immunity and novel antimicrobials. Lancet Infect Dis 1(3):156–164
Hancock R, Lehrer R (1998) Cationic peptides: a new source of antibiotics. Trends Biotechnol 16(2):82–88
Hancock R, Falla T, Brown M (1995) Cationic bactericidal peptides. Adv Microb Physiol 37:135–175
Hellberg S, Sjostrom M, Skagerberg B, Wold S (1987) Peptide quantitative structure–activity relationships, a multivariate approach. J Med Chem 30:1126–1135
Hildebrand P (2006) Zur Strukturvorhersage der Membranproteine. PhD thesis, Humboldt-Universität zu Berlin, Mathematisch-Naturwissenschaftliche Fakultät I
Hilpert K, Hancock REW (2007a) Use of luminescent bacteria for rapid screening and characterization of short cationic antimicrobial peptides synthesized on cellulose by peptide array technology. Nat Protoc 2(7):1652–1660
Hilpert K, Elliott MR, Volkmer-Engert R, Henklein P, Donini O, Zhou Q, Winkler DFH, Hancock REW (2006) Sequence requirements and an optimization strategy for short antimicrobial peptides. Chem Biol 13(10):101–1107
Hilpert K, Winkler DFH, Hancock REW (2007b) Peptide arrays on cellulose support: SPOT synthesis—a time and cost efficient method for synthesis of large numbers of peptides in a parallel and addressable fashion. Nat Protoc 2(6):1333–1349
Hilpert K, Fjell C, Cherkasov A (2008) Short linear cationic antimicrobial peptides: screening, optimizing, and prediction. Methods Mol Biol 494:127–159
Jenssen H, Hamill P, Hancock R (2006) Peptide antimicrobial agents. Clin Microbiol Rev 19(3):491–511
Lejon T, Stroem M, Svendsen J (2001) Antibiotic activity of pentadecapeptides modelled from amino acid descriptors. J Pept Sci 7(2):74–81
Mekenyan O, Nikolova N, Schmieder P (2003) Dynamic 3D QSAR techniques: applications in toxicology. J Mol Struct: THEOCHEM 622(1–2): 147–165
Mikut R, Jäkel J, Gröll L (2005) Interpretability issues in data-based learning of fuzzy systems. Fuzzy Sets Syst 150(2):179–197
Mikut R, Burmeister O, Braun S, Reischl M (2008a) The open source MATLAB toolbox Gait-CAD and its application to bioelectric signal processing. In: Proceedings of DGBMT—workshop biosignal analysis, Potsdam, pp 109–111
Mikut R, Reischl M, Ulrich A, Hilpert K (2008b) Data-based activity analysis and interpretation of small antibacterial peptides. In: Proceedings of 18th workshop computational intelligence, Universitätsverlag Karlsruhe, pp 189–203
Ostberg N, Kaznessis Y (2005) Protegrin structure–activity relationships using homology models of synthetic sequences to determine structural characteristics important for activity. Peptides 26(2):197–206
Perkins R, Fang H, Tong W, Welsh W (2003) Quantitative structure–activity relationship methods: perspectives on drug discovery and toxicology. Environ Toxicol Chem 22:1666–1679
Ros F, Pintore M, Chrétien J (2002) Molecular descriptor selection combining genetic algorithms and fuzzy logic: application to database mining procedures. Chemom Intell Lab Syst 63(1):15–26
Rosenfeld Y, Sahl H, Shai Y (2008) Parameters involved in antimicrobial and endotoxin detoxification activities of antimicrobial peptides. Biochemistry 47(24):6468–6478
Sandberg M, Eriksson L, Jonsson J, Sjöström M, Wold S (1998) New chemical descriptors relevant for the design of biologically active peptides. A multivariate characterization of 87 amino acids. J Med Chem 41(14):2481–2491
Strom M, Stensen W, Svendsen J, Rekdal O (2001) Increased antibacterial activity of 15-residue murine lactoferricin derivatives. J Pept Res 57(2):127–139
White S, Wimley W (1999) Membrane protein folding and stability: physical principles. Ann Rev Biophys Biomol Struct 28(1):319–336
Zadeh L (1965) Fuzzy sets. Inf Control 8:338–353
Acknowledgement
We thank R. A. Klady for the critical proofreading of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mikut, R., Hilpert, K. Interpretable Features for the Activity Prediction of Short Antimicrobial Peptides Using Fuzzy Logic. Int J Pept Res Ther 15, 129–137 (2009). https://doi.org/10.1007/s10989-009-9172-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10989-009-9172-5