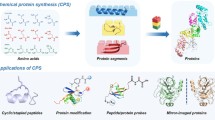
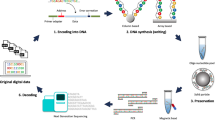
Abstract
A two-step PCR method has been developed for the robust, high-throughput production of linear templates ready for cell-free protein synthesis. The construct made from the cDNA expresses a target protein region with N- and/or C-terminal tags. The procedure consists only of mixing, dilution, and PCR steps, and is free from cloning and purification steps. In the first step of the two-step PCR, a target region within the coding sequence is amplified using two gene-specific forward and reverse primers, which contain the linker sequences and the terminal sequences of the target region. The second PCR concatenates the first PCR product with the N- and C-terminal double-stranded fragments, which contain the linker sequences as well as the sequences for the tag(s) and the initiation and termination, respectively, for T7 transcription and ribosomal translation, and amplifies it with the universal primer. Proteins can be fused with a variety of tags, such as natural poly-histidine, glutathione-S-transferase, maltose-binding protein, and/or streptavidin-binding peptide. The two-step PCR method was successfully applied to 42 human target protein regions with various GC contents (38–77%). The robustness of the two-step PCR method against possible fluctuations of experimental conditions in practical use was explored. The second PCR product was obtained at 60–120 μg/ml, and was used without purification as a template at a concentration of 2–4 μg/ml in an Escherichia coli coupled transcription-translation system. This combination of two-step PCR with cell-free protein synthesis is suitable for the rapid production of proteins in milligram quantities for genome-scale studies.







Similar content being viewed by others
Abbreviations
- GST:
-
Glutathione-S-transferase
- MBP:
-
E. coli maltose binding protein
- SBP:
-
Streptavidin binding peptide
- NHis:
-
Natural poly-histidine
- TEV:
-
Tobacco etch virus
- DMSO:
-
Dimethyl sulfoxide
- HA:
-
Hemagglutinin
References
Nameki N, Tochio N, Koshiba S, Inoue M, Yabuki T, Aoki M, Seki E, Matsuda T, Fujikura Y, Saito M, Ikari M, Watanabe M, Terada T, Shirouzu M, Yoshida M, Hirota H, Tanaka A, Hayashizaki Y, Guntert P, Kigawa T, Yokoyama S (2005) Protein Sci 14:756–764
Yamasaki K, Kigawa T, Inoue M, Tateno M, Yamasaki T, Yabuki T, Aoki M, Seki E, Matsuda T, Tomo Y, Hayami N, Terada T, Shirouzu M, Tanaka A, Seki M, Shinozaki K, Yokoyama S (2005) Plant Cell 17:944–956
Li H, Inoue M, Yabuki T, Aoki M, Seki E, Matsuda T, Nunokawa E, Motoda Y, Kobayashi A, Terada T, Shirouzu M, Koshiba S, Lin YJ, Guntert P, Suzuki H, Hayashizaki Y, Kigawa T, Yokoyama S (2005) J Biomol NMR 32:329–334
Yabuki T, Kigawa T, Dohmae N, Takio K, Terada T, Ito Y, Laue ED, Cooper JA, Kainosho M, Yokoyama S (1998) J Biomol NMR 11:295–306
Kurimoto K, Muto Y, Obayashi N, Terada T, Shirouzu M, Yabuki T, Aoki M, Seki E, Matsuda T, Kigawa T, Okumura H, Tanaka A, Shibata N, Kashikawa M, Agata K, Yokoyama S (2005) J Struct Biol 150:58–68
Arai R, Kukimoto-Niino M, Uda-Tochio H, Morita S, Uchikubo-Kamo T, Akasaka R, Etou Y, Hayashizaki Y, Kigawa T, Terada T, Shirouzu M, Yokoyama S (2005) Protein Sci 14:1888–1893
Kigawa T, Yamaguchi-Nunokawa E, Kodama K, Matsuda T, Yabuki T, Matsuda N, Ishitani R, Nureki O, Yokoyama S (2002) J Struct Funct Genomics 2:29–35
Kang SH, Kim DM, Kim HJ, Jun SY, Lee KY, Kim HJ (2005) Biotechnol Prog 21:1412–1419
Ryabova LA, Desplancq D, Spirin AS, Pluckthun A (1997) Nat Biotechnol 15:79–84
Klammt C, Schwarz D, Lohr F, Schneider B, Dotsch V, Bernhard F (2006) FEBS J 273:4141–4153
Yokoyama S, Matsuo Y, Hirota H, Kigawa T, Shirouzu M, Kuroda Y, Kurumizaka H, Kawaguchi S, Ito Y, Shibata T, Kainosho M, Nishimura Y, Inoue Y, Kuramitsu S (2000) Prog Biophys Mol Biol 73:363–376
Endo Y, Sawasaki T (2004) J Struct Funct Genomics 5:45–57
Sawasaki T, Ogasawara T, Morishita R, Endo Y (2002) Proc Natl Acad Sci USA 99:14652–14657
Woodrow KA, Airen IO, Swartz JR (2006) J Proteome Res 5:3288–3300
Kigawa T, Muto Y, Yokoyama S (1995) J Biomol NMR 6:129–134
Nakai K, Horton P (1999) Trends Biochem Sci 24:34–36
Dougherty WG, Cary SM, Parks TD (1989) Virology 171:356–364
Chaga G, Bochkariov DE, Jokhadze GG, Hopp J, Nelson P (1999) J Chromatogr A 864:247–256
Hirota Y, Katsumata A, Takeya T (1990) Nucleic Acids Res 18:6432
Keefe AD, Wilson DS, Seelig B, Szostak JW (2001) Protein Expr Purif 23:440–446
Matsuda T, Kigawa T, Koshiba S, Inoue M, Aoki M, Yamasaki K, Seki M, Shinozaki K, Yokoyama S (2006) J Struct Funct Genomics 7:93–100
Lin-Goerke JL, Robbins DJ, Burczak JD (1997) Biotechniques 23:409–412
Kigawa T, Yabuki T, Yoshida Y, Tsutsui M, Ito Y, Shibata T, Yokoyama S (1999) FEBS Lett 442:15–19
Nakano H, Kobayashi K, Ohuchi S, Sekiguchi S, Yamane T (2000) J Biosci Bioeng 90:456–458
Esposito D, Chatterjee DK (2006) Curr Opin Biotechnol 17:353–358
Labaer J, Ramachandran N (2005) Curr Opin Chem Biol 9:14–19
Griffiths AD, Tawfik DS (2006) Trends Biotechnol 24:395–402
Waldo GS, Standish BM, Berendzen J, Terwilliger TC (1999) Nat Biotechnol 17:691–695
Acknowledgements
The authors thank Dr. Daisuke Kiga (Tokyo Institute of Technology) and Dr. Yasuaki Kawarasaki (University of Shizuoka) for helpful discussions; Dr. Takayoshi Matsuda (RIKEN) for providing HSQC spectra of Ras mutants; Natsuko Matsuda, Natsumi Suzuki, and Yikkiko Fujikura for their technical assistance; and Tomoko Nakayama and Azusa Ishii for expert secretarial assistance. This work was supported by the RIKEN Structural Genomics/Proteomics Initiative (RSGI), the National Project on Protein Structural and Functional Analysis, Ministry of Education, Culture, Sports, Science and Technology of Japan.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Yabuki, T., Motoda, Y., Hanada, K. et al. A robust two-step PCR method of template DNA production for high-throughput cell-free protein synthesis. J Struct Funct Genomics 8, 173–191 (2007). https://doi.org/10.1007/s10969-007-9038-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10969-007-9038-z