Abstract
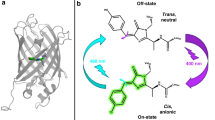
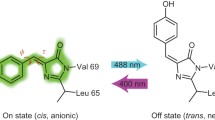
Dronpa is a green fluorescent protein homologue with a photochromic property. A green laser illumination reversibly converts Dronpa from a green-emissive bright state to a non-emissive dark state, and ultraviolet illumination converts it to the bright state. We have employed solution NMR to understand the underlying molecular mechanism of the photochromism. The detail characterization of Dronpa is hindered as it is metastable in the dark state and spontaneously converts to the bright state. To circumvent this issue, we have designed in magnet laser illumination device. By combining the device with a 150-mW argon laser at 514.5 nm, we have successfully converted and maintained Dronpa in the dark state in the NMR tube by continuous illumination during the NMR experiments. We have employed direct-detection of 13C nuclei from the carbon skeleton of the chromophore for detailed characterization of chromophore in both states of Dronpa by using the Bruker TCI cryoprobe. The results from NMR data have provided direct evidence of the double bond formation between Cα and Cβ of Y63 in the chromophore, the β-barrel structure in solution, and the ionized and protonated state of Y63 hydroxyl group in the bright and dark states, respectively. These studies have also revealed that a part of β-barrel around the chromophore becomes polymorphic only in the dark state, which may be critical to make the fluorescence dim by increasing the contribution of non-emissive vibrational relaxation pathways.







Similar content being viewed by others
References
Akke M, Palmer AG III (1996) Monitoring macromolecular motions on microsecond to millisecond time scales by R1ρ–R1 constant relaxation time NMR spectroscopy. J Am Chem Soc 118:911–912
Ando R, Mizuno H, Miyawaki A (2004) Regulated fast nucleocytoplasmic shuttling observed by reversible protein highlighting. Science 306:1370–1373
Andresen M, Stiel AC, Trowitzsch S, Weber G, Eggeling C, Wahl MC, Hell SW, Jakobs S (2007) Structural basis for reversible photoswitching in Dronpa. Proc Natl Acad Sci USA 104:13005–13009
Bargon J, Fisher H, Johnsen U (1967) Kernresonanz-emissionslinein wahrend rascher radikalreaktionen. I. aufnahmeverfahren und beispiele. Z Naturforsch A22:1551–1555
Bermel W, Felli IC, Kümmerle R, Pierattelli R (2008) 13C Direct-detection biomolecular NMR. Concepts Magn Reson 32A:183–200
Bertini I, Felli IC, Kümmerle R, Moskau D, Pierattelli R (2004) 13C–13C NOESY: an attractive alternative for studying large macromolecules. J Am Chem Soc 126:464–465
Bradbury JH, Ramesh V (1985) 1H n.m.r. studies of insulin. Assignment of resonances and properties of tyrosine residues. Biochem J 229:731–737
Craven CJ, Derix NM, Hendriks J, Boelens R, Hellingwerf KJ, Kaptein R (2000) Probing the nature of the blue-shifted intermediate of photoactive yellow protein in solution by NMR: hydrogen–deuterium exchange data and pH studies. Biochemistry 39:14392–14399
Dedecker P, Hotta J, Flors C, Sliwa M, Uji-I H, Roeffaers MB, Ando R, Mizuno H, Miyawaki A, Hofkens J (2007) Subdiffraction imaging through the selective donut-mode depletion of thermally stable photoswitchable fluorophores: numerical analysis and application to the fluorescent protein Dronpa. J Am Chem Soc 129:16132–16141
Egan W, Shindo H, Cohen JS (1978) On the tyrosine residues of ribonuclease A. J Biol Chem 253:16–17
Farrow NA, Muhandiram R, Singer AU, Pascal SM, Kay CM, Gish G, Shoelson SE, Pawson T, Forman-Kay JD, Kay LE (1994) Backbone dynamics of a free and phosphopeptide-complexed Src homology 2 domain studied by 15N NMR relaxation. Biochemistry 33:5984–6003
Flors C, Hotta J, Uji-I H, Dedecker P, Ando R, Mizuno H, Miyawaki A, Hofkens J (2007) A stroboscopic approach for fast photoactivation-localization microscopy with Dronpa mutants. J Am Chem Soc 129:13970–13977
Harper SM, Neil LC, Gardner KH (2003) Structural basis of a phototropin light switch. Science 301:1541–1544
Harper SM, Neil LC, Day IJ, Hore PJ, Gardner KH (2004) Conformational changes in a photosensory LOV domain monitored by time-resolved NMR spectroscopy. J Am Chem Soc 126:3390–3391
Kaptein R (1982) In: Berliner LJ, Reuben J (eds) Biological magnetic resonance, vol 4. Plenum Press, New York, pp 145–191
Kuhn T, Schwalbe H (2000) Monitoring the kinetics of ion-dependent protein folding by time-resolved NMR spectroscopy at atomic resolution. J Am Chem Soc 122:6169–6174
Kwon OY, Kwon IC, Song HK, Jeon H (2008) Real-time imaging of NF-AT nucleocytoplasmic shuttling with a photoswitchable fluorescence protein in live cells. Biochim Biophys Acta 1780:1403–1407
Mal TK, Masutomi Y, Zheng L, Nakata Y, Ohta H, Nakatani Y, Kokubo T, Ikura M (2004) Structural and functional characterization on the interaction of yeast TFIID subunit TAF1 with TATA-binding protein. J Mol Biol 339:681–693
Mizuno H, Mal TK, Wälchli M, Kikuchi A, Fukano T, Ando R, Jeyakanthan J, Taka J, Shiro Y, Ikura M, Miyawaki A (2008) Light-dependent regulation of structural flexibility in a photochromic fluorescent protein. Proc Natl Acad Sci USA 105:9927–9932
Mizuno H, Dedecker P, Ando R, Fukano T, Hofkens J, Miyawaki A (2010) Higher resolution in localization microscopy by slower switching of a photochromic protein. Photochem Photobiol Sci 9:239–248
Mok KH, Hore PJ (2004) Photo-CIDNP NMR methods for studying protein folding. Methods 34:75–87
Nam KH, Kwon OY, Sugiyama K, Lee WH, Kim YK, Song HK, Kim EE, Park SY, Jeon H, Hwang KY (2007) Structural characterization of the photoswitchable fluorescent protein Dronpa-C62S. Biochem Biophys Res Commun 354:962–967
Rubinstenn G, Vuister GW, Mulder FA, Düx PE, Boelens R, Hellingwerf KJ, Kaptein R (1998) Structural and dynamic changes of photoactive yellow protein during its photocycle in solution. Nat Struct Biol 5:568–570
Rubinstenn G, Vuister GW, Zwanenburg N, Hellingwerf KJ, Boelens R, Kaptein R (1999) NMR experiments for the study of photointermediates: application to the photoactive yellow protein. J Magn Reson 137:443–447
Scheffler JE, Cottrell CE, Berliner LJ (1985) An inexpensive, versatile sample illuminator for photo-CIDNP on any NMR spectrometer. J Magn Reson 63:199–201
Shroff H, Galbraith CG JA, White Helen, Gillette J, Olenych S, Davidson MW, Betzig E (2007) Dual-color superresolution imaging of genetically expressed probes within individual adhesion complexes. Proc Natl Acad Sci USA 104:20308–20313
Spera S, Bax A (1991) Empirical correlation between protein backbone conformation and Ca and Cb nuclear magnetic resonance chemical shifts. J Am Chem Soc 113:5490–5492
Stiel AC, Trowitzsch S, Weber G, Andresen M, Eggeling C, Hell SW, Jakobs S, Wahl MC (2007) 1.8 Å bright-state structure of the reversibly switchable fluorescent protein Dronpa guides the generation of fast switching variants. Biochem J 402:35–42
Ward HR, Lawler RG (1967) Nuclear magnetic resonance emission and enhanced apsorption in rapid organometallic reactions. J Am Chem Soc 89:5518–5519
Wilbur DJ, Allerhand A (1976) Titration behavior of individual tyrosine residues of myoglobins from sperm whale, horse, and red kangaroo. J Biol Chem 251:5187–5194
Wilmann PG, Turcic K, Battad JM, Wilce MC, Devenish RJ, Prescott M, Rossjohn J (2006) The 1.7 Å crystal structure of Dronpa: a photoswitchable green fluorescent protein. J Mol Biol 364:213–224
Wishart DS, Sykes BD (1994) The 13C chemical-shift index: a simple method for the identification of protein secondary structure using 13C chemical-shift data. J Biomol NMR 4:171–180
Acknowledgments
We acknowledge Dr. R. Kato for valuable advice. We thank Dr. C. Marshall for careful reading of the manuscript, and Dr. R. Ando, Ms. K.I. Tong, Ms. K. Otsuki, and Mr. M. Usui for technical assistance. This work was partly supported by grants from the Human Frontier Science Program, Molecular Ensemble Program at RIKEN, Japan MEXT Grant-in-Aid for Scientific Research on priority areas, Japan MEXT and Japan Society for the Promotion of Science for Grants-in-Aid for Scientific Research B, and Canadian Institutes for Health Research.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Mizuno, H., Mal, T.K., Wälchli, M. et al. Molecular basis of photochromism of a fluorescent protein revealed by direct 13C detection under laser illumination. J Biomol NMR 48, 237–246 (2010). https://doi.org/10.1007/s10858-010-9453-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-010-9453-5