Abstract
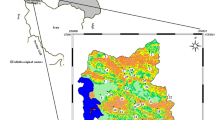
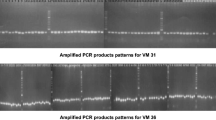
Citrullus lanatus ssp. vulgaris oleaginous type (West-African watermelon) is a crop cultivated in sub-Saharan Africa for its dried seeds reported to be rich in nutrients. In previous studies, little polymorphism was found in watermelon—cultivated for its flesh with the use of microsatellite (SSR) markers. Such study has never been applied to the oleaginous type until now. The objectives of the present study were firstly to apply the SSR markers set up for watermelon to the West-African watermelon and secondly to study the genetic structure of this type in Ivory Coast. For the first objective, 37 markers were studied on eight plants pertaining to four accessions. For the second objective, the polymorphic markers were applied on three morphologically and geographically separated accessions with twenty plants per accession. Multiple correspondence analysis (MCA), unweighted pair-group method with arithmetic averaging (UPGMA), molecular analysis of variance (AMOVA) and assignments test structure were applied. The optimal annealing temperature varied from 49 to 59°C according to the markers. Thirty-two markers that proved to amplify their respective loci were selected, but only nine of them appeared to show polymorphism on the set of 8 plants studied. The application of these markers on the three accessions revealed several features. No stucturation into sub-populations was observed inside a given accession. The genetic variance proved to be substantially higher between the different accessions than inside a given accession. Moreover this analysis is a first hint that the morphology classification does not match the genetic structure of C. lanatus. The results of this work provide the first quantitative information regarding the genetic variability of Citrullus lanatus oleaginous type. In order to sharpen our understanding of the mechanisms responsible for the genetic variance inter/intra accessions, further studies based on a larger sample of plants and accessions are required.




Similar content being viewed by others
Notes
Levi Ammon: USDA–ARS, U.S. Vegetable Laboratory, Charleston, SC 29414.
References
Bowcock AM, Ruiz-Linares A, Tomforhde J, Minch E, Kidd JR, Cavalli-Sforza LL (1994) High resolution of human evolutionary trees with polymorphic microsatellites. Lett Nat 368:454–455
Dane F, Liu J (2007) Diversity and origin of cultivated and citron type watermelon (Citrullus lanatus). Genet Resour Crop Evol 54:1255–1265
Djè Y, Tahi GC, Zoro Bi IA, Malice M, Baudoin J-P, Bertin P (2006) Optimization of ISSR markers for African edible-seeded Cucurbitaceae species genetic diversity analysis. Afr J Biotechnol 5:83–87
Ducarme V, Risterucci AM, Wesselingh RA (2008) Development of microsatellite markers in Rhinanthus angustifolius and cross-species amplification. Mol Ecol Resour 8:384–386
Ferreira MAJ, Vencovsky R, Vieira MLC, de Queiróz MA (2000) Outcrossing rate and implications for the improvement of a segregating population of watermelon. Acta Hortic 510:47–54
Guerra-Sanz J (2002) Citrullus simple sequence repeats markers from sequence databases. Mol Ecol Notes 2:223–225
Gusmini G (2005) Inheritance of fruit characteristics and disease resistance in watermelon [Citrullus lanatus (Thunb.) Matsum. and Nakai]. PhD thesis, Graduate Faculty of North Carolina State University
Gusmini G, Wehner T (2003) Polygenic inheritance of some vine traits in two segregating watermelon families. Cucurbit Genet Coop Rep 26:32–35
Gusmini G, Wehner T, Jarret R (2004) Inheritance of egusi seed type in watermelon. J Hered 95:268–270
Jarret R, Merrick L, Holms T, Evans J, Aradhya M (1997) Simple sequence repeats in watermelon Citrullus lanatus (Thunb.) Matsum. & Nakai. Genome 40:433–441
Joobeur T, Zhang X, Xu Y, Wehner T, Gusmini G, Levi A, Oliver M, Dean R (2006) Construction of a watermelon BAC library and identification of SSRs anchored to melon or Arabidopsis genomes. Theor Appl Genet 112:1553–1562
Kwon Y-S, Park EK, Lee W-S, Yi S-I, Bae K-M, An J-S, Kim H-Y (2007) Genetic assessment of watermelon (Citrullus lanatus) varieties using SSR markers developed from Cucurbit species. Korean J Genet 29:137–146
Levi A, Thomas CE (1999) An improved procedure for isolation of high quality DNA from watermelon and melon leaves. Cucurbit Genet Coop Rep 22:41–42
Levi A, Thomas C, Wehner T, Zhang X (2001) Low genetic diversity indicates the need to broaden the genetic basis of cultivated watermelon. Hortic Sci 36:1096–1101
Loehrlein M, Ray DT (2007) Triploid and tetraploid watermelon (Citrullus lanatus (Thunb.) Matsum. and Nakai) seed size and weight. Cucurbit Genet Coop Rep 22:34–37
Loukou A, Gnakri D, Djè Y, Kippré A, Malice M, Baudoin J-P, Zoro Bi IA (2007) Macronutrient composition of three cucurbit species cultivated for seed consumption in Côte d’Ivoire. Afr J Biotechnol 6:529–533
Merrick LC (1998) Cucurbits. Crop production science in horticulture. In Austin DF, Harriman NA, Johns T, Steinberg MK, Ugarte CA, Eich E, Forstner MRJ, Collins DA, Bedigian D, Leonard K, Lizarralde M. Book reviews. Econ Bot 52:428–440
Minch E, Ruiz-Linares A, Goldstein D, Feldman M, Cavalli-Sforza LL (1995) MICROSAT Version 1.4d: a computer program for calculating various statistics on microsatellite allele data. http://crick.stanford.edu/microsat/microsat.html
Navot N, Zamir D (1987) Isozyme and seed protein phylogeny of the genus Citrullus (Cucurbitaceae). Plant Syst Evol 156:61–67
Pissard A, Arbizu C, Ghislain M, Faux A-M, Paulet S, Bertin P (2008) Congruence between morphological and molecular markers inferred from the analysis of the intra-morphotype genetic diversity and the spatial structure of Oxalis tuberosa Mol. Genetica 132:71–85
Powell W, Morgante M, Andre C, Hanafey M, Vogel J, Tingey S, Rafalski A (1996) The comparison of RFLP, RAPD, AFLP and SSR (microsatellite) markers for germplasm analysis. Mol Breed 2:225–238
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Raymond M, Rousset F (1995) Genepop (version 1.2): population genetics software for exact tests and ecumenicism. J Hered 96:248–250
Ritschel PS, de Lima Lins TS, Tristan RL, Cortopassi Buso GS, Buso JA, Ferreira ME (2004) Development of microsatellite markers from an enriched genomic library for genetic analysis of melon (Cucumis melo L.). BMC Plant Biol 4:9
Tahi GC (2006) Genetic structure of African edible seeds Citrullus lanatus (Thunberg) Matsumara & Nakai var. citroides using ISSR molecular markers. Master’s thesis, Université catholique de Louvain, Faculté d’ingénierie biologique, agronomique et environnementale
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Zoro Bi IA, Koffi K, Djè Y (2003) Caractérisation botanique et agronomique de trois espèces de cucurbites consommées en sauce en Afrique de l’ouest : Citrullus sp., Cucumeropsis mannii Naudin et Lagenaria siceraria (Molina) Standl. Biotechnology, Agronomy. Soc Environ 7:189–199
Acknowledgments
This research was supported by the FSR (Fonds Spéciaux de Recherche, UCL) and by the FRIA (Fonds pour la formation à la Recherche dans l’Industrie et dans l’Agriculture, Belgium). We are grateful to P. Baret (UCL) and X. Draye (UCL) for their advice during the proofreading of this paper.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Minsart, LA., Zoro bi, I.A., Djè, Y. et al. Set up of simple sequence repeat markers and first investigation of the genetic diversity of West-African watermelon (Citrullus lanatus ssp. vulgaris oleaginous type). Genet Resour Crop Evol 58, 805–814 (2011). https://doi.org/10.1007/s10722-010-9617-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-010-9617-x