Abstract
Purpose
Alternations to the paternal epigenome, specifically the components of sperm chromatin, can lead to infertility in humans and potentially transmit aberrant information to the embryo. One key component of sperm chromatin is the post-translational modification of histones (PTMs). We previously identified a comprehensive profile of histone PTMs in normozoospermic sperm; however, only specific histone PTMs have been identified in abnormal sperm by antibody-based approaches and comprehensive changes to histone PTM profiles remain unknown. Here, we investigate if sperm with abnormalities of total motility, progressive motility, and morphology have altered histone PTM profiles compared to normozoospermic sperm samples.
Methods
Discarded semen samples from 31 men with normal or abnormal semen parameters were analyzed for relative abundance of PTMs on histone H3 and H4 by “bottom-up” nano-liquid chromatography-tandem mass spectrometry.
Results
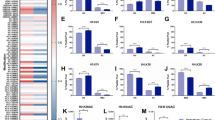
Asthenoteratozoospermic samples (abnormal motility, forward progression, and morphology, n = 6) displayed overall decreased H4 acetylation (p = 0.001) as well as alterations in H4K20 (p = 0.003) and H3K9 methylation (p < 0.04) when compared to normozoospermic samples (n = 8). Asthenozoospermic samples (abnormal motility and progression, n = 5) also demonstrated decreased H4 acetylation (p = 0.04) and altered H4K20 (p = 0.005) and H3K9 methylation (p < 0.04). Samples with isolated abnormal progression (n = 6) primarily demonstrated decreased acetylation on H4 (p < 0.02), and teratozoospermic samples (n = 6) appeared similar to normozoospermic samples (n = 8).
Conclusion
Sperm samples with combined and isolated abnormalities of total motility, progressive motility, and morphology display distinct and altered histone PTM signatures compared to normozoospermic sperm. This provides evidence that alterations in histone PTMs may be important for normal sperm function and fertility.



Similar content being viewed by others
References
Stephen EH, Chandra A. Declining estimates of infertility in the United States: 1982–2002. Fertil Steril. 2006;86:516–23.
Thonneau P, Marchand S, Tallec A, Ferial ML, Ducot B, Lansac J, et al. Incidence and main causes of infertility in a resident population (1,850,000) of three French regions (1988-1989). Hum Reprod. 1991;6:811–6.
Cooper TG, Noonan E, von Eckardstein S, Auger J, Baker HWG, Behre HM, et al. World Health Organization reference values for human semen characteristics. Hum Reprod Update Oxford University Press. 2010;16:231–45.
Larsen L. Computer-assisted semen analysis parameters as predictors for fertility of men from the general population. Hum Reprod. 2000;15:1562–7.
Nallella KP, Sharma RK, Aziz N, Agarwal A. Significance of sperm characteristics in the evaluation of male infertility. Fertil Steril. 2006;85:629–34.
Gunalp S, Onculoglu C, Gurgan T, Kruger TF, Lombard CJ. A study of semen parameters with emphasis on sperm morphology in a fertile population: an attempt to develop clinical thresholds. Hum Reprod. 2001;16:110–4.
Vawda AI, Gunby J, Younglai EV. Andrology: semen parameters as predictors of in-vitro fertilization: the importance of strict criteria sperm morphology. Hum Reprod. 1996;11:1445–50.
Verheyen G, Tournaye H, Staessen C, De Vos A, Vandervorst M, Van Steirteghem A. Controlled comparison of conventional in-vitro fertilization and intracytoplasmic sperm injection in patients with asthenozoospermia. Hum Reprod. 1999;14:2313–9.
Kornberg RD. Structure of chromatin. Annu Rev Biochem. 1977;46:931–54.
Zhao Y, Garcia BA. Comprehensive catalog of currently documented histone modifications. Cold Spring Harb Perspect Biol. 2015;7:a025064–22.
Kouzarides T. Chromatin Modifications and Their Function. Cell. 2007;128:693–705.
Tanphaichitr N, Sobhon P, Taluppeth N, Chalermisarachai P. Basic nuclear proteins in testicular cells and ejaculated spermatozoa in man. Exp Cell Res. 1978;117:347–56.
Ward WS, Coffey DS. DNA packaging and organization in mammalian spermatozoa: comparison with somatic cells. Biol Reprod. 1991;44:569–74.
Gannon JR, Emery BR, Jenkins TG, Carrell DT. The sperm epigenome: implications for the embryo. Adv Exp Med Biol New York, NY: Springer New York. 2014;791:53–66.
Gatewood J, Cook G, Balhorn R, Bradbury E, Schmid C. Sequence-specific packaging of DNA in human sperm chromatin. Science. 1987;236:962–4.
Zhang X. Sperm nuclear histone to protamine ratio in fertile and infertile men: evidence of heterogeneous subpopulations of spermatozoa in the ejaculate. J Androl. 2006;27:414–20.
Chevaillier P, Mauro N, Feneux D, Jouannet P, David G. Anomalous protein complement of sperm nuclei in some infertile men. Lancet. 1987;2:806–7.
de Mateo S, Ramos L, van der Vlag J, de Boer P, Oliva R. Improvement in chromatin maturity of human spermatozoa selected through density gradient centrifugation. Int J Androl. 2010;34:256–67.
Carrell DT, Liu L. Altered protamine 2 expression is uncommon in donors of known fertility, but common among men with poor fertilizing capacity, and may reflect other abnormalities of spermiogenesis. J Androl. 2001;22:604–10.
Carrell DT, Emery BR, Hammoud S. Altered protamine expression and diminished spermatogenesis: what is the link? Hum Reprod Update. 2007;13:313–27.
Aoki VW, Emery BR, Liu L, Carrell DT. Protamine levels vary between individual sperm cells of infertile human males and correlate with viability and DNA integrity. J Androl. 2006;27:890–8.
Aoki V, Liu L, Jones K, Hatasaka H, Gibson M, Peterson C, et al. Sperm protamine 1/protamine 2 ratios are related to in vitro fertilization pregnancy rates and predictive of fertilization ability. Fertil Steril. 2006;86:1408–15.
Hammoud SS, Nix DA, Zhang H, Purwar J, Carrell DT, Cairns BR. Distinctive chromatin in human sperm packages genes for embryo development. Nature Nature Publishing Group. 2009;460:473–8.
Arpanahi A, Brinkworth M, Iles D, Krawetz SA, Paradowska A, Platts AE, et al. Endonuclease-sensitive regions of human spermatozoal chromatin are highly enriched in promoter and CTCF binding sequences. Genome Res. 2009;19:1338–49.
Brykczynska U, Hisano M, Erkek S, Ramos L, Oakeley EJ, Roloff TC, et al. Repressive and active histone methylation mark distinct promoters in human and mouse spermatozoa. Nat Struct Mol Biol. 2010;17:679–87.
Jung YH, Sauria MEG, Lyu X, Cheema MS, Ausió J, Taylor J, et al. Chromatin states in mouse sperm correlate with embryonic and adult regulatory landscapes. Cell Rep. ElsevierCompany. 2017;18:1366–82.
Samans B, Yang Y, Krebs S, Sarode GV, Blum H, Reichenbach M, et al. Uniformity of nucleosome preservation pattern in mammalian sperm and its connection to repetitive DNA elements. Dev Cell. 2014;30:23–35.
Carone BR, Hung J-H, Hainer SJ, Chou M-T, Carone DM, Weng Z, et al. High-resolution mapping of chromatin packaging in mouse embryonic stem cells and sperm. Dev Cell 2014;1–20. Elsevier Inc.
Yuan ZF, Arnaudo AM, Garcia BA. Mass spectrometric analysis of histone proteoforms. Annu Rev Anal Chem (Palo Alto Calif). 2014;7:113–28.
Luense LJ, Wang X, Schon SB, Weller AH, Shiao EL, Bryant JM, et al. Comprehensive analysis of histone post-translational modifications in mouse and human male germ cells. Epigenetics Chromatin BioMed Central. 2016;9:1–15.
Kruger TF, Acosta AA, Simmons KF, Swanson RJ, Matta JF, Oehninger S. Predictive value of abnormal sperm morphology in in vitro fertilization. Fertil Steril. 1988;49:112–7.
Shechter D, Dormann HL, Allis CD, Hake SB. Extraction, purification and analysis of histones. Nat Protoc. 2007;2:1445–57.
Lin S, Garcia BA. Examining histone posttranslational modification patterns by high-resolution mass spectrometry. Methods Enzymol. 2012;512:3–28.
Govin J, Lestrat C, Caron C, Pivot-Pajot C, Rousseaux S, Khochbin S. Histone acetylation-mediated chromatin compaction during mouse spermatogenesis. Ernst Schering Res Found Workshop. 2006;57:155–72.
Boissonnas CC, Jouannet P, Jammes H. Epigenetic disorders and male subfertility. Fertil Steril. 2013;99:624–31.
Rathke C, Baarends WM, Awe S, Renkawitz-Pohl R. Chromatin dynamics during spermiogenesis. Biochim Biophys Acta. 1839;2014:155–68.
Goudarzi A, Shiota H, Rousseaux S, Khochbin S. Genome-scale acetylation-dependent histone eviction during spermatogenesis. J Mol Biol Elsevier Ltd. 2014;426:3342–9.
Dada R, Kumar M, Jesudasan R, Fernández JL, Gosálvez J, Agarwal A. Epigenetics and its role in male infertility. J Assist Reprod Genet. 2012;29:213–23.
Faure AK. Misregulation of histone acetylation in Sertoli cell-only syndrome and testicular cancer. Mol Hum Reprod. 2003;9:757–63.
Sonnack V, Failing K, Bergmann M, Steger K. Expression of hyperacetylated histone H4 during normal and impaired human spermatogenesis. Andrologia. 2002;34:384–90.
van Nuland R, Gozani O. Histone H4 lysine 20 (H4K20) methylation, expanding the signaling potential of the proteome one methyl moiety at a time. Mol Cell Proteomics. 2016;15:755–64.
Jørgensen S, Schotta G, Sørensen CS. Histone H4 lysine 20 methylation: key player in epigenetic regulation of genomic integrity. Nucleic Acids Res Oxford University Press. 2013;41:2797–806.
Grewal SIS, Jia S. Heterochromatin revisited. Nat Rev Genet. 2007;8:35–46.
Okada Y, Scott G, Ray MK, Mishina Y, Zhang Y. Histone demethylase JHDM2A is critical for Tnp1 and Prm1 transcription and spermatogenesis. Nature. 2007;450:119–23.
Peters A, O'Carroll D, Scherthan H, Mechtler K. Loss of the Suv39h histone methyltransferases impairs mammalian heterochromatin and genome stability. Cell. 2001;107:323–37.
van de Werken C, van der Heijden GW, Eleveld C, Teeuwssen M, Albert M, Baarends WM, et al. Paternal heterochromatin formation in human embryos is H3K9/HP1 directed and primed by sperm-derived histone modifications. Nat Commun. 2014;5:5868.
Steilmann C, Paradowska A, Bartkuhn M, Vieweg M, Schuppe HC, Bergmann M, et al. Presence of histone H3 acetylated at lysine 9 in male germ cells and its distribution pattern in the genome of human spermatozoa. Reprod Fertil Dev. 2011;23:997–15.
Vieweg M. Methylation analysis of histone H4K12ac-associated promoters in sperm of healthy donors and subfertile patients. Clin Epigenetics BioMed Central. 2015;7:1–17.
Hammoud SS, Nix DA, Hammoud AO, Gibson M, Cairns BR, Carrell DT. Genome-wide analysis identifies changes in histone retention and epigenetic modifications at developmental and imprinted gene loci in the sperm of infertile men. Hum Reprod. 2011;26:2558–69.
Acknowledgements
We would like to thank all of the staff at the Penn Fertility Care Andrology laboratory for their assistance, time, and effort in the de-identification of samples used in this study. In addition, we would like to thank The Penn Center for the Study of Epigenetics in Reproduction.
Funding
This work was funded by the National Institutes of Health Grants P50HD06817 (S.L.B./M.S.B./C.C/L.J.L), T32HD040135 (S.B.S/C.C), 5K12HD065257-07 (S.B.S), F32HD086939 (L.J.L.), and GM110174 (B.A.G).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. Approval for this study was obtained from the University of Pennsylvania Institutional Review Board (Protocol 815929).
Informed consent
As all samples were discarded and de-identified, the University of Pennsylvania Institutional Review Board determined that this study was exempt from requiring informed consent.
Electronic supplementary material
ESM 1
(DOCX 19 kb)
Rights and permissions
About this article
Cite this article
Schon, S.B., Luense, L.J., Wang, X. et al. Histone modification signatures in human sperm distinguish clinical abnormalities. J Assist Reprod Genet 36, 267–275 (2019). https://doi.org/10.1007/s10815-018-1354-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10815-018-1354-7