Abstract
The KB-Rank tool was developed to help determine the functions of proteins. A user provides text query and protein structures are retrieved together with their functional annotation categories. Structures and annotation categories are ranked according to their estimated relevance to the queried text. The algorithm for ranking first retrieves matches between the query text and the text fields associated with the structures. The structures are next ordered by their relative content of annotations that are found to be prevalent across all the structures retrieved. An interactive web interface was implemented to navigate and interpret the relevance of the structures and annotation categories retrieved by a given search. The aim of the KB-Rank tool is to provide a means to quickly identify protein structures of interest and the annotations most relevant to the queries posed by a user. Informational and navigational searches regarding disease topics are described to illustrate the tool’s utilities. The tool is available at the URL http://protein.tcmedc.org/KB-Rank.
Similar content being viewed by others
Introduction
The ability to search for proteins of interest via text query is a standard utility of protein biomedical resources such as NCBI Protein [1], MMDB [2], UniProt [3], sites created for the Protein Data Bank (PDB) by the members of the wwPDB [4] (RCSB Protein Data Bank [5], PDBe [6], BMRB [7], and PDBj [8]), and the Structural Biology Knowledgebase (SBKB) [9]. These resources offer search services over a variety of annotations. For example, NCBI protein has curated information regarding protein sequences that is available for text query. UniProt hosts text searches over of a collection of annotation records of the protein sequences, which were collected based a review of the associations documented in the literature and/or were derived computationally. The SBKB provides searches for protein structures over summary text fields from the primary literature citations. The fields includes abstracts and associated terms such as medical subject headings or MeSH terms [9]. The wwPDB websites offer a variety of searches that include those over the collections of text fields from primary literature citations and the cross-referenced annotations from other protein databases [5, 8]. As examples, searches for ligands contained within the protein structures have also been implemented [5, 6, 8]. With these and related protein resources, users have at their disposal a means to search for protein structures based on a collection of associated protein annotations and attributes. A recent review of protein databases and some of the associated searches that are available therein is provided by Chen et al. [10].
The presentation of the results of a text query within a protein database can be done in which users can browse all the entries that match any of the text fields or browse only entries that have matches within specified fields. For example, UniProt allows a user to retrieve matches based on all the annotations collected within the UniProt data files or restrict the search to matches within a particular annotation fields. Annotation fields in UniProt include the protein or organism name fields. Similarly, the RCSB PDB provides a list of all the protein structures found based on matches across all the available text fields or the results for searches that are restricted to matches within particular annotation categories, such as the enzyme type or a Gene Ontology term category. Given that text searches may produce a large collection of annotations and structures that may possibly be browsed, the user may ask the following. Which structures are the most relevant to my query? Of the annotations retrieved, which ones are the most relevant? These questions are analogous to those commonly made for website searches with regard to which topics and which web pages are estimated to be the relevant to a given query. User demand to expand the utilities of web search engines has lead to the development of more efficient and effective methods to retrieve the most relevant topics and web pages to a given text query [11].
With the goal to achieve improved efficiency and effectiveness for searches for protein structures and their associated annotation categories, a ranking tool, KB-Rank, is described. The KB-Rank tool provides a means to retrieve a list of protein structural chains and annotation categories that are relevant to the provided text query. Structural chains within each retrieved category are ranked according to their estimated relevance to the queried text. The annotation categories are also presented according to their estimated relevance. These utilities can be used to address a variety of searches that are conducted by users of protein structural databases. The tool facilitates informational searches to learn more about particular topics, e.g. the retrieval of information associated with a particular disease. An example of an informational search example is to gain a better understanding of the pathogenic mechanisms of asthma. Navigational searches are also enabled that provide a means to identify specific structural chains that can be used to address particular research questions. One such type of search is to find structures that may be used in a structure based drug design protocol, for example protein structural chains may be used in drug design strategies for the treatment of melanoma.
Materials and methods
Annotation assembly
The assembly and integration of the protein annotations from open sources is done weekly to coordinate with the release of new protein structures and to ensure that the analysis is up date for all available structures. Annotations are mapped to protein structures at the level of the protein structural chain. A full list of protein structural chains is available from the ftp site at a URL at the PDB <ftp://ftp.wwpdb.org/pub/pdb/derived_data/pdb_seqres.txt>. The following annotations are assembled. Cellular and biochemical pathway assignments were extracted from BioCyc [12], CellMap [13], HumanCyc [13], INOH [14], and the NCI Pathway Interaction Database (PID) [15]. Small molecule associations were from BioCyc, BindingDB [16], HumanCyc, DrugBank [17], ChEBI [18], ChEMBL [19], and SMPDB [20]. SNPs3D [21] and OMIM [22, 23] provided disease associations. Molecular functions, biological processes, cellular components were from the Gene Ontology (GO) classification system [24], as assigned in SIFTS [25]. Enzyme classifications were from the EC2PDB database [26]. Structural domains assignments were provided through the CATH [27] and SCOP [28] databases. Sequence domain assignments were identified through the Pfam database [29]. Further structural groups were based on the jFatCat alignment algorithm [30, 31]. The FEATURE resource provided predictions of functional sites [32]. The annotations utilized in the current study have been described previously for the purpose of the prediction of protein function [33], and more complete description of their assembly is provided therein.
Query and presentation of protein structures and annotation categories
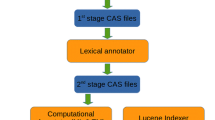
At the first stage of the text query, the search is over the text fields associated with the primary literature citations of the protein structures and text associated with domains within the structures as retrieved by the Pfam protein family database [29]. The fields from the primary literature citations include the title, author list, abstract, medical subject headings or MeSH terms, and the substance list. Text from the Pfam database is from the description and comment fields. A search over all of the fields provides a means to identify and rank structure entries as a list of PDBIDs. Such a search using text from the primary literature fields has been implemented in the SBKB [34]. The scoring for the ranking is implemented using MySQL’s “Match…Against” method [35]. The following gives a summary of the formula utilized as discussed at the URL <http://forge.mysql.com/wiki/MySQL_Internals_Algorithms>.
In the equation, the variable dtf is the number of times the term appears in the document, sumdtf is the sum of (log (dtf) + 1)’s for all terms in the same document; U is the number of unique terms in the document; N is the total number of documents; and nf is the number of documents that contain the term. Based on the keyword match within the text fields and using equation A, structural entries or PDBIDs are retrieved and ranked. The first 200 entries found by the text search are saved for further analysis.
The next task is to order the protein chains within the retrieved structural entries. From all the structural chains within the entries retrieved by a given text search, a nonredundant set is obtained by identifying representative chains that are nonidentical in primary amino acid sequence. All the annotations associated with these nonidentical structural chains are then retrieved. For each of the structural chains, an array of values that corresponds to each of the annotations retrieved was created. If a structural chain had a given annotation, as found in any of the representative chains, a one is entered for that position in the array. If not, a zero is entered at that position. A matrix with the arrays of the structural chains versus annotation presence or absence is then generated. To rank the structural chains, the array for each structural chain is multiplied by the entire matrix created from the representative set of chains in primary sequence. All the elements of product matrix are summarized to get a rank value. Structural chains are ordered according to value of the rank, which referred to as the relevance score. Annotation categories are also ranked according to the average relevance score of the structural chains in each category. See Fig. 1 for an illustration of the method. A comparable method was used to find a relevant transcription factor binding sites among potential promoter sequences [36].
A truncated annotation profile matrix, structures versus annotations, provides a schematic of the method that is used to order the protein structural chains according to their relevance to a queried text. If an annotation is associated with a given structure, the entry in the corresponding point in the matrix was a one and zero otherwise. Structural chains are ranked based on the product of its annotation profile array multiplied by the entire annotation profile matrix that was created for the given search. The product for each chain is referred to as the chain’s relevance score for the given text search. See text for details. In the diagram shown, structure with PDBID 1HZI chain A is ranked highest, followed by 2ZEC chain A, and so on. The annotations are: Annotation A-CATH homologous family Trysin-like serine proteases, Annotation B—SCOP superfamily Ferritin-like, Annotation C—GO cellular component extracellular region, Annotation D—SNPs3D disease association hypertension, Annotation E—ChEMBL small molecule association l-Serine, and Annotation F—jFatCat structural group with similarity to structure with PDBID 3GOV, chain B
Results
The web interface
An interactive web interface for the KB-Rank tool was created to search and browse the protein structural chains retrieved and their associated annotation categories. The main page is shown in Fig. 2. A text search box is provided whereby the user can initiate a query. Annotation categories that are available for browsing include cellular pathways retrieved from the National Cancer Institute’s Pathway Interaction Database [15]; superfamily designations provided from the SCOP database [28]; small molecule associations and metabolic pathway associations as assigned in BioCyc [12]; enzyme classification assignments from EC2PDB [26, 37]; molecular function, biological process, and cellular component term designations as found in the Gene Ontology term hierarchy [24]; and small associations that are assigned within the ChEBI [18], ChEMBL [38], SMPDB [20], and DrugBank [17] resources. The resources are listed in the tabs at the top of the search results page where one can choose to retrieve results based on each. As the annotation categories are presented on the web interface, links are forged to the annotation provider’s website. That is done either to the home page of the resource or to the page that describes the annotation category selected, whichever is appropriate. A legend is provided on the results page to give a summary of each resource and what annotations are utilized from each.
The landing or main page of the KB-Rank tool describes its utility as a means to identify protein structures and biomedical annotations via text search. Types of search terms are given as examples are protein functions or disease associations of the protein structures. The search term shown is asthma
A utility of the KB-Rank query tool is that annotation categories and structural chains are ordered and presented according to their estimated relevance to the queried text. Relevance scores are used as described in Materials and Methods. To make the interpretation of the relevance scores more visually intuitive, colors are used to indicate where each annotation category or structural chain lies within the entire ranges of the scores. As an analogy to a traffic light, a green color indicates that a category or structural chain is most associated with the queried text while a red color indicates that is least relevant. Colors in between are used to indicate intermediate scores and corresponding relevance. The coloring method is comparable with that used within the protein modeling portal [39], where model quality for a predicted structure, rather than relevance to text query, is similarly assessed.
User case scenarios
Illustrative user case scenarios are now described. For the first scenario, the aim is to perform an informational search on a particular topic. An example search is regarding the disease asthma. The user wants to learn more about that disease based on a review of the protein structures and annotation categories that are found to be relevant. Upon executing the text search, the user selects from annotation categories that can be browsed. A selection of the Gene Ontology resource for the categories within the ontology domain of molecular function is shown in Fig. 3, panel A. The highest ranked molecular function category retrieved is interleukin-4 receptor binding, GO + 0005136. A review of the primary literature shows that interactions of interleukin-4 are involved in the proinflammatory response in asthma; and interleukin-4 protein mediates the development of allergic reactions [40, 41]. For the cellular component ontology domain of GO, the results show that the highest ranking category is the extracellular matrix, GO + 0031012 (Fig. 3, panel B). Based on a literature review, it is known that in asthmatic patients, abnormal extracellular matrix components are deposited [42]. Also, in fatal cases of asthma, the fractional area of the extracellular matrix within airway smooth muscle is larger [43]. The categories of interleukin-4 binding and extracellular matrix were found to be the highest ranking in their respective Gene Ontology domains for queried text asthma. That corresponds well with each category’s importance regarding the pathogenesis of the disease. The utility of the tool in this case is that it aids the user in efficiently collecting relevant information about the disease.
Panel A resolution of the structures retrieved by the queried text asthma into molecular function categories as assigned in the Gene Ontology hierarchy. The results indicate that for protein structures associated with asthma the molecular functions associated with cytokine activity are prevalent. Also prevalent are protease activities. Panel B resolution of the structures retrieved by the queried text asthma into cellular component categories as assigned in the GO hierarchy. The results of the annotation category show that the protein participants in the disease engage in activity at the extracellular matrix. The ranking of the categories indicates that the component extracellular matrix, colored with a green correspondence, is relatively more relevant to the disease than the component stored secretory granule, which is shown in orange-yellow
A second utility of the KB-Rank tool is that it orders the protein structural chains within each annotation category according to their relative relevance to the queried text. That utility can aid the user in identifying the relatively more important chains, among all those retrieved, to a queried topic. The text search for asthma is further used to demonstrate that utility. As shown in Fig. 3 in panel A, cytokine activity, GO + 0005125, is ranked fifth among the list molecular function categories retrieved. The link provided for the 26 structural chains within that category can be expanded. In Fig. 4 is the structure of interleukin-5, PDB + 1HUL chain A, is listed, which is in the middle portion of the list. The associated orange color is used to indicate that the structure is estimated to have intermediate relevance to the queried text. At the top of the list retrieved but not shown in the Fig. 4, is interleukin-4, PDB + 1HZI, chain A. It has an associated bright green color that corresponds to the highest ranking structural chain found for the cytokine activity annotation category.
A list of the structures retrieved by the queried text asthma ordered according to their relative relevance to the disease. The results indicate that the cytokines eotaxin and IL-5, which are shown in orange, are estimated to be relatively less relevant than the cytokine IL-4. IL-4 is much higher in the list and has an associated green color. The PDBID and chain designation are given for each entry
Based on a review of the literature, the importance of IL-4 and IL-5 in the development of asthma can be assessed. It is known that that IL-4 contributes in a variety of ways to the development of asthma, one of which is the stimulation of Th0 lymphocytes to Th2 lymphocytes [40, 44]. Th2 lymphocytes secrete other cytokines that include additional IL-4, IL-5, IL-9, and IL-13. IL-5 thereby has a secondary role to disease development as compared to IL-4 in terms of the sequence of the disease mechanism. Further, IL-4 based therapies for asthma have shown improved clinical outcome for the treatment of asthma while IL-5 based therapies have not [45]. The results indicate the relative importance of the two cytokines in the pathogenesis of asthma, and that matches with relevance ranking found by the KB-Rank search tool. The ranking of the structures by the tool thereby provides a starting point for further understanding of the disease mechanism with regard the important protein players and their roles.
In addition to providing informational searches that utilize the ranking of the structural chains and annotation categories, navigational searches are also possible with the KB-Rank tool. In a navigational search, the purpose is to identify a particular structural chain that can be used for further investigation and research. An example type of a navigational search is for the identification of structural chains that can be used in a structure based drug design (SBDD) protocol and virtual screening. For that application, a user searches for a potential drug target that is particularly important to the disease of interest [46]. Selection is further made to find those protein structures that are druggable, i.e. protein structures that have binding pockets and/or that can accommodate a drug molecule [47].
As an illustrative example of a navigational search for SBDD, a search was made to identify those that can be targeted to treat melanoma. See Fig. 5 where the text query is melanoma. Based in part on information from the DrugBank resource, the highest ranked small molecule found by the search is 5-Bromo-N-(2,3-Dihydroxypropoxy)-3,4-Difluoro-2-[(2-Fluoro-4-Iodophenyl)Amino] Benzamide, DrugBank + DB03115. The entry in DrugBank for the small molecule indicates that it is an experimental molecule, and the protein target is MEK1, PDB + 3E8N chain A. The finding that the MEK1 structure binds to small, drug-like molecule indicates that it is likely a druggable target. The next step was to verify that MEK1 plays an important role in the mechanism of disease in melanoma. The primary citation for the structure of MEK1, PDB + 3E8N chain A, indicates it is targeted for the treatment of various types of cancer including lung, colon, melanoma, pancreatic, and prostate cancer [48, 49]. The MEK1 protein is within a signal transduction cascade, the RAS-RAF-mitogen-activated protein kinase (MAPK)/extracellular signal-regulated kinase (ERK) kinase (MEK)-ERK pathway, that leads to cancer [50].
Structures associated with the disease melanoma were searched. A resolution of the drug associations of the structures retrieved was done based on the DrugBank resource. The structure of the B-Raf kinase is found to interact with the drug Sorafenib. The search demonstrates an application for the identification of structures that can be used for structure based drug design for the treatment of melanoma. The result of the search provides a protein of known three dimensional structure that is known to be a druggable protein target for the disease
Upon examination of the other small molecules found from DrugBank for the query of melanoma, we see that the second molecule listed is the drug Sorafenib. It inhibits another protein along the RAS-RAF-MEK-ERK pathway, B-Raf kinase35, PDB + 3C4D chain B. Inhibition with Sorafenib has not proven to be effective in clinical trials for melanoma [51, 52]. But the protein structure is demonstrated to be druggable, and further inhibitors of that target have been developed and demonstrated to have effective anti-melanoma effects in humans [53, 54]. These results indicate that applications of SBDD for the B-Raf kinase target are ongoing and yielding effective results. The identification of the structures of B-Raf kinase and MEK1 with the KB-Rank tool as structures can be used for SBDD for the treatment of melanoma demonstrates that the tool provides a point of entry for the identification known and potential protein structural targets. The high ranks of the viable targets found, based on the text searches, illustrate the utility of the tool for that purpose.
Discussion
The KB-Rank tool provides a means to attach a relevance score to structural chains and/or associated categories retrieved by a given a text query. It is anticipated that as more annotations are utilized for the ranking process, e.g. through the addition of more annotations associated with the primary amino acid sequence and/or the three-dimensional structures of the protein chains, the display order will more accurately reflect the order of their relevance to the queried text. The annotation categories can be expanded within the types of annotations that have already been assembled. These types include additional three dimensional structural characteristics, small molecule interaction assignments, functional site assignments, and cellular/biochemical pathway designations. The resultant granularity for the searches and subsequent ranking is at the chain level rather than at the level of the structural entry as found in the PDB. That has the advantage of narrowing down a search to particular chain within an entry that has multiple chains. It has the ability to identify relevant protein chain that resides within a complex that may not be directly relevant to the text searched.
The relationship between function and disease are anticipated topics for searches. At the first stage of the search, a text search is implemented over the summary fields extracted from PubMed abstracts of the primary citations of the protein structures and the descriptive fields of constituent domains of the structures as extracted from the Pfam database. In the second stage, an integrated set of annotations are used for categorizing the functional roles of the protein structural chains, and to subsequently rank the retrieved chains by an expected relevance to the queried text. The annotations used for the final ranking need not contain a match with the queried text; they need only be prevalent in the structures retrieved by the text search. The prevalence of a given annotation within the structural chains retrieved is used as an indicator that it is relevant to the queried text. Structures with a relatively larger number of the prevalent will be ranked relatively higher. Also, structures that have been well characterized with a relatively larger number of any annotations will tend to be ranked higher as well. That tendency is analogous to what is found for webpage ranking where the interest level in web pages, as reflected number of its links and its link structure [55], is used to facilitate the ranking.
Data integration effort forms the substrate for the search tool and connections forged between the annotations further lend utility to the search tool. For example, UniProt entries are connected with chain entries from the PDB; and DrugBank entries are connected with UniProt entries within the integrated database that is utilized by the KB-Rank tool. A result is that for the melanoma search example, the user can identify a small molecule in DrugBank that is associated with melanoma and be provided with a relevant protein structural chain. The result demonstrates the utility of the data integration aspect of the tool as an important component of the tool’s functionality and utility. To complete the data integration, sequence comparisons are done to map the protein chain to annotations. The mapping of entries in BindingDB to the structural chains was done by finding the corresponding sequences with greater than 90% sequence identify through sequence comparison using the BLAST program [56]. The SIFTs resource and the UniProt data files are also utilized to provide connections between the protein structural chains with a host of sequence and functional information [3, 25, 57].
To improve the calculation of the rank score, sequence redundancy of the protein chains was considered. Repetition of the same annotation profile due to the inclusion of chains identical in primary sequence ultimately causes such chains to be ranked unduly higher in the search results. A study by Devos and Valencia demonstrated that protein chains with as high as 95% identity can have a different annotation profiles [58]. To remove chains redundant in primary sequence but limit the loss of annotations, chains were considered redundant if they were identical in sequence [33]. As discussed in the “Materials and methods” section, the representatives of these redundant sequence groups were used to calculate the relevance scores. Through the removal of chains identical in primary sequence, the annotation profile matrix created for a given search became more accurately weighted and when used so generated more accurate relevance scores.
The organization of the text search of the KB-Rank tool application is user-friendly, intuitive, and interactive. As part of the web tool, computer applications access specified annotations only at the required times. The search process itself is implemented in steps that are organized in hierarchical fashion; and each step is run according to user’s request. The organization makes the tool scalable with regard to the further addition of informative annotations from a variety of data sources.
The results of performing a search with the KB-Rank tool include an ordered list of annotation categories and an ordered list of protein structural chains within each category. As each protein structural chain is displayed, links are provided that include a redirection to the corresponding annotation page with chain specific information at the SBKB. At the corresponding page in the SBKB, annotations that are specific to the chain at a resolution below the annotation category can be retrieved. As examples, such links include more specific structural domains contained within the chains, and the differential tissue specific expression patterns that can be found through resources that are linked to the SBKB. In that way, the KB-Rank tool can be used in conjunction with the SBKB to retrieve annotations at different levels of granularity.
Conclusion
The KB-Rank tool provides a means to improve the efficiency and accuracy of searches to identify relevant protein structural chains and functional annotations relevant to a given text query. User search scenarios were described that demonstrate the tool’s utilities for informational and navigational searches. An example informational search was an examination of the protein structures and functional annotations that have a role in the disease asthma. An example navigational search identified potential structures that may be used to further investigate potential treatments for melanoma via a structure based drug design strategy. We demonstrate, through the illustrative examples, how annotations from different data sources were integrated from biomedical resources to enable research. Features of the tool include a staged integration of biomedical text information and the subsequent use of annotations of protein structural chains. It allows the user to effectively identify protein structural chains and annotation categories given a text search regarding protein functional or disease associations.
Abbreviations
- NCBI:
-
National Center for Biotechnology Information
- MMDB:
-
Molecular Modeling Database
- UniProt:
-
Universal Protein Resource
- PDB:
-
Protein Data Bank
- wwPDB:
-
Worldwide Protein Data Bank
- RCSB:
-
Research Collaboratory for Structural Bioinformatics
- PDBe:
-
Protein Data Bank in Europe
- BMRB:
-
Biological Magnetic Resonance Bank
- PDBj:
-
Protein Data Bank Japan
- SBKB:
-
Structural Biology Knowledgebase
- CellMap:
-
The Cancer Cell Map
- INOH:
-
Integrated Network Objects with Hierarchies
- NCI:
-
National Cancer Institute
- PID:
-
Pathway Interaction Database
- BindingDB:
-
The Binding Database
- DrugBank:
-
Open Data Drug and Drug, Target Database
- ChEBI:
-
Chemical entities of biological interest at the European Bioinformatics Institute
- ChEMBL:
-
Chemical database of bioactive drug-like small molecules of the European Molecular Biology Laboratory
- SMPDB:
-
The Small Molecule Pathway Database
- SNPs3D:
-
Single Nucleotide Polymorphisms modeled on protein structures
- OMIM:
-
Online Mendelian Inheritance in Man
- GO:
-
Gene Ontology
- SIFTS:
-
Structure integration with function, taxonomy and sequence
- EC2PDB:
-
Enzyme commission to Protein Data Bank
- CATH:
-
Class architecture topology homology
- SCOP:
-
Structural Classification of Protein
- Pfam:
-
Protein family database
- MeSH:
-
Medical Subject Headings
- KB-Rank:
-
Knowledge Base Ranking Tool
- IL:
-
Interleukin
- SBDD:
-
Structure based drug design
- MEK1:
-
Mitogen activated kinase 1/extracellular-signal-regulated kinase 1 kinase 1
- RAS-RAF-MEK-ERK pathway:
-
RAS-RAF-mitogen-activated protein kinase (MAPK)/extracellular signal-regulated kinase (ERK) kinase (MEK)-ERK
- PDBID:
-
Protein Data Bank identification code
References
Sayers EW, Barrett T, Benson DA, Bryant SH, Canese K, Chetvernin V, Church DM, Dicuccio M, Edgar R, Federhen S, Feolo M, Geer LY, Helmberg W, Kapustin Y, Landsman D, Lipman DJ, Madden TL, Maglott DR, Miller V, Mizrachi I, Ostell J, Pruitt KD, Schuler GD, Sequeira E, Sherry ST, Shumway M, Sirotkin K, Souvorov A, Starchenko G, Tatusova TA, Wagner L, Yaschenko E, Ye J (2009) Database resources of the national center for biotechnology information. Nucleic Acids Res 37:D5–D15
Wang Y, Addess KJ, Chen J, Geer LY, He J, He S, Lu S, Madej T, Marchler-Bauer A, Thiessen PA, Zhang N, Bryant SH (2007) MMDB: annotating protein sequences with Entrez’s 3D-structure database. Nucleic Acids Res 35:D298–D300
Magrane M, Consortium U (2011) UniProt Knowledgebase: a hub of integrated protein data. Database (Oxford) 2011:bar009
Berman H, Henrick K, Nakamura H, Markley JL (2007) The worldwide Protein Data Bank (wwPDB): ensuring a single, uniform archive of PDB data. Nucleic Acids Res 35:D301–D303
Rose PW, Beran B, Bi C, Bluhm WF, Dimitropoulos D, Goodsell DS, Prlic A, Quesada M, Quinn GB, Westbrook JD, Young J, Yukich B, Zardecki C, Berman HM, Bourne PE (2011) The RCSB Protein Data Bank: redesigned web site and web services. Nucleic Acids Res 39:D392–D401
Velankar S, Alhroub Y, Alili A, Best C, Boutselakis HC, Caboche S, Conroy MJ, Dana JM, Van Ginkel G, Golovin A, Gore SP, Gutmanas A, Haslam P, Hirshberg M, John M, Lagerstedt I, Mir S, Newman LE, Oldfield TJ, Penkett CJ, Pineda-Castillo J, Rinaldi L, Sahni G, Sawka G, Sen S, Slowley R, Sousa da Silva AW, Suarez-Uruena A, Swaminathan GJ, Symmons MF, Vranken WF, Wainwright M, Kleywegt GJ (2010) PDBe: Protein Data Bank in Europe. Nucleic Acids Res 39: D402–10
Ulrich EL, Akutsu H, Doreleijers JF, Harano Y, Ioannidis YE, Lin J, Livny M, Mading S, Maziuk D, Miller Z, Nakatani E, Schulte CF, Tolmie DE, Kent Wenger R, Yao H, Markley JL (2008) BioMagResBank. Nucleic Acids Res 36:D402–D408
Kinjo AR, Yamashita R, Nakamura H (2010) PDBj Mine: design and implementation of relational database interface for Protein Data Bank Japan. Database (Oxford) 2010:baq021
Gabanyi MJ, Adams PD, Arnold K, Bordoli L, Carter LG, Flippen-Andersen J, Gifford L, Haas J, Kouranov A, McLaughlin WA, Micallef DI, Minor W, Shah R, Schwede T, Tao YP, Westbrook JD, Zimmerman M, Berman HM (2011) The Structural Biology Knowledgebase: a portal to protein structures, sequences, functions, and methods. J Struct Funct Genomics 12:45–54
Chen C, Huang H, Wu CH (2011) Protein bioinformatics databases and resources. Methods Mol Biol 694:3–24
Page L, Brin S (1998) The anatomy of a large-scale hypertextual Web search engine. Comput Netw ISDN Syst 30:107–117
Karp PD, Ouzounis CA, Moore-Kochlacs C, Goldovsky L, Kaipa P, Ahren D, Tsoka S, Darzentas N, Kunin V, Lopez-Bigas N (2005) Expansion of the BioCyc collection of pathway/genome databases to 160 genomes. Nucleic Acids Res 33:6083–6089
Cerami EG, Gross BE, Demir E, Rodchenkov I, Babur O, Anwar N, Schultz N, Bader GD, Sander C (2011) Pathway Commons, a web resource for biological pathway data. Nucleic Acids Res 39:D685–D690
Fukuda K (2008) INOH pathway database: curation, annotation, integration. InterOntology08 1:47–50
Schaefer CF, Anthony K, Krupa S, Buchoff J, Day M, Hannay T, Buetow KH (2009) PID: the pathway interaction database. Nucleic Acids Res 37:D674–D679
Liu T, Lin Y, Wen X, Jorissen RN, Gilson MK (2007) BindingDB: a web-accessible database of experimentally determined protein-ligand binding affinities. Nucleic Acids Res 35:D198–D201
Wishart DS, Knox C, Guo AC, Cheng D, Shrivastava S, Tzur D, Gautam B, Hassanali M (2008) DrugBank: a knowledgebase for drugs, drug actions and drug targets. Nucleic Acids Res 36:D901–D906
Degtyarenko K, de Matos P, Ennis M, Hastings J, Zbinden M, McNaught A, Alcantara R, Darsow M, Guedj M, Ashburner M (2008) ChEBI: a database and ontology for chemical entities of biological interest. Nucleic Acids Res 36:D344–D350
Gaulton A, Bellis LJ, Bento AP, Chambers J, Davies M, Hersey A, Light Y, McGlinchey S, Michalovich D, Al-Lazikani B, Overington JP (2012) ChEMBL: a large-scale bioactivity database for drug discovery. Nucleic Acids Res 40(Database issue):D1100–D1107
Frolkis A, Knox C, Lim E, Jewison T, Law V, Hau DD, Liu P, Gautam B, Ly S, Guo AC, Xia J, Liang Y, Shrivastava S, Wishart DS (2010) SMPDB: The small molecule pathway database. Nucleic Acids Res 38:D480–D487
Yue P, Melamud E, Moult J (2006) SNPs3D: candidate gene and SNP selection for association studies. BMC Bioinformatics 7:166
McKusick VA (2007) Mendelian Inheritance in Man and its online version, OMIM. Am J Hum Genet 80:588–604
Hamosh A, Scott AF, Amberger JS, Bocchini CA, McKusick VA (2005) Online Mendelian Inheritance in Man (OMIM), a knowledgebase of human genes and genetic disorders. Nucleic Acids Res 33:D514–D517
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, Harris MA, Hill DP, Issel-Tarver L, Kasarskis A, Lewis S, Matese JC, Richardson JE, Ringwald M, Rubin GM, Sherlock G (2000) Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25:25–29
Velankar S, McNeil P, Mittard-Runte V, Suarez A, Barrell D, Apweiler R, Henrick K (2005) E-MSD: an integrated data resource for bioinformatics. Nucleic Acids Res 33:D262–D265
Bairoch A (2000) The ENZYME database in 2000. Nucleic Acids Res 28:304–305
Orengo CA, Pearl FM, Bray JE, Todd AE, Martin AC, Lo Conte L, Thornton JM (1999) The CATH Database provides insights into protein structure/function relationships. Nucleic Acids Res 27:275–279
Murzin AG, Brenner SE, Hubbard T, Chothia C (1995) SCOP: a structural classification of proteins database for the investigation of sequences and structures. J Mol Biol 247:536–540
Finn RD, Mistry J, Tate J, Coggill P, Heger A, Pollington JE, Gavin OL, Gunasekaran P, Ceric G, Forslund K, Holm L, Sonnhammer EL, Eddy SR, Bateman A (2010) The Pfam protein families database. Nucleic Acids Res 38:D211–D222
Ye Y, Godzik A (2003) Flexible structure alignment by chaining aligned fragment pairs allowing twists. Bioinformatics 19(Suppl 2):ii246–ii255
Prlic A, Bliven S, Rose PW, Bluhm WF, Bizon C, Godzik A, Bourne PE (2010) Pre-calculated protein structure alignments at the RCSB PDB website. Bioinformatics 26:2983–2985
Halperin I, Glazer DS, Wu S, Altman RB (2008) The FEATURE framework for protein function annotation: modeling new functions, improving performance, and extending to novel applications. BMC Genomics 9(Suppl 2):S2
Julfayev ES, McLaughlin RJ, Tao YP, McLaughlin WA (2011) A new approach to assess and predict the functional roles of proteins across all known structures. J Struct Funct Genomics 12:9–20
Berman HM, Westbrook JD, Gabanyi MJ, Tao W, Shah R, Kouranov A, Schwede T, Arnold K, Kiefer F, Bordoli L, Kopp J, Podvinec M, Adams PD, Carter LG, Minor W, Nair R, Baer JL (2008) The protein structure initiative structural genomics knowledgebase. Nucleic Acids Res 37:D365–D368
Pachev AS (2007) Understanding MySQL internals. Sebastopol, CA, O’Reilly, Beijing
Bucher P (1990) Weight matrix descriptions of four eukaryotic RNA polymerase II promoter elements derived from 502 unrelated promoter sequences. J Mol Biol 212:563–578
Laskowski RA, Chistyakov VV, Thornton JM (2005) PDBsum more: new summaries and analyses of the known 3D structures of proteins and nucleic acids. Nucleic Acids Res 33:D266–D268
Arnold K, Kiefer F, Kopp J, Battey JN, Podvinec M, Westbrook JD, Berman HM, Bordoli L, Schwede T (2009) The protein model portal. J Struct Funct Genomics 10:1–8
Borish LC, Nelson HS, Lanz MJ, Claussen L, Whitmore JB, Agosti JM, Garrison L (1999) Interleukin-4 receptor in moderate atopic asthma. A phase I/II randomized, placebo-controlled trial. Am J Respir Crit Care Med 160:1816–1823
Chatila TA (2004) Interleukin-4 receptor signaling pathways in asthma pathogenesis. Trends Mol Med 10:493–499
Araujo BB, Dolhnikoff M, Silva LF, Elliot J, Lindeman JH, Ferreira DS, Mulder A, Gomes HA, Fernezlian SM, James A, Mauad T (2008) Extracellular matrix components and regulators in the airway smooth muscle in asthma. Eur Respir J 32:61–69
Bai TR, Cooper J, Koelmeyer T, Pare PD, Weir TD (2000) The effect of age and duration of disease on airway structure in fatal asthma. Am J Respir Crit Care Med 162:663–669
Abehsira-Amar O, Gibert M, Joliy M, Theze J, Jankovic DL (1992) IL-4 plays a dominant role in the differential development of Tho into Th1 and Th2 cells. J Immunol 148:3820–3829
Levine SJ, Wenzel SE (2010) Narrative review: the role of Th2 immune pathway modulation in the treatment of severe asthma and its phenotypes. Ann Intern Med 152:232–237
Anderson AC (2003) The process of structure-based drug design. Chem Biol 10:787–797
Pitt WR, Higueruelo AP, Groom CR (2009) Structural bioinformatics in drug discovery. In: Gu J, Bourne PE (eds) Structural bioinformatics, 2nd edn. Wiley-Blackwell, Hoboken
Iverson C, Larson G, Lai C, Yeh LT, Dadson C, Weingarten P, Appleby T, Vo T, Maderna A, Vernier JM, Hamatake R, Miner JN, Quart B (2009) RDEA119/BAY 869766: a potent, selective, allosteric inhibitor of MEK1/2 for the treatment of cancer. Cancer Res 69:6839–6847
Wang D, Boerner SA, Winkler JD, Lorusso PM (2007) Clinical experience of MEK inhibitors in cancer therapy. Biochim Biophys Acta 1773:1248–1255
Adjei AA (2001) Blocking oncogenic Ras signaling for cancer therapy. J Natl Cancer Inst 93:1062–1074
Flaherty KT, Puzanov I, Kim KB, Ribas A, McArthur GA, Sosman JA, O’Dwyer PJ, Lee RJ, Grippo JF, Nolop K, Chapman PB (2010) Inhibition of mutated, activated BRAF in metastatic melanoma. N Engl J Med 363:809–819
Eisen T, Ahmad T, Flaherty K, Gore M, Kaye S, Marais R, Gibbens I, Hackett S, James M, Schuchter L (2006) Sorafenib in advanced melanoma: a phase II randomised discontinuation trial analysis. Br J Cancer 95:581–586
Arkenau HT, Kefford R, Long GV (2011) Targeting BRAF for patients with melanoma. Br J Cancer 104:392–398
Ji Z, Flaherty KT, Tsao H (2011) Targeting the RAS pathway in melanoma. Trends Mol Med 18:27–35
Page L, Brin S, Motwani R, Winograd T (1998) The PageRank citation ranking: bringing order to the web. Stanford Digital Libraries Working Paper
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Consortium U (2011) Ongoing and future developments at the Universal Protein Resource. Nucleic Acids Res 39:D214–D219
Devos D, Valencia A (2000) Practical limits of function prediction. Proteins 41:98–107
Acknowledgments
We thank Raship Shah at Rutgers University for a template of cascading style sheets that are used in the tool’s web design. We thank John Westbrook, Margaret Gabanyi, and Helen Berman at Rutgers University who provided assistance with access to data collections of the SBKB and computer code for text search over primary citation text fields. We are grateful to Dr. Peter Karp and the rest of the BioCyc team. They provided the necessary access to the protein and pathway data for each organism. Support was provided in part by a grant from the National Institute of General Medical Sciences [grant number 5U01 GM093324-02].
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Julfayev, E.S., McLaughlin, R.J., Tao, YP. et al. KB-Rank: efficient protein structure and functional annotation identification via text query. J Struct Funct Genomics 13, 101–110 (2012). https://doi.org/10.1007/s10969-012-9125-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10969-012-9125-7