Abstract
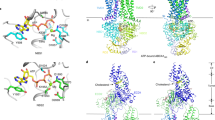
Many of the 48 or 49 human ABC proteins are involved in lipid homeostasis and in defence against hydrophobic substances in food and the environment. Defects in their functions cause various diseases, suggesting that they play very important roles in human health; however, the mechanism of how they handle enormous numbers of hydrophobic compounds with various structures and molecular weights, or phospholipids and cholesterol, major components of cellular membranes, is not known. We compared the functions of drug-transporting and lipid-transporting ABC proteins, and found that (1) ABC proteins, either lipid or drug transporters, have a similar substrate binding site which recognizes PL and cholesterol, or drugs and cholesterol; (2) Cholesterol in membranes binds to various ABC proteins together with PL or drugs, and plays an important role in substrate recognition, especially by ABCB1/MDR1, where cholesterol fills the empty space in the substrate binding site when small drugs bind to it. ABC proteins exert very flexible substrate recognition, i.e., one-to-many interaction rather than the conventional rigid one-to-one interaction. We propose calling the mechanism the “cholesterol fill-in model”.
Similar content being viewed by others
References
Bodzioch M, Orso E, Klucken J, Langmann T, Bottcher A, Diederich W, Drobnik W, Barlage S, Buchler C, Porsch-Ozcurumez M, Kaminski W, Hahmann H, Oette K, Rothe G, Aslanidis C, Lackner K, Schmitz G (1999) Nat Genet 22:347–351
Brooks-Wilson A, Marcil M, Clee S, Zhang L, Roomp K, van Dam M, Yu L, Brewer C, Collins J, Molhuizen H, Loubser O, Ouelette B, Fichter K, Ashbourne-Excoffon K, Sensen C, Scherer S, Mott S, Denis M, Martindale D, Frohlich J, Morgan K, Koop B, Pimstone S, Kastelein J, Hayden M (1999) Nat Genet 22:336–345
Fielding PE, Nagao K, Hakamata H, Chimini G, Fielding CJ (2000) Biochemistry 39:14113–14120
Fitzgerald ML, Morris AL, Rhee JS, Andersson LP, Mendez AJ, Freeman MW (2002) J Biol Chem 277:33178–33187
Hanada K, Kumagai K, Yasuda S, Miura Y, Kawano M, Fukasawa M, Nishijima M (2003) Nature 426:803–809
Hayashi M, Abe-Dohmae S, Okazaki M, Ueda K, Yokoyama S (2005) J Lipid Res 46:1703–1711
Kimura Y, Kioka N, Kato H, Matsuo M, Ueda K (2007) Biochem J 401:597–605
Kino K, Taguchi Y, Yamada K, Komano T, Ueda K (1996) FEBS Lett 399:29–32
Kobayashi A, Takanezawa Y, Hirata T, Shimizu Y, Misasa K, Kioka N, Arai H, Ueda K, Matsuo M (2006) J Lipid Res 47:1791–1802
Koseki M, Hirano KI, Masuda D, Ikegami C, Tanaka M, Ota A, Sandoval JC, Nakagawa-Toyama Y, Sato SB, Kobayashi T, Shimada Y, Ohno-Iwashita Y, Matsuura F, Shimomura I, Yamashita S (2007) J Lipid Res 48:299–306
Landry YD, Denis M, Nandi S, Bell S, Vaughan AM, Zha X (2006) J Biol Chem 281:36091–36101
Loo TW, Bartlett MC, Clarke DM (2006) Biochem J 22:22
Morita SY, Kobayashi A, Takanezawa Y, Kioka N, Handa T, Arai H, Matsuo M, Ueda K (2007) Hepatology 46:188–199
Nagao K, Takahashi K, Hanada K, Kioka N, Matsuo M, Ueda K (2007) J Biol Chem 282:14868–14874
Ruetz S, Gros P (1994) Cell 77:1071–1081
Rust S, Rosier M, Funke H, Real J, Amoura Z, Piette J, Deleuze J, Brewer H, Duverger N, Denefle P, Assmann G (1999) Nat Genet 22:352–355
Sano O, Kobayashi A, Nagao K, Kumagai K, Kioka N, Hanada K, Ueda K, Matsuo M (2007) J Lipid Res 48:2377–2384
Smith AJ, van Helvoort A, van Meer G, Szabo K, Welker E, Szakacs G, Varadi A, Sarkadi B, Borst P (2000) J Biol Chem 275:23530–23539
Taguchi Y, Kino K, Morishima M, Komano T, Kane SE, Ueda K (1997a) Biochemistry 36:8883–8889
Taguchi Y, Morishima M, Komano T, Ueda K (1997b) FEBS Lett 413:142–146
Takahashi K, Kimura Y, Kioka N, Matsuo M, Ueda K (2006) J Biol Chem 281:10760–10768
Tanaka AR, Ikeda Y, Abe-Dohmae S, Arakawa R, Sadanami K, Kidera A, Nakagawa S, Nagase T, Aoki R, Kioka N, Amachi T, Yokoyama S, Ueda K (2001) Biochem Biophys Res Commun 283:1019–1025
Tanaka AR, Abe-Dohmae S, Ohnishi T, Aoki R, Morinaga G, Okuhira KI, Ikeda Y, Kano F, Matsuo M, Kioka N, Amachi T, Murata M, Yokoyama S, Ueda K (2003) J Biol Chem 278:8815–8819
Ueda K, Cardarelli C, Gottesman MM, Pastan I (1987) Proc Natl Acad Sci U S A 84:3004–3008
van der Bliek AM, Kooiman PM, Schneider C, Borst P (1988) Gene 71:401–411
Wang N, Silver D, Costet P, Tall A (2000) J Biol Chem 275:33053–33058
Wang N, Silver DL, Thiele C, Tall AR (2001) J Biol Chem 276:23742–23747
Wang N, Lan D, Gerbod-Giannone M, Linsel-Nitschke P, Jehle AW, Chen W, Martinez LO, Tall AR (2003) J Biol Chem 278:42906–42912
Wang J, Sun F, Zhang DW, Ma Y, Xu F, Belani JD, Cohen JC, Hobbs HH, Xie XS (2006) J Biol Chem 281:27894–27904
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kimura, Y., Kodan, A., Matsuo, M. et al. Cholesterol fill-in model: mechanism for substrate recognition by ABC proteins. J Bioenerg Biomembr 39, 447–452 (2007). https://doi.org/10.1007/s10863-007-9109-7
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10863-007-9109-7