Abstract
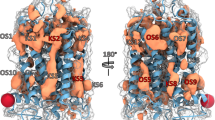
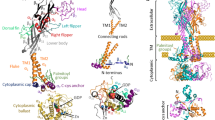
Ongoing efforts to model P2Y receptors for extracellular nucleotides, i.e., endogenous ADP, ATP, UDP, UTP, and UDP-glucose, were summarized and correlated for the eight known subtypes. The rhodopsin-based homology modeling of the P2Y receptors is supported by a growing body of site-directed mutagenesis data, mainly for P2Y1 receptors. By comparing molecular models of the P2Y receptors, it was concluded that nucleotide binding could occur in the upper part of the helical bundle, with the ribose moiety accommodated between transmembrane domain (TM) 3 and TM7. The nucleobase was oriented towards TM1, TM2, and TM7, in the direction of the extracellular side of the receptor. The phosphate chain was oriented towards TM6, in the direction of the extracellular loops (ELs), and was coordinated by three critical cationic residues. In particular, in the P2Y1, P2Y2, P2Y4, and P2Y6 receptors the nucleotide ligands had very similar positions. ADP in the P2Y12 receptor was located deeper inside the receptor in comparison to other subtypes, and the uridine moiety of UDP-glucose in the P2Y14 receptor was located even deeper and shifted toward TM7. In general, these findings are in agreement with the proposed binding site of small molecules to other class A GPCRs.
Similar content being viewed by others
References
Abbracchio, MP, Burnstock, G, Boeynaems, JM, Barnard, EA, Boyer, JL, Kennedy, C, Fumagalli, M, King, BF, Gachet, C, Jacobson, KA, Weisman, GA. International Union of Pharmacology. Update of the P2Y G protein-coupled nucleotide receptors: from molecular mechanisms and pathophysiology to therapy. Pharmacol Rev (in press)
Costanzi S, Mamedova L, Gao ZG, Jacobson KA (2004) Architecture of P2Y nucleotide receptors: Structural comparison based on sequence analysis, mutagenesis, and homology modeling. J Med Chem 47:5393
Visiers I, Ballesteros JA, Weinstein H (2002) Three-dimensional representations of G protein-coupled receptor structures and mechanisms. Methods Enzymol 343:329
Van Rhee AM, Fischer B, van Galen PJM, Jacobson KA (1995) Modelling the P2Y purinoceptor using rhodopsin as template. Drug Des Discov 13:133
Moro S, Guo D, Camaioni E, Boyer JL, Harden TK, Jacobson KA (1998) Human P2Y1 receptor: Molecular modeling and site-directed mutagenesis as tools to identify agonist and antagonist recognition sites. J Med Chem 41:1456
Moro S, Hoffmann C, Jacobson KA (1999) Role of the extracellular loops of G protein-coupled receptors in ligand recognition: A molecular modeling study of the human P2Y1 receptor. Biochemistry 38:3498
Jiang Q, Guo D, Lee BX, van Rhee AM, Kim YC, Nicholas RA, Schachter J, Harden TK, Jacobson KA (1997) A mutational analysis of residues essential for ligand recognition at the human P2Y1 receptor. Mol Pharmacol 52:499
Kim HS, Barak D, Harden TK, Boyer JL, Jacobson KA (2001) Acyclic and cyclopropyl analogues of adenosine bisphosphate antagonists of the P2Y1 receptor: structure-activity relationships and receptor docking. J Med Chem 44:3092
Major DT, Nahum V, Wang Y, Reiser G, Fischer B (2004) Molecular recognition in purinergic receptors. 2. Diastereoselectivity of the h-P2Y1-receptor. J Med Chem 47:4405
Major DT, Nahum V, Wang Y, Reiser G, Fischer B (2004) Molecular recognition in purinergic receptors. 1. A comprehensive computational study of the h-P2Y1-receptor. J Med Chem 47:4391
Guo D, von Kügelgen I, Moro S, Kim YC, Jacobson KA (2002) Evidence for the recognition of non-nucleotide antagonists within the transmembrane domains of the human P2Y1 receptor. Drug Devel Res 57:173
Nandanan E, Jang SY, Moro S, Kim H, Siddiqui MA, Russ P, Marquez VE, Busson R, Herdewijn P, Harden TK, Boyer JL, Jacobson KA (2000) Synthesis, biological activity, and molecular modeling of ribose-modified adenosine bisphosphate analogues as P2Y1 receptor ligands. J Med Chem 43:829
Ohno M, Costanzi S, Kim HS, Kempeneers V, Vastmans K, Herdewijn P, Maddileti S, Gao ZG, Harden TK, Jacobson KA (2004) Nucleotide analogues containing 2-oxa-bicyclo[2.2.1]heptane and L-α-threofuranosyl ring systems: Interactions with P2Y receptors. Biooorg Med Chem 12:5619
Hoffmann C, Moro S, Nicholas RA, Harden TK, Jacobson KA (1999) The role of amino acids in extracellular loops of the human P2Y1 receptor in surface expression and activation processes. J Biol Chem 274:14639
Moro S, Jacobson KA (2002) Molecular modeling as a tool to investigate molecular recognition in P2Y receptors. Curr Pharmaceut Design 8:99
Jacobson KA, Costanzi S, Ivanov AA, Tchilibon S, Besada P, Gao Z-G, Maddileti S, Harden TK (2005) Structure activity and molecular modeling analyses of ribose- and base-modified uridine 5′-triphosphate analogues at the human P2Y2 and P2Y4 receptors. Biochem Pharmacol 71:540
Costanzi S, Joshi BV, Maddileti S, Mamedova L, Gonzalez-Moa MJ, Marquez VE, Harden TK, Jacobson KA (2005) Human P2Y6 receptor: molecular modeling leads to the rational design of a novel agonist based on a unique conformational preference. J Med Chem 48:8108
Besada P, Shin DH, Costanzi S, Ko HJ, Mathé C, Gagneron J, Gossselin G, Maddileti S, Harden TK, Jacobson KA (2006) Structure activity relationship of uridine 5′-diphosphate analogues at the human P2Y6 receptor. J Med Chem (in press)
Ivanov AA, Jacobson KA. Molecular dynamics simulation of the human P2Y14 receptor and study of ligand–receptor interactions, 231st Am. Chem. Soc. National Meeting, Atlanta, GA, March 26–30 (2006) Abstract COMP 217
Šali A, Blundell TL (1993) Comparative protein modelling by satisfaction of spatial restraints. J Mol Biol 234:779
Šali A, Overington JP (1994) Derivation of rules for comparative protein modeling from a database of protein structure alignments. Protein Sci 3:1582
Mohamadi FN, Richards GJ, Guida WC, Liskamp R, Lipton M, Caufield C, Chang G, Hendrickson T, Still WC MacroModel––an integrated software
El-Tayeb A, Qi A, Müller CE (2006) Synthesis and structure-activity relationships of base-modified UDP and UTP analogues at the human P2Y2, P2Y4, and P2Y6 receptors. Purinergic Signalling 2:304
Hoffmann K, Algaier I, von Kügelgen I (2006) Evidence for the involvement of basic amino acid residues in transmembrane regions 6 and 7 of the human platelet P2Y12-receptor in ligand recognition. Purinergic Signalling 2:199
Acknowledgments
We acknowledge support from the Intramural Research Program of the NIH, National Institute of Diabetes and Digestive and Kidney Diseases. We thank Prof. Christa Müller and Prof. Ivan von Kügelgen (Univ. of Bonn, Germany) and Prof. T. K. Harden (Univ. of North Carolina) for helpful discussion.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ivanov, A.A., Costanzi, S. & Jacobson, K.A. Defining the nucleotide binding sites of P2Y receptors using rhodopsin-based homology modeling. J Comput Aided Mol Des 20, 417–426 (2006). https://doi.org/10.1007/s10822-006-9054-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-006-9054-2