Abstract
Purpose
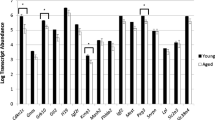
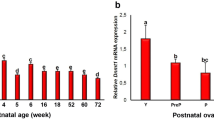
This study was designed to evaluate DNA methylation and the expression of DNA methyltransferases (Dnmt1, Dnmt3a, Dnmt3b and Dnmt3L) in metaphaseII (MII) oocytes and the DNA methylation of pre-implantation embryos during mouse aging to address whether such aging-related changes are associated with decreased reproductive potential in aged mice.
Methods
Oocytes (MII) from 6 to 8 weeks old female mice are referred to as the ‘young group’; oocytes from the same group that were maintained until 35–40 weeks old are referred to as the ‘old group.’ The oocytes were fertilized both in vitro and in vivo to obtain embryos. The DNA methylation levels in the oocytes (MII) and pre-implantation embryos were assessed using fluorescence staining. The expression levels of the Dnmt genes in the oocytes (MII) were assessed using Western blotting.
Results
The DNA methylation levels in the oocytes and pre-implantation embryos (in vivo and in vitro) decreased significantly during the aging of the mice. The expression levels of all of the examined Dnmt proteins in the old group were lower than young group. Both the cleavage and blastocyst rate were significantly lower in the oocytes of the older mice (69.9 % vs. 80.9 %, P < 0.05; 33.9 % vs. 56.4 %, P < 0.05). The pregnancy rate of the old mice was lower than that of the young mice (46.7 % vs. 100 %, P < 0.05). The stillbirth and fetal malformation rate was significantly higher in the old group than in the young group (17.2 % vs. 2.9 %, P < 0.05).
Conclusions
The decreased expression of Dnmt1, Dnmt3a, Dnmt3b and Dnmt3L in oocytes (MII) and the change of genome-wide DNA methylation in oocytes and pre-implantation embryos due to aging may be related to lower reproductive potential in old female mice.



Similar content being viewed by others
References
Armstrong DT. Effects of maternal age on oocyte developmental competence. Theriogenology. 2001;55(6):1303–22.
Klein J, Sauer MV. Assessing fertility in women of advanced reproductive age. Am J Obstet Gynecol. 2001;185(3):758–70.
Assisted reproductive technology in the United States: 1997 results generated from the American Society for Reproductive Medicine/Society for Assisted Reproductive Technology Registry. Fertil Steril. 2000;74(4):641–653; discussion 653–644.
van Kooij RJ, Looman CW, Habbema JD, Dorland M, te Velde ER. Age-dependent decrease in embryo implantation rate after in vitro fertilization. Fertil Steril. 1996;66(5):769–75.
Plachot M, Veiga A, Montagut J, et al. Are clinical and biological IVF parameters correlated with chromosomal disorders in early life: a multicentric study. Hum Reprod. 1988;3(5):627–35.
Hamatani T, Falco G, Carter MG, et al. Age-associated alteration of gene expression patterns in mouse oocytes. Hum Mol Genet. 2004;13(19):2263–78.
Wise PM, Smith MJ, Dubal DB, et al. Neuroendocrine modulation and repercussions of female reproductive aging. Recent Prog Horm Res. 2002;57:235–56.
Yeh J, Kim BS, Peresie J. Ovarian vascular endothelial growth factor and vascular endothelial growth factor receptor patterns in reproductive aging. Fertil Steril. 2008;89(5 Suppl):1546–56.
Thorneycroft IH, Soderwall AL. The nature of the litter size loss in senescent hamsters. Anat Rec. 1969;165(3):343–8.
Finn CA. Reproductive ageing and the menopause. Int J Dev Biol. 2001;45(3):613–7.
te Velde ER, Pearson PL. The variability of female reproductive ageing. Hum Reprod Update. 2002;8(2):141–54.
Ito M, Muraki M, Takahashi Y, et al. Glutathione S-transferase theta 1 expressed in granulosa cells as a biomarker for oocyte quality in age-related infertility. Fertil Steril. 2008;90(4):1026–35.
Pan H, Ma P, Zhu W, Schultz RM. Age-associated increase in aneuploidy and changes in gene expression in mouse eggs. Dev Biol. 2008;316(2):397–407.
Chiang T, Duncan FE, Schindler K, Schultz RM, Lampson MA. Evidence that weakened centromere cohesion is a leading cause of age-related aneuploidy in oocytes. Curr Biol. 2010;20(17):1522–8.
Antinori S, Versaci C, Gholami GH, Panci C, Caffa B. Oocyte donation in menopausal women. Hum Reprod. 1993;8(9):1487–90.
Sauer MV, Paulson RJ, Lobo RA. Reversing the natural decline in human fertility. An extended clinical trial of oocyte donation to women of advanced reproductive age. JAMA. 1992;268(10):1275–9.
Fraga MF, Esteller M. Epigenetics and aging: the targets and the marks. Trends Genet. 2007;23(8):413–8.
Fraga MF, Agrelo R, Esteller M. Cross-talk between aging and cancer: the epigenetic language. Ann N Y Acad Sci. 2007;1100:60–74.
Munoz-Najar U, Sedivy JM. Epigenetic control of aging. Antioxid Redox Signal. 2011;14(2):241–59.
Trasler JM. Gamete imprinting: setting epigenetic patterns for the next generation. Reprod Fertil Dev. 2006;18(1–2):63–9.
Robertson KD, Wolffe AP. DNA methylation in health and disease. Nat Rev Genet. 2000;1(1):11–9.
Jurkowska RZ, Jurkowski TP, Jeltsch A. Structure and function of mammalian DNA methyltransferases. ChemBioChem. 2010;12(2):206–22.
Reik W, Dean W, Walter J. Epigenetic reprogramming in mammalian development. Science. 2001;293(5532):1089–93.
Golbus J, Palella TD, Richardson BC. Quantitative changes in T cell DNA methylation occur during differentiation and ageing. Eur J Immunol. 1990;20(8):1869–72.
Wilson VL, Smith RA, Ma S, Cutler RG. Genomic 5-methyldeoxycytidine decreases with age. J Biol Chem. 1987;262(21):9948–51.
Vanyushin BF, Nemirovsky LE, Klimenko VV, Vasiliev VK, Belozersky AN. The 5-methylcytosine in DNA of rats. Tissue and age specificity and the changes induced by hydrocortisone and other agents. Gerontologia. 1973;19(3):138–52.
Issa JP, Ottaviano YL, Celano P, Hamilton SR, Davidson NE, Baylin SB. Methylation of the oestrogen receptor CpG island links ageing and neoplasia in human colon. Nat Genet. 1994;7(4):536–40.
Eads CA, Lord RV, Wickramasinghe K, et al. Epigenetic patterns in the progression of esophageal adenocarcinoma. Cancer Res. 2001;61(8):3410–8.
Kang GH, Lee HJ, Hwang KS, Lee S, Kim JH, Kim JS. Aberrant CpG island hypermethylation of chronic gastritis, in relation to aging, gender, intestinal metaplasia, and chronic inflammation. Am J Pathol. 2003;163(4):1551–6.
Shen L, Ahuja N, Shen Y, et al. DNA methylation and environmental exposures in human hepatocellular carcinoma. J Natl Cancer Inst. 2002;94(10):755–61.
Tsuchiya T, Tamura G, Sato K, et al. Distinct methylation patterns of two APC gene promoters in normal and cancerous gastric epithelia. Oncogene. 2000;19(32):3642–6.
Waki T, Tamura G, Sato M, Motoyama T. Age-related methylation of tumor suppressor and tumor-related genes: an analysis of autopsy samples. Oncogene. 2003;22(26):4128–33.
Tang WY, Ho SM. Epigenetic reprogramming and imprinting in origins of disease. Rev Endocr Metab Disord. 2007;8(2):173–82.
Morgan HD, Santos F, Green K, Dean W, Reik W. Epigenetic reprogramming in mammals. Hum Mol Genet. 2005;14:R47–58.
Worrad DM, Ram PT, Schultz RM. Regulation of gene expression in the mouse oocyte and early preimplantation embryo: developmental changes in Sp1 and TATA box-binding protein, TBP. Development. 1994;120(8):2347–57.
Aoki F, Worrad DM, Schultz RM. Regulation of transcriptional activity during the first and second cell cycles in the preimplantation mouse embryo. Dev Biol. 1997;181(2):296–307.
Agarwal S, Rao A. Modulation of chromatin structure regulates cytokine gene expression during T cell differentiation. Immunity. 1998;9(6):765–75.
Richardson B. Impact of aging on DNA methylation. Ageing Res Rev. 2003;2(3):245–61.
Feinberg AP. DNA methylation, genomic imprinting and cancer. Curr Top Microbiol Immunol. 2000;249:87–99.
Feinberg AP, Cui H, Ohlsson R. DNA methylation and genomic imprinting: insights from cancer into epigenetic mechanisms. Semin Cancer Biol. 2002;12(5):389–98.
Wilson VL, Jones PA. DNA methylation decreases in aging but not in immortal cells. Science. 1983;220(4601):1055–7.
Cedar H, Razin A. DNA methylation and development. Biochim Biophys Acta. 1990;1049(1):1–8.
Dahl C, Guldberg P. DNA methylation analysis techniques. Biogerontology. 2003;4(4):233–50.
Eden S, Hashimshony T, Keshet I, Cedar H, Thorne AW. DNA methylation models histone acetylation. Nature. 1998;394(6696):842.
Suo L, Meng QG, Pei Y, et al. Changes in acetylation on lysine 12 of histone H4 (acH4K12) of murine oocytes during maternal aging may affect fertilization and subsequent embryo development. Fertil Steril. 2010;93(3):945–51.
Bachman KE, Park BH, Rhee I, et al. Histone modifications and silencing prior to DNA methylation of a tumor suppressor gene. Canc Cell. 2003;3(1):89–95.
Lehnertz B, Ueda Y, Derijck AA, et al. Suv39h-mediated histone H3 lysine 9 methylation directs DNA methylation to major satellite repeats at pericentric heterochromatin. Curr Biol. 2003;13(14):1192–200.
Strunnikova M, Schagdarsurengin U, Kehlen A, Garbe JC, Stampfer MR, Dammann R. Chromatin inactivation precedes de novo DNA methylation during the progressive epigenetic silencing of the RASSF1A promoter. Mol Cell Biol. 2005;25(10):3923–33.
Tamaru H, Selker EU. A histone H3 methyltransferase controls DNA methylation in Neurospora crassa. Nature. 2001;414(6861):277–83.
Manosalva I, Gonzalez A. Aging changes the chromatin configuration and histone methylation of mouse oocytes at germinal vesicle stage. Theriogenology. 2010;74(9):1539–47.
Li E, Bestor TH, Jaenisch R. Targeted mutation of the DNA methyltransferase gene results in embryonic lethality. Cell. 1992;69(6):915–26.
Hata K, Okano M, Lei H, Li E. Dnmt3L cooperates with the Dnmt3 family of de novo DNA methyltransferases to establish maternal imprints in mice. Development. 2002;129(8):1983–93.
Okano M, Bell DW, Haber DA, Li E. DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell. 1999;99(3):247–57.
Feil R, Khosla S. Genomic imprinting in mammals: an interplay between chromatin and DNA methylation? Trends Genet. 1999;15(11):431–5.
Lopes FL, Fortier AL, Darricarrere N, Chan D, Arnold DR, Trasler JM. Reproductive and epigenetic outcomes associated with aging mouse oocytes. Hum Mol Genet. 2009;18(11):2032–44.
Santos F, Dean W. Using immunofluorescence to observe methylation changes in mammalian preimplantation embryos. Meth Mol Biol. 2006;325:129–37.
Acknowledgments
This work was supported by the National Natural Science Foundation Project of China (No. 30972102); China Agricultural University Graduate Scientific Research and Innovation Special Project (No. kycx09027). We thank Professor William Hohenboken, Professor Tiantian Zhang and Dr. QingGang Meng for proofreading the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Capsule
Aging caused a significant decrease in the expression of four key Dnmts and genome-wide DNA methylation in oocytes and pre-implantation embryos.
Ming-xing Yue and Xiang-wei Fu contributed equally to this work.
Rights and permissions
About this article
Cite this article
Yue, Mx., Fu, Xw., Zhou, Gb. et al. Abnormal DNA methylation in oocytes could be associated with a decrease in reproductive potential in old mice. J Assist Reprod Genet 29, 643–650 (2012). https://doi.org/10.1007/s10815-012-9780-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10815-012-9780-4