Abstract
Purpose
DNA methylation is an epigenetic mechanism that plays critical roles during mammalian oocyte and preimplantation embryo development. It is achieved by adding a methyl group to the fifth carbon atom of cytosine residues within cytosine-phosphate-guanine (CpG) and non-CpG dinucleotide sites using DNA methyltransferase (DNMT) enzymes for de novo and maintenance methylation processes. DNMT1, DNMT3A, and DNMT3B play important roles in establishing methylation of developmentally related genes in oocytes and early embryos. The purpose of this study is to identify the effect of superovulation on the expression and subcellular localizations of these three DNMT enzymes in the mouse oocytes and early embryos.
Methods
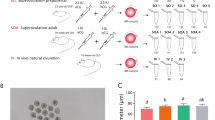
Three groups composed of control, normal dose [5 IU pregnant mare serum gonadotropin (PMSG) and 5 IU human chorionic gonadotropin (hCG)], and high dose [7.5 IU PMSG and 7.5 IU hCG] were created from 4–5-week-old female BALB/c mice. The relative expression and subcellular localizations of the DNMT proteins in the control and experiment groups have been characterized by using immunofluorescence staining subsequently analyzed in detailed.
Results
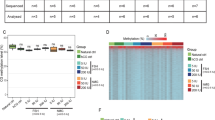
DNMT1, DNMT3A, and DNMT3B protein expression in the germinal vesicle and metaphase II oocytes and in one-cell and two-cell embryos differed significantly when some of the normal- and high-dose groups were compared with the control counterparts.
Conclusion
This study has demonstrated for the first time that superovulation alters expression levels of the DNMT proteins, a finding that indicates that certain developmental defects in superovulated oocytes and early embryos may result from impaired DNA methylation processes.




Similar content being viewed by others
References
Waldman E. Cultural priorities revealed: the development and regulation of assisted reproduction in the United States and Israel. Health Matrix. 2006;16:65–106.
Nyboe Andersen A, Goossens V, Bhattacharya S, Ferraretti AP, Kupka MS, de Mouzon J, et al. Assisted reproductive technology and intrauterine inseminations in Europe, 2005: results generated from European registers by ESHRE: ESHRE. The European IVF Monitoring Programme (EIM), for the European Society of Human Reproduction and Embryology (ESHRE). Hum Reprod. 2009;24:1267–87.
Huffman SR, Pak Y, Rivera RM. Superovulation induces alterations in the epigenome of zygotes, and results in differences in gene expression at the blastocyst stage in mice. Mol Reprod Dev. 2015;82:207–17.
Hansen M, Kurinczuk JJ, Bower C, Webb S. The risk of major birth defects after intracytoplasmic sperm injection and in vitro fertilization. N Engl J Med. 2002;346:725–30.
Schieve LA, Meikle SF, Ferre C, Peterson HB, Jeng G, Wilcox LS. Low and very low birth weight in infants conceived with use of assisted reproductive technology. N Engl J Med. 2002;346:731–7.
Halliday J, Oke K, Breheny S, Algar E, Amor DJ. Beckwith-Wiedemann syndrome and IVF: a case-control study. Am J Hum Genet. 2004;75:526–8.
Jiang Z, Wang Y, Lin J, Xu J, Ding G Huang H. Genetic and epigenetic risks of assisted reproduction. Best Pract Res Clin Obstet Gynaecol. 2017.
Meijerink AM, Oomen RE, Fleischer K, IntHout J, Woldringh GH, Braat DD. Effect of maternal and treatment-related factors on the prevalence of birth defects after PESA-ICSI and TESE-ICSI: a retrospective cohort study. Acta Obstet Gynecol Scand. 2015;94:1245–53.
Jwa J, Jwa SC, Kuwahara A, Yoshida A, Saito H. Risk of major congenital anomalies after assisted hatching: analysis of three-year data from the national assisted reproduction registry in Japan. Fertil Steril. 2015;104:71–8.
Pinborg A, Loft A, Romundstad LB, Wennerholm UB, Soderstrom-Anttila V, Bergh C, et al. Epigenetics and assisted reproductive technologies. Acta Obstet Gynecol Scand. 2016;95:10–5.
Uysal F, Akkoyunlu G, Ozturk S. Dynamic expression of DNA methyltransferases (DNMTs) in oocytes and early embryos. Biochimie. 2015;116:103–13.
Turek-Plewa J, Jagodzinski PP. The role of mammalian DNA methyltransferases in the regulation of gene expression. Cell Mol Biol Lett. 2005;10:631–47.
Bestor TH. The DNA methyltransferases of mammals. Hum Mol Genet. 2000;9:2395–402.
Barau J, Teissandier A, Zamudio N, Roy S, Nalesso V, Herault Y, et al. The DNA methyltransferase DNMT3C protects male germ cells from transposon activity. Science. 2016;354:909–12.
Fatemi M, Hermann A, Gowher H, Jeltsch A. Dnmt3a and Dnmt1 functionally cooperate during de novo methylation of DNA. Eur J Biochem. 2002;269:4981–4.
Margot JB, Ehrenhofer-Murray AE, Leonhardt H. Interactions within the mammalian DNA methyltransferase family. BMC Mol Biol. 2003;4:7.
Deplus R, Brenner C, Burgers WA, Putmans P, Kouzarides T, de Launoit Y, et al. Dnmt3L is a transcriptional repressor that recruits histone deacetylase. Nucleic Acids Res. 2002;30:3831–8.
Tajima S, Suetake I, Takeshita K, Nakagawa A, Kimura H. Domain structure of the Dnmt1, Dnmt3a, and Dnmt3b DNA methyltransferases. Adv Exp Med Biol. 2016;945:63–86.
Goll MG, Kirpekar F, Maggert KA, Yoder JA, Hsieh CL, Zhang X, et al. Methylation of tRNAAsp by the DNA methyltransferase homolog Dnmt2. Science. 2006;311:395–8.
El Hajj N, Haaf T. Epigenetic disturbances in in vitro cultured gametes and embryos: implications for human assisted reproduction. Fertil Steril. 2013;99:632–41.
Nelissen EC, Dumoulin JC, Daunay A, Evers JL, Tost J, van Montfoort AP. Placentas from pregnancies conceived by IVF/ICSI have a reduced DNA methylation level at the H19 and MEST differentially methylated regions. Hum Reprod. 2013;28:1117–26.
Li L, Wang L, Le F, Liu X, Yu P, Sheng J, et al. Evaluation of DNA methylation status at differentially methylated regions in IVF-conceived newborn twins. Fertil Steril. 2011;95:1975–9.
Whidden L, Martel J, Rahimi S, Chaillet JR, Chan D, Trasler JM. Compromised oocyte quality and assisted reproduction contribute to sex-specific effects on offspring outcomes and epigenetic patterning. Hum Mol Genet. 2016;25:4649–60.
Wang LY, Wang N, Le F, Li L, Lou HY, Liu XZ, et al. Superovulation induced changes of lipid metabolism in ovaries and embryos and its probable mechanism. PLoS One. 2015;10:e0132638.
Fauque P, Jouannet P, Lesaffre C, Ripoche MA, Dandolo L, Vaiman D, et al. Assisted reproductive technology affects developmental kinetics, H19 imprinting control region methylation and H19 gene expression in individual mouse embryos. BMC Dev Biol. 2007;7:116.
Akkoyunlu G, Ustunel I, Demir R. The distribution of transglutaminase in the rat oocytes and embryos. Theriogenology. 2007;68:834–41.
Ozturk S, Yaba-Ucar A, Sozen B, Mutlu D, Demir N. Superovulation alters embryonic poly(A)-binding protein (Epab) and poly(A)-binding protein, cytoplasmic 1 (Pabpc1) gene expression in mouse oocytes and early embryos. Reprod Fertil Dev. 2016;28:375–83.
Lawrence LT, Moley KH. Epigenetics and assisted reproductive technologies: human imprinting syndromes. Semin Reprod Med. 2008;26:143–52.
Van der Auwera I, D’Hooghe T. Superovulation of female mice delays embryonic and fetal development. Hum Reprod. 2001;16:1237–43.
Krisher RL. The effect of oocyte quality on development. J Anim Sci. 2004;82 E-Suppl:E14–23.
Ertzeid G, Storeng R. The impact of ovarian stimulation on implantation and fetal development in mice. Hum Reprod. 2001;16:221–5.
Whidden L, Martel J, Rahimi S, Richard Chaillet J, Chan D, Trasler JM. Compromised oocyte quality and assisted reproduction contribute to sex-specific effects on offspring outcomes and epigenetic patterning. Hum Mol Genet. 2016.
de Waal E, Vrooman LA, Fischer E, Ord T, Mainigi MA, Coutifaris C, et al. The cumulative effect of assisted reproduction procedures on placental development and epigenetic perturbations in a mouse model. Hum Mol Genet. 2015;24:6975–85.
Anckaert E, Fair T. DNA methylation reprogramming during oogenesis and interference by reproductive technologies: studies in mouse and bovine models. Reprod Fertil Dev. 2015;27:739–54.
Fortier AL, McGraw S, Lopes FL, Niles KM, Landry M, Trasler JM. Modulation of imprinted gene expression following superovulation. Mol Cell Endocrinol. 2014;388:51–7.
de Waal E, Yamazaki Y, Ingale P, Bartolomei MS, Yanagimachi R, McCarrey JR. Gonadotropin stimulation contributes to an increased incidence of epimutations in ICSI-derived mice. Hum Mol Genet. 2012;21:4460–72.
Allen WR, Moor RM. The origin of the equine endometrial cups. I. Production of PMSG by fetal trophoblast cells. J Reprod Fertil. 1972;29:313–6.
Kumar TR, Wang Y, Lu N, Matzuk MM. Follicle stimulating hormone is required for ovarian follicle maturation but not male fertility. Nat Genet. 1997;15:201–4.
Meduri G, Charnaux N, Driancourt MA, Combettes L, Granet P, Vannier B, et al. Follicle-stimulating hormone receptors in oocytes? J Clin Endocrinol Metab. 2002;87:2266–76.
Patsoula E, Loutradis D, Drakakis P, Kallianidis K, Bletsa R, Michalas S. Expression of mRNA for the LH and FSH receptors in mouse oocytes and preimplantation embryos. Reproduction. 2001;121:455–61.
Patsoula E, Loutradis D, Drakakis P, Michalas L, Bletsa R, Michalas S. Messenger RNA expression for the follicle-stimulating hormone receptor and luteinizing hormone receptor in human oocytes and preimplantation-stage embryos. Fertil Steril. 2003;79:1187–93.
Wada M, Seeger RC, Mizoguchi H, Koeffler HP. Maintenance of normal imprinting of H19 and IGF2 genes in neuroblastoma. Cancer Res. 1995;55:3386–8.
Market-Velker BA, Zhang L, Magri LS, Bonvissuto AC, Mann MR. Dual effects of superovulation: loss of maternal and paternal imprinted methylation in a dose-dependent manner. Hum Mol Genet. 2010;19:36–51.
Denomme MM, Zhang L, Mann MR. Embryonic imprinting perturbations do not originate from superovulation-induced defects in DNA methylation acquisition. Fertil Steril. 2011;96:734–8. e2.
Liang XW, Cui XS, Sun SC, Jin YX, Heo YT, Namgoong S, et al. Superovulation induces defective methylation in line-1 retrotransposon elements in blastocyst. Reprod Biol Endocrinol. 2013;11:69.
Funding
This study was supported by the Akdeniz University Scientific Research Projects Coordination Unit (Project Number: 2014.02.0122.013).
Author information
Authors and Affiliations
Contributions
F. Uysal performed the experiments. F. Uysal, S. Ozturk, and G. Akkoyunlu analyzed the data. F. Uysal, S. Ozturk, and G. Akkoyunlu wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Uysal, F., Ozturk, S. & Akkoyunlu, G. Superovulation alters DNA methyltransferase protein expression in mouse oocytes and early embryos. J Assist Reprod Genet 35, 503–513 (2018). https://doi.org/10.1007/s10815-017-1087-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10815-017-1087-z