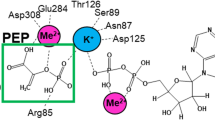
Saccharomyces cerevisiae phosphoenolpyruvate (PEP) carboxykinase catalyzes the reversible formation of oxaloacetate and adenosine triphosphate from PEP, adenosine diphosphate and carbon dioxide, and uses Mn2+ as the activating metal ion. Comparison with the crystalline structure of homologous Escherichia coli PEP carboxykinase [Tari et al. (1997) Nature Struct. Biol. 4, 990–994] shows that Lys213 is one of the ligands to Mn2+ at the enzyme active site. Coordination of Mn2+ to a lysyl residue is not common and suggests a low pK a value for the ε-NH2 group of Lys213. In this work, we evaluate the role of neighboring Phe216 in contributing to provide a low polarity microenvironment suitable to keep the ε-NH2 of Lys213 in the unprotonated form. Mutation Phe216Tyr shows that the introduction of a hydroxyl group in the lateral chain of the residue produces a substantial loss in the enzyme affinity for Mn2+, suggesting an increase of the pK a of Lys213. In agreement with this interpretation, theoretical calculations indicate an alkaline shift of 2.8 pH units in the pK a of the ε-amino group of Lys213 upon Phe216Tyr mutation.


Similar content being viewed by others
Abbreviations
- CD:
-
circular dichroism spectroscopy
- HEPES:
-
N-(2-hydroxyethyl)piperazine-N′-2(ethanesulfonic acid)
- HPLC:
-
high-performance liquid chromatography
- MD:
-
molecular dynamics
- MOPS:
-
3-(N-morpholino)propanesulfonic acid
- OAA:
-
oxaloacetic acid
- PEP:
-
phosphoenolpyruvate
- SDS-PAGE:
-
sodium dodecylsulfate-polyacrylamide gel electrophoresis
References
Cotelesage J.J., Prasad L., Zeikus J.G., Laivenieks M., Delbaere L.T. (2005) Int. J. Biochem. Cell Biol. 37:1829–1837
Dunten P., Belunis C., Crowther R., Hollfelder K., Kammlott U., Levin W., Michel H., Ramsey G.B., Swain A., Weber D., Wertheimer S.J. (2002) J. Mol. Biol. 316:257–264
González-Nilo F.D., Krautwurst H., Yévenes A., Cardemil E., Cachau R. (2002) Biochim. Biophys. Acta 1599:65–71
Holyoak T.,Sullivan S.M., Nowak T. (2006) Biochemistry 45:8254–8263
Jacob L.R., Vollert H., Rose M., Entian K.D., Bartunik L.J., Bartunik H. D. (1992) J. Chromatogr. 625:47–54
Krautwurst H., Encinas M.V., Marcus F., Latshaw S.P., Kemp R.G., Frey P.A., Cardemil E. (1995). Biochemistry 34:6382–6388
Krautwurst H., Bazaes S., González F.D., Jabalquinto A.M., Frey P.A., Cardemil E. (1998) Biochemistry 37:6295–6302
Krautwurst H., Roschzttardtz H., Bazaes S., Gonzalez-Nilo F.D., Nowak T., Cardemil E. (2002) Biochemistry 41:12763–12770
Lee M.H., Hebda C.A., Nowak T. (1981) J. Biol. Chem. 256:12793–12801
Lee F.S., Chu Z.T., Warshel A. (1993) J. Comput. Chem. 14:161–185
Llanos L., Briones R., Yévenes A., González-Nilo F.D., Frey P.A., Cardemil E. (2001). FEBS Letters 493:1–5
Martel, A.E., and Smith, R.M. (1998). NIST standard references database 46, version 5.0
Matte A, Tari L.W., Goldie H., Delbaere L.T.J. (1997) J. Biol. Chem. 272:8105–8108
Müller M., Müller H., Holzer H. (1981) J. Biol. Chem. 256:723–727
Perella F.W. (1988) Anal. Biochem. 174:437–447
Sham Y.Y., Chu Z.T., Warshel A. (1997) J. Phys. Chem. 101:4458–4472
Sugahara M., Ohshima N., Ukita Y., Sugahara M., Kunishima N. (2005) Acta Cryst. D61:1500–1507
Tari L.W., Matte A., Goldie H., Delbaere L.T.J. (1997) Nature Struct. Biol. 4:990–994
Trapani S., Linss J., Goldenberg S., Fische H., Craievich A.F., Oliva G. (2001) J. Mol. Biol. 313:1059–1072
Utter M.F., Kolenbrander H.M. (1972) The Enzymes. 3rd, Academic Press, New York, 6:117–168
Yévenes A., Espinoza R., Rivas-Pardo J.A., Villarreal J.M., González-Nilo F.D., Cardemil E. (2006) Biochimie 88:663–672
Acknowledgments
This work was supported by research grants FONDECYT 2010041 (AY) and 1030760 (EC). CD experiments were carried out at the Biophysics Instrumentation Facility of the University of Wisconsin-Madison, which was established by funding from NSF (BIR-9512577), NIH (S10RR3790), and the University of Wisconsin.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yévenes, A., González-Nilo, F. & Cardemil, E. Relevance of phenylalanine 216 in the affinity of Saccharomyces cerevisiae phosphoenolpyruvate carboxykinase for Mn(II). Protein J 26, 135–141 (2007). https://doi.org/10.1007/s10930-006-9054-z
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10930-006-9054-z