Abstract
The “Cheminformatics aspects of high throughput screening (HTS): from robots to models” symposium was part of the computers in chemistry technical program at the American Chemical Society National Meeting in Denver, Colorado during the fall of 2011. This symposium brought together researchers from high throughput screening centers and molecular modelers from academia and industry to discuss the integration of currently available high throughput screening data and assays with computational analysis. The topics discussed at this symposium covered the data-infrastructure at various academic, hospital, and National Institutes of Health-funded high throughput screening centers, the cheminformatics and molecular modeling methods used in real world examples to guide screening and hit-finding, and how academic and non-profit organizations can benefit from current high throughput screening cheminformatics resources. Specifically, this article also covers the remarks and discussions in the open panel discussion of the symposium and summarizes the following talks on “Accurate Kinase virtual screening: biochemical, cellular and selectivity”, “Selective, privileged and promiscuous chemical patterns in high-throughput screening” and “Visualizing and exploring relationships among HTS hits using network graphs”.
Similar content being viewed by others
Introduction
This article introduces the symposium while providing a background of why this symposium was of interest to the computational community and the topics covered at the meeting; the authors of the mini-series articles contributed to the symposium.
The drug discovery process in the past 20 years has been greatly accelerated by the utilization of robotic automation for key aspects. Specifically, the ability to screen a chemically diverse compound library consisting of tens of thousands of compounds in a single pass for a disease target and then analyzing the plethora of biological and chemical information to aid in the early stage of drug discovery. In 2005 the National Institutes of Health (NIH) established the $88.9 million dollar Molecular Libraries Screening Centers Network1 (MLSCN) to aid academic researchers. The goal of this initiative was to build a collaborative research network to leverage high-throughput screening methods. Small molecules identified by HTS at MLSCN can then be used as research tools to enable academic researchers the ability to explore new ideas and further projects on a broad front against human diseases. The MLSCN is also part of the NIH Roadmap for medical research and “New Pathways to Discovery” initiative, whose goal is to advance the understanding of biological systems and provide a better “toolbox” for medical researchers. One of the most powerful aspects of this NIH effort is the availability of the generated high-throughput assay data to researchers in both the public and private sectors through the PubChem database at the National Library of Medicine of NIH (http://pubchem.ncbi.nlm.nih.gov/).
The potential to analyze a large quantity of biological endpoints for a set of small molecules that have been tested for activity across different bioassays, opens a new era of understanding with respect to the mechanisms of disease and identification of potential targets for new therapies. The available HTS data generated by the NIH funded screening centers is composed of more than 300,000 compounds from commercial sources and synthesized by academic and government researchers. For the first time in the history, a large amount of molecular data—chemical structures, biological systems, and endpoints—is provided in an unencumbered fashion to the public for study and analysis. The excellent design and user-friendliness of the PubChem Substance and Bioassay databases enables academic and government researcher groups along with small biotech companies a place to contribute their HTS results for a defined set of small molecules against a specific disease (biological target).
Though symposia regarding HTS and cheminformatics paradigms have been organized in the past, there has not been a symposium that bridges these two unique and vital aspects. The “Cheminformatics aspects of high throughput screening (HTS): from robots to models” symposium provided researchers the opportunity to address and discuss general topics, specifically: the data-infrastructure of HTS centers, the methods underlining the management of HTS data to guide iterative screening and hit-finding, together with the potential exchange of data, methods, or computational models between academic, government, and industrial groups.
Topics covered in “Cheminformatics aspects of high throughput screening (HTS): from robots to models”
The “Cheminformatics aspects of high throughput screening (HTS): from robots to models” symposium aimed to foster a discussion between the scientists that work in the areas of high throughput screening, cheminformatics and molecular modeling. A state-of-the-art HTS pipeline relies on each scientist’s talents, and improvements to the entire process require better cheminformatics techniques. This symposium was organized within the Computers in Chemistry (COMP) division’s technical program and co-sponsored by the Chemical Information (CINF) and Medicinal Chemistry (MEDI) divisions of the American Chemistry Society.
The symposium had two half-day sessions beginning with the introductory remarks of Y. Jane Tseng, Ph.D. from National Taiwan University to describe the aims and scope of this symposium. The first speaker, Stephen Bryant, Ph.D. of the NIH, where he is PubChem’s Database Leader, started the discussion by providing an introduction to the “Open repository for chemical structure and biological activity information”. Sivaraman Dandapani, Ph.D. from the Broad Institute of MIT and Harvard demonstrated a superbly designed medicinal screening platform infrastructure for the screening of stereoisomers using the Broad Chemical Biology Informatics Platform for analyzing the data. S. Joshua Swamidass, Ph.D. from Washington University in St. Louis presented an interesting and intriguing data mining study through the use of multiple HTS bioassays on probabilistic substructure mining from small-molecule screens.
The second half of the symposium included real-world examples from the Novartis Institutes for Biomedical Research’s Global Discovery Chemistry division presented by Eric Martin, Ph.D. Dr. Martin presented a great example from the pharmaceutical industry that demonstrated the application of iterative and accurate kinase virtual screens—constructed from partial training data—to handle the biochemical, cellular, and selectivity problems without the aid of solved crystal structures of the kinase receptor of interest. Cristian G. Bologa, Ph.D. from the University of New Mexico Center for Molecular Discovery presented a very commonly seen but hard to deal with obstacle that is experienced at all HTS centers: promiscuous chemical patterns in HTS. Anang Shelat, Ph.D. from the Chemical Biology and Therapeutics Department at St. Jude Children’s Research Hospital, discussed ways to visualize and explore the varied relationships among HTS hits using network graphs. Visualization techniques are important when dealing with large quantities of data for better interpretation and understanding.
The symposium was concluded with an open panel discussion. Detailed articles on several of the talks presented in the symposium are presented in— “An Informatic Pipeline for Managing High-Throughput Screening Experiments and Analyzing Data from Stereochemically Diverse Libraries” by C. Mulrooney et al. and “Managing Missing Measurements in Small-Molecule Screens” by M. R. Browning and coworkers. The discussions of the open panel discussion in the symposium as well as the summary of the following talks on the topics of “Accurate Kinase virtual screening: biochemical, cellular and selectivity” by Dr. Eric Martin, “Selective, privileged and promiscuous chemical patterns in high-throughput screening” by Dr. Cristian Bologa, and “Visualizing and exploring relationships among HTS hits using network graphs” by Dr. Anang Shelat are included in this article. Detailed articles on several of the talks presented in the symposium are presented elsewhere in this issue.
Summary of “accurate kinase virtual screening: biochemical, cellular and selectivity”
“Accurate Kinase Virtual Screening: Biochemical, Cellular and Selectivity” was presented by Dr. Eric Martin. The talk presented a collection of uniquely accurate protein family–based kinase virtual screening methods for predicting biochemical and cellular activity and selectivity for new kinase targets without requiring the protein structure, and with or without training data.
Experimental high throughput screening (HTS) has provided most drug leads for over 20 years. However, a typical HTS of a large corporate archive takes 6-9 months and can cost nearly $1,000,000. Thus, fast, inexpensive “virtual-screening” is a long-standing goal. Unfortunately, conventional docking and QSAR lack the predictive power to serve as an adequate alternative. This presentation described 4 novel Protein-Family Virtual Screening (PFVS) methodologies specifically for kinases: Profile-QSAR, Kinase-Kernel, AutoShim and Surrogate AutoShim. These computational methods combine a modest amount of IC50 data for a new kinase with a vast kinase knowledgebase from all previous Novartis kinase projects, giving unparalleled IC50 predictions rivaling the accuracy of experimental HTS.
Profile-QSAR [1], is a 2D substructure-based meta-QSAR. It requires about 500 experimental IC50s for a new kinase, used to train a new model using predicted activities from 100 historical kinase Quantitative Structure–Activity Relationships (QSARs) as compound descriptors. Every prediction is informed by over 1.5 million historical IC50s from over 130,000 compounds, resulting in unprecedented accuracy and extrapolation power. Profile-QSAR also predicts cellular activity, selectivity among kinases, and entire kinase profiles for 115 kinases with sufficient training data.
However, there are nearly 400 additional kinases of potential pharmaceutical interest. Kinase-Kernel [2] models interpolate between each new kinase’s nearest neighbors from among the 115 with trained Profile-QSAR models, based on active-site sequence similarity. Thus, activity for the remaining kinases with no training data can also be predicted.
Profile-QSAR and Kinase Kernel models, based only on the 2D substructures of the ligands, have unmatched accuracy. However an orthogonal docking method, which instead uses 3D interactions with the protein active site, could find additional active compounds. Unfortunately, docking suffers from 3 key limitations: it is slow, requires a protein structure for the target, and cannot predict reliable IC50s. Our AutoShim [3] method trains highly accurate customized scoring functions for each new protein target by adjusting the weights of pharmacophore interaction—“shims”, added to the binding site, to reproduce 500 training IC50s. AutoShim has excellent accuracy, and is not restricted to kinases, but is still slow and requires a protein structure.
However, for kinases, Surrogate AutoShim [4] uses a “Universal Kinase Surrogate Receptor” ensemble of 16 diverse kinase crystal structures as a proxy for the actual kinase of interest. Four million internal and commercial compounds have been pre-docked into this surrogate receptor, and the docking scores and pharmacophore interactions have been extracted and stored for the billions of docked poses. Surrogate AutoShim models for new kinases can be trained, and accurate IC50′s of these 4 million compounds predicted, in just hours rather than weeks, without further docking. Combined, Profile QSAR, Kinase-Kernel and Surrogate AutoShim identify most of the actives in a compound collection. Activity has been predicted for these 4 million internal and commercial compounds across all 500 + human kinases and numerous cell lines, so an initial PFVS for any kinase is now just a table lookup. To do this experimentally would take many years and cost at least a billion dollars!
PFVS has been successfully applied to about 50 recent Novartis kinase projects, with external R2 = 0.35–0.7, enrichments of 20x–50x, and hit rates of 25–80 % even for completely novel compounds. Applications have covered a wide range of project stages: finding tool compounds for target validation ahead of HTS, virtual screening where protein is too scarce for full HTS, “catch-up” screens for compounds added to the archive since an earlier HTS, rescuing HTS false-negatives, triaging experimental HTS hits for cellular potency or specific selectivities, selecting compounds from commercial vendors, and enhancing the archive based on kinase profile diversity.
Since the symposium, PFVS has been extended to additional families: GPCRs, Ser/Cis proteases, and many non-kinase adenosine nucleotide binding proteins from many families including chaperones, carboxylases, phosphodiesterases, ABC transporters, pyrophosphatases, ion channels, ubiquitin-congugators and others. No longer a kinase-specific niche technology, PFVS now applies to roughly ½ of the estimated druggable genome (Fig. 1).
Summary of “Selective, privileged and promiscuous chemical patterns in high-throughput screening”
The talk “Selective, privileged and promiscuous chemical patterns in high-throughput screening” was presented by Dr. Cristian Bologa. The talk was focused on the lessons learned in the process of finding successful chemical probes at the University of New Mexico Center for Molecular Discovery.
The University of New Mexico Center for Molecular Discovery (UNMCMD) is a screening center within the Molecular Libraries Probe Production Centers (MLPCN) focusing on multiplexed, high throughput flow cytometry. The UNMCMD was part of the NIH Roadmap for The Molecular Libraries Program (MLP) since 2005. UNMCMD has been screening more than 10 million chemical samples in more than 300 assays and has identified 14 chemical probes by August 2011. The UNM Biocomputing facility is focused on bioinformatics, cheminformatics, ligand- and structure-based molecular design and supports high-throughput screening (HTS) data mining and screening informatics for UNMCMD. All the screening results of UNMCMD are made public in the PubChem database. Of all the chemical probes developed by the MLP program, the UNMCMD GPR30 agonist G-1 [5] has got the highest number of citations at the time of this presentation, followed by two chemical probes from the Scripps Research Institute, an agonist and an antagonist of the S1P1 receptor.
In the author’s opinion, five factors have a great influence on finding a successful chemical probe—“hot” target, high potency, proper physico-chemical properties, demonstrated in vivo activity, and selectivity on the intended target. The last mentioned factor, selectivity, is not the most important property for drugs, which can produce their desired pharmacological effect by acting on multiple targets, but it is one of the most important ones for a chemical probe, used to investigate the role of the intended target in biological pathways or medical diseases. In a general library screening, the goal of HTS is to produce candidate molecular scaffolds for lead/probe optimization. However, making sense of large scale (and noisy) HTS results is not always easy. Understanding the potential of each scaffold very early in the project is critical for success and for avoiding later troubles. There are roughly four categories of scaffolds that can be defined based on the screening results in many HTS assays: not active (yet), selective, privileged, and promiscuous. Not active scaffolds are those found in compounds not active on any tested targets. Selective scaffold are those active on only one or very few specific targets. Privileged scaffolds are defined as those active on multiple targets from the same family (such as GPCR, kinases, ion channels, etc.). The last and also the focus of this presentation are the promiscuous scaffolds—those found to be “active” in assays on several targets from multiple families.
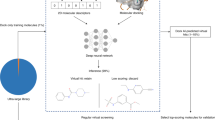
The problem of false positives in HTS has been studied in the past [6–14], and diverse sets of structural filters have been proposed to remove these promiscuous compounds from screening libraries, or from the list of the active hits that will further advance in the project. One question to be asked is whether we use the right filters to select the right chemical scaffolds for probe optimization. While that might be true, the filters would probably not cover newly synthesized promiscuous scaffolds that were not available when those structural filters were designed. For this reason, an unbiased retrospective analysis of the primary screening assay results of our current chemical library in a large number of assays can show us what are the problematic scaffolds in this library. In order to perform this analysis, we extracted from the Pubchem database only those assays that had more than 20,000 MLSMR substances tested (575 assays total), with the physicochemical profiling assays excluded from this list. The total number of substances tested in any of the 575 assays was 374,062, and 261,462 of those being found active in at least one assay. There were a total number of 126,528,538 samples tested and 931,567 of them were found active, giving an average hit rate for “samples” of around 0.7 %. If we compute the same sample hit rate for the compound sets selected using literature published substructure filters for removing false positives, we would expect that value to be significantly higher. However, we found actual hit rates between 0.7 and 17 %, with a weighted average of 1.3 %. That simply says that not all, but many of the structural filters published in the literature remove very little of the potential false positives, and at the same time remove a lot of potentially good compounds.
In order to prioritize chemical scaffolds for probe optimization, we have focused our analysis on identification of promiscuous scaffolds. Scaffold generation has been researched in the past, and we have re-implemented a recently published method [15] (re-implemented source code available at http://code.google.com/p/unm-biocomp-hscaf). For each of the unique scaffold extracted from all the MLSMR substances, we then computed a “promiscuity score” based on the following numbers: number of substances containing that scaffold tested (in any of the 575 selected HTS assays), number of substances found active, number of assays where compounds containing that scaffold have been tested, number of assays where some of those compounds where found active, number of samples containing that scaffold tested, and number of samples active. Examples of most promiscuous scaffolds detected using this analysis are shown in Fig. 2. UNM Biocomputing has also implemented a BioActivity Data Associative Promiscuity Pattern Learning Engine (“badapple”) at http://pasilla.health.unm.edu/tomcat/biocomp/badapple to evaluate HTS hits and warn if they contain any promiscuous scaffolds.
An alternative strategy to the use of scaffolds in promiscuity analysis is to use matched molecular pairs (MMPs) [16] to extract information regarding structural changes that induce promiscuity. Once identified, that information could be used, for example, for designing of more selective compounds by replacing the promiscuous part(s). The following steps have been used for finding the most promiscuous substructures: (1) downloading and cleaning of the MLSMR library from PubChem by removing salt and solvent molecules, and also normalizing tautomers; (2) identifying all non-redundant single, double, and triple cut MMPs; (3) mapping “active” MMPs to HTS assays; and (4) computing a promiscuity score similar to the one used for ranking scaffolds. Some of the top promiscuous fragments are presented in Fig. 3.
Finally, there are many questions that can be answered by performing a similar analysis, but have not been asked yet—for example, which scaffolds are found active more often in: one screening center compared to the other centers; in yeast versus mammalian cells assays; in bioluminescent assays versus other assays; in a specific class of targets versus all the other—the classic case of privileged patterns, (see some examples in Fig. 4); in assays with or without a certain reagent (DTT); or in phenotypic versus target-based screens? Unfortunately, some of these questions cannot be answered easily at this moment due to the curation effort needed by the fact that most of this type of information is available in Pubchem only in free text format.
To summarize some of the lessons learned from UNMCMD: it is easy to find non-selective actives but hard to find the selective ones; it is very important to have the right controls in every assay instead of purely rely on the computational filters; physico-chemical profiling assays are helpful only if run under the same assay conditions as the main assay; and it is also important to have robust counter screens and secondary assays in place.
Summary of “visualizing and exploring relationships among HTS hits using network graphs”
The talk “Visualizing and exploring relationships among HTS hits using network graphs” was presented by Dr. Anang Shelat. The talk describes how network graph techniques were used to analyze hits from a high-throughput screen (HTS) for antimalarial agents [19] and to guide subsequent SAR studies that resulted in the identification of several novel scaffold series.
The motivation for using network graphs to explore HTS results was inspired by the work of Schuffenhauer et al. [18, 23] and Wetzel et al. [17].
An HTS primary screen ‘hit’ is less likely to be a false positive if it belongs to a well-defined SAR series: a scaffold that demonstrates significant changes to potency (ideally > 2 log) with small structural variation. To identify such scaffolds, we routinely employ an SAR-by-catalog approach to construct analog sets from our internal library and commercially-available databases. At this stage in the discovery process, we place a premium on interpretability and opt for building models that are based on ‘obvious’ topological relationships between molecules. Network graphs are a powerful tool for visualizing and understanding biological activity in the context of these topological relationships.
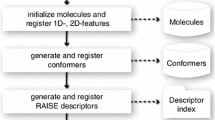
To build the network graph, input molecules are subject to several rounds of abstraction to yield representations with decreasing complexity. The input molecules and scaffolds define the nodes of the graph, and nodes that are topologically related are connected by edges. Figure 5 describes the abstraction process in detail. First, the Bemis-Murcko fragmentation algorithm [20] is used to truncate side chains while keeping intact contiguous ring systems plus any chains that link two or more rings. In this ‘scaffold’ representation, the alpha atom at side chain substitution points is preserved as a generic carbon atom. At the next abstraction level, a ‘topology’ is generated by removing the alpha atom to allow substitutions at all positions, and by saturating all bonds. We have found that bond saturation is an effective way to ensure that tautomers, especially those from dispersed pi systems that sometimes confound canonical tautomerization algorithms, are assigned to the same parent node. The final level of abstraction involves hierarchically clustering topologies using the McQuitty algorithm with the ECFP_4 fingerprint and the Dixon-Koehler dissimilarity metric [22]. This abstraction procedure is performed by a program written in the Pipeline Pilot environment (Accelrys), using native functions except for clustering which requires an R script that uses the hclust function [21].
The input molecules and entities resulting from the abstraction process represent the nodes in the network graph. The nodes are connected by edges according to the following rules: (a) molecules are connected to other molecules if they share the same scaffold; (b) molecules are connected to their parent scaffold; (c) scaffolds are connected to a ‘parent’ scaffold if a sub-structure relationship exists; (d) scaffolds not connected to a ‘parent’ scaffold are connected to their ‘parent’ topology; (e) topologies are connected to each other according to the dendrogram generated from hierarchical clustering. This algorithm generates a planar tree graph: every pair of nodes has exactly one edge and no edges cross. Nodes annotations and relationships are stored in a relational database as parent–child pairs, facilitating efficient lookup during network construction.
The Cytoscape program [25] is used to visualize and explore the network graph. In addition to having a number of useful network layout algorithms and interactive tools, this application makes it easy to annotate graph attributes and then visualize these annotations within the context of the graph structure. For example, Fig. 6 shows a single branch from the network graph created for the antimalarial HTS (refer to [19] for the full network graph). Nodes representing HTS hits are colored according to potency against the K1 strain of Plasmodium falciparum, and sized according to potency against the 3D7 strain. Edge color indicates topological relationships. The layering of multidimensional biological data and chemical topological information facilitates a deeper understanding of the SARs present within the 2,3-diaminonaphthalene-1,4-dione cluster.
Visual interrogation of the antimalarial HTS network graph helped identify well-defined SAR series, such as the 2,3-diaminonaphthalene-1,4-dione scaffold series, but also pinpointed hits from poorly sampled regions of chemical space. Such singletons were further explored using the SAR-by-catalog workflow described in Fig. 7. We expanded the network graph to include node relationships from molecules beyond the original HTS screening library, including our entire in-house chemical library (N = 543,022) and the Screening Compounds Directory ('SCD', N = 9,533,508, Accelrys). We retrieved analogs by walking up the branches of validated hits to identify similar scaffolds and topologies, and then walking down the branches of those nodes to identify analogs of our hits. Longer walks away from the hit node result in a larger number of retrieved compounds, but also result in less similar analogs.
In the malaria project, we queried our network graph database using 228 high quality hits that were derived from 156 scaffolds. We identified 1,213 novel analogs in our in-house library and 4,017 novel analogs from the SCD after walking up to the scaffold level and then down new branches. After acquiring and screening 1,056 novel compounds, we were able to establish well-defined SARs for 13 scaffolds (data in preparation for publication).
Our network graph approach enabled facile interpretation of HTS results and rapid construction of SAR series for subsequent study. However, during implementation, we encountered the following limitations: (a) the Bemis-Murcko fragmentation algorithm requires at least one ring and does not process linear molecules, and is unable to deal with situations where an aliphatic side chain contains important pharmacophore elements; (b) the abstraction process can miss obvious relationships between multi-ring compounds differing by a single ring or compounds with rings differing by a single heteroatom; (c) topologies are not real molecules and can sometimes be difficult to interpret. We are currently working to address (b) by combining Bemis-Murcko fragmentation with the Schuffenhauer decomposition [8, 23], and (c) by implementing an improved canonical tautomerization algorithm that avoids having to force bond saturation.
Furthermore, we are expanding the use of network graphs to describe relationships beyond chemical topology. For example, the Anatomical Therapeutic Chemical (ATC) classification system [24] classifies compounds at five levels based on physiological or therapeutic effect and chemical structure. As of 2011, Level 1 includes 14 anatomical groups; level 2 includes the therapeutic main groups; level 3 contains the therapeutic-pharmacological subgroups; level 4 contains the chemical-therapeutic-pharmacological subgroups; and level 5 contains actual chemical substances. A network graph can be constructed using this hierarchy, and visualized in a manner similar to that used in the antimalarial HTS study. Figure 8 shows an example of a branch from a network graph constructed from an HTS screen against wild-type and mutant tumor cells (unpublished data). The active molecules in this subset include compounds with scaffolds quite dissimilar to the three artermisins depicted, yet share common biological activity as chemotherapeutics targeting eukaryotic protozoa.
In summary, network graphs are powerful tools for exploring multidimensional relationships among biological and chemical properties, and they enable rapid and efficient evolution of HTS hits.
Final open panel discussion
Two main topics were raised in the open panel discussion: availability of data from non-NIH funded HTS centers, and how to deal with the noise that can interfere with the HTS data management.
Current non-NIH funded HTS centers, especially from non-profit organizations and the private sector, have no obligation to deposit data or make data available to the public. Steve Bryant commented on the issue and explained that there is no restriction from the NIH Pubchem database that prevents anyone from depositing assays having even just a few compounds. The NIH Pubchem database does not own the copyright of the data, but rather serves just as a public repository. The private sector does have intellectual property issues that prevent them from depositing data to public repositories such as the PubChem database. An audience member from an NIH funded HTS center asked whether it is possible to deposit the molecular models or virtual screening models developed using the HTS data. That is, instead of directly depositing the data (compounds), the private sector deposits the models, which will not reveal the compound structures belonging to the companies. Steve Bryant commented on this interesting proposal and stated that there were no rules preventing depositors from depositing the models in the PubChem database, although there was no such precedent. An audience member noted that there is a public database of biological models called BioModel.net [26], but that a similar scheme for molecular models might be more difficult to implement due to reliance on proprietary software applications. Eric Martin commented that the Profile-QSAR and AutoShim IC50 virtual screening models are trained using high-quality IC50 assay data. They typically require 400 + IC50 s from a single assay, or from combined assays shown to correlate well, and with at least 25 active compounds. They have not used public data to train their models yet, but if the data met those criteria it could be used. Anang Shelat lamented the lack of quality models in the public domain, but noted that pharmaceutical companies are increasingly granting access to their chemical tools and subsets of their screening libraries, and that perhaps enabling broader use of some of their computational models would follow.
The second discussion in the panel was on handling noise in the HTS data. HTS data, especially from primary screening, contains a significant amount of noise from the experimental design that affects further cheminformatics results. Cristian Bologa of UNM commented that these HTS artifacts might have multiple causes: mechanical—compound carryover, solvent evaporation, edge effects, etc.; optical—compound fluorescence or fluorescence quenching, light scattering by insoluble compounds; membrane permeabilization or cytotoxicity, in cellular assays; reactive or redox cycling compounds; and other assay specific artifacts. For example some of the most promiscuous compounds in MLSMR are known redox cycling compounds, and the active compounds in the assay “NIH Library Profiling”: Compound and DTT Dependent Redox Cycling H2O2 Generation (Pubchem AID: 828) have an overall hit rate of 8.2 % in all other MLSMR assays, much higher than 0.7 % for the compounds inactive in that assay. Not the same can be said about other physico-chemical profiling assays designed to catch false positives. For example, there is very little difference between the hit rate of actives (1 %; 1.5 %) and inactives (0.7 %) in the “aggregator” profiling assays (AIDs: 585 and 1,476). These HTS artifact problems encountered in the HTS screening campaigns are best handled from the beginning of the assay development and HTS experimental design: developing of robust assays which include verified positive and negative controls, screening in duplicate, triplicate or at multiple concentrations, having additional counterscreens and secondary screens that use a different detection method, etc. After the screening has been performed, a cheminformatics post-HTS analysis can and should be used to filter out known false positives, try to recover false negatives, and prioritize the most promising set of scaffolds for further testing and development. Eric Martin commented that they find that the most important factor in the quality of their Profile-QSAR and AutoShim IC50 virtual screening models is the dynamic range of the training data. Single concentration HTS percent inhibition data are not only noisy, they also have very little dynamic range, so they have not trained models on HTS data. They would rather build a model on 500 IC50 s (assuming at least 25 active compounds) than 2 million HTS percent inhibitions. Conversely, some of the key uses of their models are to help triage noisy HTS data, recovering of false negatives, predicting which HTS hits are highly potent or highly ligand efficient, and predicting selectivity over anti-targets for which they have models. In addition, medicinal chemists can start working immediately on early chemical leads from a virtual screen, allowing the biologists time to develop more sensitive, accurate and relevant HTS assays, rather than rushing the HTS to get chemistry something to work on.
References
Martin E, Mukherjee P, Sullivan D, Jansen J (2011) J Chem Inf Model 51(8):1942
Martin E, Mukherjee P (2011) J Chem Inf Model 52(1):156
Martin EJ, Sullivan DC (2008) J Chem Inf Model 48(4):861
Axerio-Cilies P, Castañeda I, Mirza A, Reynisson J (2009) Eur J Med Chem 44(3):1128
Bologa CG, Revankar CM, Young SM, Edwards BS, Arterburn JB, Kiselyov AS, Parker MA, Tkachenko SE, Savchuck NP, Sklar LA, Oprea TI, Prossnitz ER (2006) Nat Chem Biol 2(4):207
Huth JR, Song D, Mendoza RR, Black-Schaefer CL, Mack JC, Dorwin SA, Ladror US, Severin JM, Walter KA, Bartley DM, Hajduk PJ (2007) Chem Res Toxicol 20(12):1752
Di L, Kerns EH (2009) Chem Biodivers 6(11):1875
McGovern SL, Helfand BT, Feng B, Shoichet BK (2003) J Med Chem 46(20):4265
Rishton GM (1997) Drug Discovery Today 2(9):382
Jadhav A, Ferreira RS, Klumpp C, Mott BT, Austin CP, Inglese J, Thomas CJ, Maloney DJ, Shoichet BK, Simeonov A (2009) J Med Chem 53(1):37
Baell JB, Holloway GA (2010) J Med Chem 53(7):2719
Metz J, Huth J, Hajduk P (2007) J Comput-Aid Mol Des 21(1–3):139
Thorne N, Auld DS, Inglese J (2010) Curr Opin Chem Biol 14(3):315
Huth JR, Mendoza R, Olejniczak ET, Johnson RW, Cothron DA, Liu Y, Lerner CG, Chen J, Hajduk PJ (2004) J Am Chem Soc 127(1):217
Wilkens S, Janes J, Su A (2005) J Med Chem 48(9):3182
Leach AG, Jones HD, Cosgrove DA, Kenny PW, Ruston L, MacFaul P, Wood JM, Colclough N, Law B (2006) J Med Chem 49(23):6672
Wetzel S, Klein K, Renner S, Rauh D, Oprea T, Mutzel P, Waldmann H (2009) Nat Chem Biol 5(8):581
Schuffenhauer A, Ertl P, Roggo S, Wetzel S, Koch M, Waldmann H (2007) J Chem Inf Model 47(1):47
Guiguemde W, Shelat A, Bouck D, Duffy S, Crowther G, Davis P, Smithson D, Connelly M, Clark J, Zhu F, Jiménez-Díaz M, Martinez M, Wilson E, Tripathi A, Gut J, Sharlow E, Bathurst I, El Mazouni F, Fowble J, Forquer I, McGinley P, Castro S, Angulo-Barturen I, Ferrer S, Rosenthal P, Derisi J, Sullivan D, Lazo J, Roos D, Riscoe M, Phillips M, Rathod P, Van Voorhis W, Avery V, Guy R (2010) Nature 465(7296):311
Bemis GW, Murcko MA (1996) J Med Chem 39(15):2887
R: A Language and Environment for Statistical Computing (2013) R Core Team, R Foundation for Statistical Computing, Vienna, Austria. http://www.R-project.org
Dixon SL, Koehler RT (1999) J Med Chem 42(15):2887
Schuffenhauer A, Brown N, Ertl P, Jenkins JL, Selzer P, Hamon J (2007) J Chem Inf Model 47(2):325
Structure and principles Accessed, http://www.whocc.no/atc/structure_and_principles/
Smoot M, Ono K, Ruscheinski J, Wang P-L, Ideker T (2011) Cytoscape 2.8: new features for data integration and network visualization. Bioinformatics (Oxford, England) 27(3):431–432. doi:10.1093/bioinformatics/btq675
Acknowledgments
Y. Jane Tseng would like to acknowledge the travel funding from the Taiwan National Science Council, grant numbers 100-2627-B-002-016, and 100-2325-B-009-001-. Eric Martin would like to acknowledge David Sullivan, Prasenjit Mukherjee, and Li Tian for their work developing protein-family virtual screening and applying it to dozens of projects. Cristian Bologa would like to acknowledge Jeremy J Yang for the implementation of the scaffold extraction algorithm, and the development of the BADAPPLE system; and Oleg Ursu for the implementation, analysis of results, and preparing the slides for the molecular matching pairs part of the presentation. Work on the identification of promiscuous patterns has been supported by the NIH U54MH084690 and NCRR P20 RR016480 grants. Anang Shelat would like to acknowledge Armand Guiguemde and David Smithson for their contributions to the network graph algorithm, and Cindy Nelson and Heather Ross for their work as part of the Compound Management group at SJCRH.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Jane Tseng, Y., Martin, E., G. Bologa, C. et al. Cheminformatics aspects of high throughput screening: from robots to models: symposium summary. J Comput Aided Mol Des 27, 443–453 (2013). https://doi.org/10.1007/s10822-013-9646-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-013-9646-6