Abstract
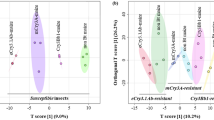
Genetically modified organisms (GMOs) need to be evaluated for safety before their release. Metabolome techniques provide effective methods to evaluate unintended effects. In this study, the grain metabolome of six transgenic rice lines containing secticidal cry and glyphosate tolerance epsps genes were tested by using an non-targeted metabolite profiling, and 161 and 138 metabolites were identified in grains at grain-filling stage and mature stage, respectively. The metabolic profiles of genetically modified (GM) lines and non-GM lines were all significantly different from each other at grain-filling stage. Although the levels of many metabolites were significantly changed in each transgenic line, only seven of them were simultaneously changed in all the six lines compared with those in non-GM lines at grain-filling stage. The number of significantly changed metabolites was much less at mature stage than that at grain-filling stage. Besides, none of these metabolites was simultaneously changed in all the six lines, suggesting that the metabolites of GMOs are different at different stages of GMO development. This study provides useful information about the metabolic variation between GMO and non-GMO in two development stages of rice grains. These findings suggest that non-targeted metabolite profiling can strengthen the assessment of risk based on metabolites.



Similar content being viewed by others
References
Aarab S, Arakrak A, Ollero FJ, Megias M, Gomes DF, Ribeiro RA, Hungria M (2016) Draft genome sequence of pseudomonas fluorescens strain ET76, isolated from rice Rhizosphere in Northwestern Morocco. Genome Announc. https://doi.org/10.1128/genomeA.00356-16
Azevedo RA, Lancien M, Lea PJ (2006) The aspartic acid metabolic pathway, an exciting and essential pathway in plants. Amino Acids 30:143–162
Barros E, Lezar S, Anttonen MJ, van Dijk JP, Rohlig RM, Kok EJ, Engel KH (2010) Comparison of two GM maize varieties with a near-isogenic non-GM variety using transcriptomics, proteomics and metabolomics Plant. Biotechnol J 8:436–451. https://doi.org/10.1111/j.1467-7652.2009.00487.x
Caldovic L, Tuchman M (2003) N-acetylglutamate and its changing role through evolution. Biochem J 372:279–290
Catchpole GS et al (2005) Hierarchical metabolomics demonstrates substantial compositional similarity between genetically modified and conventional potato crops. Proc Natl Acad Sci USA 102:14458–14462. https://doi.org/10.1073/pnas.0503955102
Coll A et al (2009) Gene expression profiles of MON810 and comparable non-GM maize varieties cultured in the field are more similar than are those of conventional lines. Transgenic Res 18:801–808. https://doi.org/10.1007/s11248-009-9266-z
Coll A, Nadal A, Collado R, Capellades G, Kubista M, Messeguer J, Pla M (2010) Natural variation explains most transcriptomic changes among maize plants of MON810 and comparable non-GM varieties subjected to two N-fertilization farming practices. Plant Mol Biol 73:349–362. https://doi.org/10.1007/s11103-010-9624-5
Dong W et al (2008) GMDD: a database of GMO detection methods. BMC Bioinform 9:260. https://doi.org/10.1186/1471-2105-9-260
Farag KM, Palta JP (2010) Use of lysophosphatidylethanolamine, a natural lipid, to retard tomato leaf and fruit senescence. Physiol Plant 87:515–521
Gamir J, Pastor V, Cerezo M, Flors V (2012) Identification of indole-3-carboxylic acid as mediator of priming against Plectosphaerella cucumerina. Plant Physiol Biochem 61:169–179
Hong JH, Sungkee H, Gukhoon C (2008) Influence of lysophosphatidylethanolamine on reactive oxygen species, ethylene biosynthesis, and auxin action in plant tissue. Kor J Hortic Sci Technol 26:209–214
Horai H et al (2010) MassBank: a public repository for sharing mass spectral data for life sciences. J Mass Spectrom 45:703–714. https://doi.org/10.1002/jms.1777
Hu C et al (2016) Identification of conserved and diverse metabolic shifts during rice grain. Dev Sci Rep 6:20942. https://doi.org/10.1038/srep20942
Hu C, Zhao H, Wang W, Xu M, Shi J, Nie X, Yang G (2018) Identification of conserved and diverse metabolic shift of the stylar, intermediate and peduncular segments of cucumber fruit during development. Int J Mol Sci https://doi.org/10.3390/ijms19010135
Kalamaki MS et al (2009) Over-expression of a tomatoN-acetyl-L-glutamate synthase gene (SlNAGS1) in Arabidopsis thaliana results in high ornithine levels and increased tolerance in salt and drought stresses. J Exp Bot 60:1859–1871
Ladics GS et al (2015) Genetic basis and detection of unintended effects in genetically modified crop plants. Transgenic Res 24:587–603. https://doi.org/10.1007/s11248-015-9867-7
Montero M, Coll A, Nadal A, Messeguer J, Pla M (2011) Only half the transcriptomic differences between resistant genetically modified and conventional rice are associated with the transgene. Plant Biotechnol J 9:693–702. https://doi.org/10.1111/j.1467-7652.2010.00572.x
Negre F et al (2003) Regulation of methylbenzoate emission after pollination in snapdragon and petunia flowers. Plant Cell 15:2992
Ozgen M, Palta JP (1999) Use of lysophosphatidylethanolamine (LPE), a natural lipid, to accelerate ripening and enhance shelf life of cranberry fruit. Hortscience 34:538
Özgen M, SerçE S, AkçA Y, Hong JH (2015) Lysophosphatidylethanolamine (LPE) improves fruit size, color, quality and phytochemical contents of sweet cherry cv. ‘0900 Ziraat’. Wonye kwahak kisulchi = Korean J Hortic Sci Technol 33:196–201
Paxton J (1980) A new working definition of the term “Phytoalexin”. Plant Dis 64:734
Peng C, Chen X, Wang X, Xu X, Wei W, Wang C, Xu J (2018) Comparative analysis of miRNA expression profiles in transgenic and non-transgenic rice using miRNA. Seq Sci Rep 8:338. https://doi.org/10.1038/s41598-017-18723-x
Ricroch AE, Berge JB, Kuntz M (2011) Evaluation of genetically engineered crops using transcriptomic, proteomic, and metabolomic profiling techniques. Plant Physiol 155:1752–1761. https://doi.org/10.1104/pp.111.173609
Schnell J et al (2015) A comparative analysis of insertional effects in genetically engineered plants: considerations for pre-market assessments. Transgenic Res 24:1–17. https://doi.org/10.1007/s11248-014-9843-7
Ubaidillah M et al (2016) Roles of plant hormones and anti-apoptosis genes during drought stress in rice (Oryza sativa L.). Biotech 6:247
Wang XJ, Zhang X, Yang JT, Wang ZX (2018) Effect on transcriptome and metabolome of stacked transgenic maize containing insecticidal cry and glyphosate tolerance epsps genes. Plant J 93:1007–1016. https://doi.org/10.1111/tpj.13825
Yamamura C et al (2016) Diterpenoid phytoalexin factor, a bHLH transcription factor, plays a central role in the biosynthesis of diterpenoid phytoalexins in rice. Plant J 84:1100–1113
Zhou J et al (2009) Metabolic profiling of transgenic rice with cryIAc and sck genes: an evaluation of unintended effects at metabolic level by using GC-FID and GC-MS. J Chromatogr 877:725–732. https://doi.org/10.1016/j.jchromb.2009.01.040
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant Nos. 31501280, 31671753) and the Natural Science Foundation of Zhejiang Province (Grant No. LQ15C130002).
Author information
Authors and Affiliations
Contributions
CP and JX designed the research. LD and XW carried out the field tests. CP and XC performed sample preparation. CH and CP carried out the metabolic profiling, the annotation of the metabolites and data analysis. CP, CH and JX wrote the manuscript. XC supervised the research.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Peng, C., Ding, L., Hu, C. et al. Effect on metabolome of the grains of transgenic rice containing insecticidal cry and glyphosate tolerance epsps genes. Plant Growth Regul 88, 1–7 (2019). https://doi.org/10.1007/s10725-019-00482-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-019-00482-6