Abstract
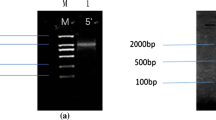
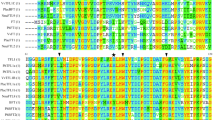
AP1/SQUA-like gene plays important roles in the development of inflorescences and flowers. In this study, an APETALA1 (AP1) homologue, EjAP1, was isolated from loquat (Eriobotrya japonica Lindl.), an economically important subtropical fruit of the Rosaceae. EjAP1 was sequenced and found to be 3,229 bp in length with 6 introns and 7 exons and encoded 239 amino acids. The deduced amino acid sequence contained the typical CaaX-motif of the AP1 functional proteins and shared high homology with the other AP1 genes. Phylogenetic analysis at the amino acid level indicated that EjAP1 belongs to the AP1/SQUA subfamily. Transcriptions of EjAP1 were detected in inflorescence bud, flower bud and flower, but not in vegetative tissues such as vegetative bud. In floral organs, highest expression of EjAP1 was detected in sepal compared to lower transcription level in petal and stamen, and no transcription in pistil. When ectopic expressed in ap1-1 mutant of Arabidopsis, EjAP1 can partially complement the mutant mainly in that it can restore the sepal and petal formation of ap1-1 mutant. These results indicated that EjAP1 gene may play a similar role as Arabidopsis AP1 in the floral development of loquat.









Similar content being viewed by others
References
Abe M, Kobayashi Y, Yamamoto S, Daimon Y, Yamaguchi A, Ikeda Y, Ichinoki H, Notaguchi M, Goto K, Araki T (2005) FD, a bZIP protein mediating signals from the floral pathway integrator FT at the shoot apex. Science 309:1052–1056
Amasino R (2010) Seasonal and developmental timing of flowering. The Plant J 61:1001–1013
Asif MH, Dhawan P, Math P (2000) A simple procedure for the isolation of high quality RNA from ripening banana fruit. Plant Mol Biol Rep 18:109–115
Badenes ML, Lin S, Yang X, Liu C, Huang X (2009) Loquat (Eriobotrya Lindl.). In: Folta KM and Gardiner SE (ed) Genetics and genomics of Rosaceae. Springer Science & Business Media, New York. pp525–538
Becker A, Theißen G (2003) The major clades of MADS-box genes and their role in the development and evolution of flowering plants. Mol Phylogenet Evol 29:464–489
Bowman JL, Smyth DR, Meyerowitz EM (1991) Genetic interactions among floral homeotic genes of Arabidopsis. Development 112:1–20
Bowman JL, Alvarez J, Weigel D, Meyerowizt EM, Smyth DR (1993) Control of flower development in Arabidopsis thaliana by APETALA1 and interacting genes. Development 119:721–743
Calonje M, Cubas P, Martínez-Zapater JM, Carmona MJ (2004) Floral meristem identity genes are expressed during tendril development in grapevine. Plant Physiol 135:1491–1501
Chi Y, Huang F, Liu H, Yang S, Yu D (2011) An APETALA1-like gene of soybean regulates flowering time and specifies floral organs. J Plant Physiol 168:2251–2259
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Coen ES, Meyerowitz EM (1991) The war of the whorls: genetic interactions controlling flower development. Nature 353:31–37
De Bodt S, Raes J, Van de Peer YV, Theißen G (2003) And then there were many: MADS goes genomic. Trends Plant Sci 8:475–483
Egea-Cortines M, Saedler H, Sommer H (1999) Ternary complex formation between the MADS-box proteins SQUAMOSA, DEFICIENS and GLOBOSA is involved in the control of floral architecture in Antirrhinum majus. EMBO J 18:5370–5379
Flachowsky H, Peil A, Sopanen T, Elo A, Hanke V (2007) Overexpression of BpMADS4 from silver birch (Betula pendula Roth.) induces early flowering in apple (Malus × domestica Borkh.). Plant Breed 126:137–145
Higgins D, Thompson J, Gibson T, Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22(22):4673–4680
Huijser P, Klein J, Lönnig WE, Meijer H, Saedler H, Sommer H (1992) Bracteomania, an inflorescence anomaly, is caused by the loss of function of the MADS-box gene squamosa in Antirrhinum majus. EMBO J 11:1239–1249
Irish VF, Sussex IM (1990) Function of the apetala-1 gene during Arabidopsis floral development. Plant Cell 2:741–753
Kaufmann K, Melzer S, Theissen G (2005) MIKC-type MADS-domain proteins: structural modularity, protein interactions and network evolution in land plants. Gene 347:183–198
Kaufmann K, Wellmer F, Muiño JM, Ferrier T, Wuest SE, Kumar V, Serrano-Mislata A, Madueño F, Krajewski P, Meyerowitz EM, Angenent GC, Riechmann JL (2010) Orchestration of floral initiation by APETALA1. Science 328:85–89
Kieffer M, Davies MB (2001) Developmental programmes in floral organ formation. Semin Cell Dev Biol 12:373–380
Kotoda N, Wada M, Komori S, Kidou S, Abe K, Masuda T, Soejima J (2000) Expression pattern of homologues of floral meristem identity genes LFY and AP1 during flower development in apple. J Am Soc Hortic Sci 125:398–403
Kotoda N, Wada M, Kusaba S, Murakami YK, Masuda T, Soejima J (2002) Overexpression of MdMADS5, an APETALA1-like gene of apple, causes early flowering in transgenic Arabidopsis. Plant Sci 162:679–687
Kotoda N, Iwanami H, Takahashi S, Abe K (2006) Antisense expression of MdTFL1, a TFL1-like gene, reduces the juvenile phase in apple. J Am Soc Hortic Sci 131:74–81
Li C, Xie H, Zhang L, Xu Y, Li YF, Ma RC (2012) Molecular characterization of the PpMADS1 gene from peach. Tree Genet Genome 8(4):831–840
Lin SQ, Sharpe RH, Janick J (1999) Loquat: botany and horticulture. Hortic Rev 23:233–276
Litt A, Irish VF (2003) Duplication and diversification in the APETALA1/FRUITFULL floral homeotic gene lineage: implications for the evolution of floral development. Genetics 165:821–833
Lu SJ, Wei H, Wang Y, Wang HM, Yang RF, Zhang XB, Tu JM (2012) Overexpression of a transcription factor OsMADS15 modifies plant architecture and flowering time in rice (Oryza sativa L.). Plant Mol Biol Rep 30(6):1461–1469
Mandel MA, Yanofsky MF (1995) A gene triggering flower formation in Arabidopsis. Nature 377:522–524
Mandel MA, Gustafson-Brown C, Savidge B, Yanofsky MF (1992) Molecular characterization of the Arabidopsis floral homeotic gene APETALA-1. Nature 360:273–277
Masiero S, Imbriano C, Ravasio F, Favaro R, Pelucchi N, Gorla MS, Mantovani R, Colombo L, Kater MM (2002) Ternary complex formation between MADS-box transcription factors and the histone fold protein NF-YB. J Biol Chem 277:26429–26435
Peña L, Martin-Trillo M, Juarez J, Pina JA, Navarro L, Martinez-Zapater JM (2001) Constitutive expression of Arabidopsis LEAFY or APETALA1 genes in citrus reduces their generation time. Nat Biotech 19:263–267
Pidkowich MS, Klenz JE, Haughn GW (1999) The making of a flower: control of floral meristem identity in Arabidopsis. Trends Plant Sci 4:64–70
Pillitterri LJ, Lovatt CJ, Walling LL (2004) Isolation and characterization of LEAFY and APETALA1 homologues from Citrus sinensis L. Osbeck ‘Washington’. J Am Soc Hortic Sci 129:846–856
Shan H, Zhang N, Liu C, Xu G, Zhang J, Chen Z, Kong H (2007) Patterns of gene duplication and functional diversification during the evolution of the AP1/SQUA subfamily of plant MADS-box genes. Mol Phylogenet Evol 44:26–41
Simpson GG, Gendall AR, Dean C (1999) When to switch to flowering. Annu Rev Cell Dev Biol 15:519–550
Srikanth A, Schmid M (2011) Regulation of flowering time: all roads lead to Rome. Cell Mol Life Sci 68:2013–2037
Sung SK, Yu GH, An G (1999) Characterization of MdMADS2, a member of the SQUAMOSA subfamily of genes, in apple. Plant Physiol 120:969–978
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28(10):2731–2739
Theissen G (2001) Development of floral organ identity: stories from the MADS house. Curr Opin Plant Biol 4:75–85
Tränkner C, Lehmann S, Hoenicka H, Hanke M, Fladung M, Lenhardt D, Dunemann F, Gau A, Schlangen K, Malnoy M, Flachowsky H (2010) Over-expression of an FT-homologous gene of apple induces early flowering in annual and perennial plants. Planta 232:1309–1324
Tsaftaris AS, Pasentsis K, Iliopoulos I, Polidoros AN (2004) Isolation of three homologous AP1-like MADS-box genes in crocus (Crocus sativus L.) and characterization of their expression. Plant Sci 166:1235–1243
van Dijk ADJ, Morabito G, Fiers M, van Ham RCHJ, Angenent GC, Immink RGH (2010) Sequence motifs in MADS transcription factors responsible for specificity and diversification of protein protein interaction. PLoS Comput Biol 6:e1001017
Wagner D, Sablowski RWM, Meyerowitz EM (1999) Transcriptional activation of APETALA1 by LEAFY. Science 285:582–584
Wellmer F, Riechmann JL (2010) Gene networks controlling the initiation of flower development. Trends Genet 26:519–527
Wigge PA, Kim MC, Jaeger KE, Busch W, Schmid M, Lohmann J, Weigel D (2005) Integration of spatial and temporal information during floral induction in Arabidopsis. Science 309:1056–1059
Yalovsky S, Rodriguez-Concepcion M, Bracha K, Toledo-Ortiz G, Gruissem W (2000) Prenylation of the floral transcription factor APETALA1 modulates its function. Plant Cell 12:1257–1266
Zik M, Irish VF (2003) Flower development: initiation, differentiation, and diversification. Annu Rev Cell Dev Biol 19:119–140
Acknowledgments
This study was financially supported by grants of Key Laboratory of Innovation and Utilization for Germplasm Resources in Horticultural Crops in South China of Guangdong Higher Education Institutes, South China Agricultural University (no. KBL11008).
Author information
Authors and Affiliations
Corresponding author
Additional information
Yuexue Liu and Huwei Song: contributed equally to this work.
Rights and permissions
About this article
Cite this article
Liu, Y., Song, H., Liu, Z. et al. Molecular characterization of loquat EjAP1 gene in relation to flowering. Plant Growth Regul 70, 287–296 (2013). https://doi.org/10.1007/s10725-013-9800-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-013-9800-0