Abstract
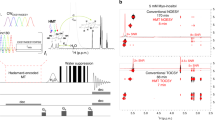
The NMR spectra of nucleic acids suffer from severe peak overlap, which complicates resonance assignments. 4D NMR experiments can overcome much of the degeneracy in 2D and 3D spectra; however, the linear increase in acquisition time with each new dimension makes it impractical to acquire high-resolution 4D spectra using standard Fourier transform (FT) techniques. The filter diagonalization method (FDM) is a numerically efficient algorithm that fits the entire multi-dimensional time-domain data to a set of multi-dimensional oscillators. Selective 4D constant-time HCCH-COSY experiments that correlate the H5–C5–C6–H6 base spin systems of pyrimidines or the H1′–C1′–C2′–H2′ spin systems of ribose sugars were acquired on the 13C-labeled iron responsive element (IRE) RNA. FDM-processing of these 4D experiments recorded with only 8 complex points in the indirect dimensions showed superior spectral resolution than FT-processed spectra. Practical aspects of obtaining optimal FDM-processed spectra are discussed. The results here demonstrate that FDM-processing can be used to obtain high-resolution 4D spectra on a medium sized RNA in a fraction of the acquisition time normally required for high-resolution, high-dimensional spectra.







Similar content being viewed by others
References
Addess KJ, Basilion JP, Klausner RD, Rouault TA, Pardi A (1997) Structure and dynamics of the iron responsive element RNA: Implications for binding of the RNA by iron regulatory binding proteins. J Mol Biol 274:72–83
Armstrong GS, Cano KE, Mandelshtam VA, Shaka AJ, Bendiak B (2004) Rapid 3D NMR using the filter diagonalization method: application to oligosaccharides derivatized with 13C-labeled acetyl groups. J Magn Reson 170:156–163
Armstrong GS, Mandelshtam VA, Shaka AJ, Bendiak B (2005) Rapid high-resolution four-dimensional NMR spectroscopy using the filter diagonalization method and its advantages for detailed structural elucidation of oligosaccharides. J Magn Reson 173:160–168
Bax A, Clore GM, Driscoll PC, Gronenborn AM, Ikura M, Kay LE (1990) Practical aspects of proton carbon carbon proton 3-dimensional correlation spectroscopy of C-13-labeled proteins. J Magn Reson 87:620–627
Bruschweiler R (2004) Theory of covariance nuclear magnetic resonance spectroscopy. J Chem Phys 121:409–414
Bruschweiler R, Zhang FL (2004) Covariance nuclear magnetic resonance spectroscopy. J Chem Phys 120:5253–5260
Chen J, Mandelshtam VA, Shaka AJ (2000) Regularization of the two-dimensional filter diagonalization method: FDM2K. J Magn Reson 146:363–368
Chen J, De Angelis AA, Mandelshtam VA, Shaka AJ (2003) Progress on the two-dimensional filter diagonalization method. An efficient doubling scheme for two-dimensional constant-time NMR. J Magn Reson 162:74–89
Chen J, Nietlispach D, Shaka AJ, Mandelshtam VA (2004) Ultra-high resolution 3D NMR spectra from limited-size data sets. J Magn Reson 169:215–224
Coggins BE, Zhou P (2006) Polar Fourier transforms of radially sampled NMR data. J Magn Reson 182:84–95
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293
Emsley L, Bodenhausen G (1990) Phase-shifts induced by transient Bloch-Siegert effects in NMR. Chem Phys Lett 168:297–303
Geen H, Freeman R (1991) Band-selective radiofrequency pulses. J Magn Reson 93:93–141
Goddard TD, Kneller DG (1998) SPARKY 3. University of California, San Francisco
Hu H, Van QN, Mandelshtam VA, Shaka AJ (1998) Reference deconvolution, phase correction, and line listing of NMR spectra by the 1D filter diagonalization method. J Magn Reson 134:76–87
Hu H, De Angelis AA, Mandelshtam VA, Shaka AJ (2000) The multidimensional filter diagonalization method. J Magn Reson 144:357–366
Hyberts SG, Heffron GJ, Tarragona NG, Solanky K, Edmonds KA, Luithardt H, Fejzo J, Chorev M, Aktas H, Colson K, Falchuk KH, Halperin JA, Wagner G (2007) Ultrahigh-resolution (1)H-(13)C HSQC spectra of metabolite mixtures using nonlinear sampling and forward maximum entropy reconstruction. J Am Chem Soc 129:5108–5116
Jeschke G, Mandelshtam VA, Shaka AJ (1999) Pure absorption electron spin echo envelope modulation spectra by using the filter-diagonalization method for harmonic inversion. J Magn Reson 137:221–230
Kay LE, Ikura M, Bax A (1990) Proton proton correlation via carbon–carbon couplings––a 3-dimensional NMR approach for the assignment of aliphatic resonances in proteins labeled with C-13. J Am Chem Soc 112:888–889
Kay LE, Ikura M, Zhu G, Bax A (1991) 4-Dimensional heteronuclear triple-resonance NMR of isotopically enriched proteins for sequential assignment of backbone atoms. J Magn Reson 91:422–428
Kim S, Szyperski T (2003) GFT NMR, a new approach to rapidly obtain precise high-dimensional NMR spectral information. J Am Chem Soc 125:1385–1393
Kim S, Szyperski T (2004) GFT NMR experiments for polypeptide backbone and 13Cβ chemical shift assignment. J Biomol NMR 28:117–130
Kupce E, Freeman R (2003) Projection-reconstruction of three-dimensional NMR spectra. J Am Chem Soc 125:13958–13959
Kupce E, Freeman R (2004) Projection-reconstruction technique for speeding up multidimensional NMR spectroscopy. J Am Chem Soc 126:6429–6440
Mandelshtam VA (2000) The multidimensional filter diagonalization method––I. Theory and numerical implementation. J Magn Reson 144:343–356
Mandelshtam VA (2001) FDM: the filter diagonalization method for data processing in NMR experiments. Prog NMR Spec 38:159–196
Mandelshtam VA, Taylor HS (1997) Harmonic inversion of time signals and its applications. J Chem Phys 107:6756–6769
Mandelshtam VA, Taylor HS, Shaka AJ (1998) Application of the filter diagonalization method to one- and two-dimensional NMR spectra. J Magn Reson 133:304–312
Marion D, Ikura M, Tschudin R, Bax A (1989) Rapid recording of 2D NMR spectra without phase cycling––application to the study of hydrogen-exchange in proteins. J Magn Reson 85:393–399
McCallum SA, Pardi A (2003) Refined solution structure of the iron-responsive element RNA using residual dipolar couplings. J Mol Biol 326:1037–1050
Nikonowicz EP, Pardi A (1992) Application of 4-dimensional heteronuclear NMR to the structure determination of a uniformly C-13 labeled RNA. J Am Chem Soc 114:1082–1083
Orekhov VY, Ibraghimov I, Billeter M (2003) Optimizing resolution in multidimensional NMR by three-way decomposition. J Biomol NMR 27:165–173
Pardi A (1995) Multidimensional heteronuclear NMR experiments for structure determination of isotopically labeled RNA. Methods Enzymol 261:350–380
Schmieder P, Stern AS, Wagner G, Hoch JC (1993) Application of nonlinear sampling schemes to COSY-type spectra. J Biomol NMR 3:569–576
Stephenson DS (1988) Linear prediction and maximum entropy methods in NMR spectroscopy. Prog Nucl Magn Reson Spectrosc 20:515–526
Vranken WF, Boucher W, Stevens TJ, Fogh RH, Pajon A, Llinas M, Ulrich EL, Markley JL, Ionides J, Laue ED (2005) The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins 59:687–696
Wijmenga SS, van Buuren BNM (1998) The use of NMR methods for conformational studies of nucleic acids. Prog Nucl Magn Reson Spectrosc 32:287–387
Zhu G, Bax A (1990) Improved linear prediction for truncated signals of known phase. J Magn Reson 90:405–410
Acknowledgements
This work was supported in part by NIH grants AI33098 (Arthur Pardi), GM68928 (Geoffrey S. Armstrong), NSF MCB-0236103 (Brad Bendiak) and Michael P. Latham was supported in part by an NIH Training Grant T32 GM65103.
Author information
Authors and Affiliations
Corresponding author
Additional information
Justin T. Douglas and Michael P. Latham contributed equally to this work.
Electronic supplementary Materials
Rights and permissions
About this article
Cite this article
Douglas, J.T., Latham, M.P., Armstrong, G.S. et al. High-resolution pyrimidine- and ribose-specific 4D HCCH-COSY spectra of RNA using the filter diagonalization method. J Biomol NMR 41, 209–219 (2008). https://doi.org/10.1007/s10858-008-9253-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-008-9253-3