Abstract
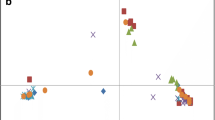
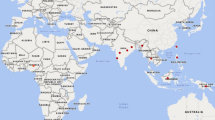
Worldwide genetic diversity in 200 individuals comprising 41 castor bean accessions was assessed using amplified fragment polymorphisms (AFLPs) and simple sequence repeats (SSRs). We found that, despite surveying five continents and 35 countries, genetic diversity in castor bean germplasm is relatively low (overall H e = 0.126 for AFLPs and 0.188 for SSRs) compared to estimates of genetic diversity in other plant species. Our data also show no geographic structuring of genotypes across continents or countries within continents. An assessment of the congruence between AFLP and SSRs indicates a low correlation (R 2 = 0.19) between the two data sets, but each marker class nonetheless shows similar patterns of low-genetic diversity and a lack of geographic structure. Our data do suggest that SSRs yield a higher percentage of polymorphic loci, higher heterozyosity and a greater range of genetic distances, and are therefore more informative than are AFLPs on a locus-by-locus basis. Based on comparisons with numerous other plant species, we suggest that the lower genetic variation in this worldwide collection may be due to one or more factors including: sampling strategies that have not captured the full extent of genetic variation in the species; artifactual variation due to long-term germplasm storage and seed regeneration; or intense selection followed by domestic cultivation of a limited number of castor bean genotypes, which are widely propagated for their horticultural and agro-economic value.



Similar content being viewed by others
References
Allard RW, Alvim P De T, Ashri A, Barton JH (1991) Managing global genetic resources; the U. S. national plant germplasm system: elements of the national plant germplasm system. The National Academies Press, Washington, DC, pp 43–86
Allender CJ, Easterday WR, Van Ert MN, Wagner DM, Keim P (2004) High-throughput extraction of arthropod vector and pathogen DNA using bead milling. Biotechniques 37:730–734
Bandelj D, Jakse J, Javornik B (2004) Assessment of genetic variability of olive varieties by microsatellite and AFLP markers. Euphytica 136:93–102
Barkley NA, Roose ML, Krueger RR, Federici CT (2006) Assessing genetic diversity and population structure in a citrus germplasm collection utilizing simple sequence repeat markers (SSRs). Theor Appl Genet 112:1519–1531
Bataillon TM, David JL, Schoen DJ (1996) Neutral genetic markers and conservation genetics: Simulated germplasm collections. Genetics 144:409–417
Botstein D, White RL, Skolnick M, Davis RW (1980) Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet 32:314–331
Brigham RD 1967 Natural outcrossing in dwarf-internode castor, Ricinus communis L. Crop Sci 7:353–355
Brown AHD (1978) Isozymes, plant population genetic structure and genetic conservation. Theor Appl Genet 52:145–157
Brown AHD (1989) Core collections: a practical approach to genetic resources management. Genome 31:818–824
Chavarriaga-Aguirre P, Maya M, Tohme J, Duque MC, Iglesias C, Bonierbale MW, Kresovich S, Kochert G (1999) Using microsatellites, isozymes and AFLPs to evaluate genetic diversity and redundancy in the cassava core collection and to assess the usefulness of DNA-based markers to maintain germplasm collections. Mol Breed 5:263–273
Colombo C, Second G, Charrier A (2000) Diversity within American cassava germplasm based on RAPD markers. Genet Mol Biol 23:189–199
Crossa J, Hernandez CM, Bretting P, Eberhart SA, Taba S (1993) Statistical genetic considerations for maintaining germplasm collections. Theor Appl Genet 86:673–678
Endo YK, Mitsui K, Motizuki M, Tsurugi K (1987) The mechanism of action of ricin and related toxic lectins on eukaryotic ribosomes: the site and the characteristics of the modification in 28 S ribosomal RNA caused by the toxins. J Biol Chem 262:5908–5917
Figueiredo-Goulart M, Ribeiro SP, Lovato MB (2005) Genetic, morphological and spatial characterization of two populations of Mabea fistulifera Mart. (Euphorbiaceae), in different successional stages. Braz Arch Biol Technol 48:275–284
Frankel OH, Brown AHD (1984) Plant genetic resources today: a critical appraisal. In: Holden JHW, Williams JT (eds) Crop genetic resources: conservation and evaluation. Allen and Unwin, London, pp 149–257
Gaudeul MI, Till-Bottraud I, Barjon F, Manel S (2004) Genetic diversity and differentiation in Eryngium alpinum L. (Apiaceae): comparison of AFLP and microsatellite markers Heredity 92:508–518
Gilbert JE, Lewis RV, Wilkinson MJ, Caligari PDS (1999) Developing an appropriate strategy to assess genetic variability in plant germplasm collections. Theor Appl Genet 98:1125–1131
Gitzendanner MA, Soltis PS (2000) Patterns of genetic variation in rare and widespread plant congeners. Am J Bot 87:783–792
Govaerts R, Frodin DG, Radcliffe-Smith A (2000) World checklist and bibliography of Euphorbiaceae (with Pandaceae). Redwood Books Limited, Trowbridge, Wiltshire
Hamrick JL (1989) Isozymes and analyses of genetic structure of plant populations. Discorides Press, Portland, OR
Harlan JR (1987) In: Yeatman CW, Kafton D, Wilkes G (eds) Plant genetic resources: a conservation imperative. Westview, Boulder, CO, pp 111–129
He G, Prakash C (2001) Evaluation of genetic relationships among botanical varieties of cultivated peanut (Arachis hypogaea L.) using AFLP markers. Genet Resour Crop Evol 48:347–352
Hilu KW (1994) Evolution of domesticated plants. Encyclopedia Agric Sci 2:117–127
Hopkins MS, Casa AM, Wang T, Mitchell SE, Dean RE, Kochert GD, Kresovich S (1999) Discovery and characterization of polymorphic simple sequence repeats (SSRs) in peanut. Crop Sci 39:1243–1247
Hyten DL, Song Q, Zhu Y, Choi I, Nelson RL, Costa JM, Specht JE, Shoemaker RC, Cregan PB (2006) Impacts of genetic bottlenecks on soybean genome diversity. PNAS 103:16666–16671
Jakse JK, Kindlhofer K, Javornik B (2001) Assessment of genetic variation and differentiation of hop genotypes by microsatellite and AFLP markers. Genome 44:773–782
Knight B (1979) Ricin – a potent homicidal poison. Br Med J 278:350–351
Kochert G, Halwart T, Branch WD (1991) RFLP variability in peanut (Arachis hypogea L.) cultivars and wild species. Theor Appl Genet 81:565–570
Lanham PG, Fennell S, Moss JP (1992) Detection of polymorphic loci in Arachis germplasm using random amplified polymorphic DNAs. Genome 35:885–889
Levi A, Thomas CE (2001) Low genetic diversity indicates the need to broaden the genetic base of cultivated watermelon. J Hortic Sci 36:1096–1101
Mariette SV, Le Corre V, Austerlitz F, Kremer A (2002) Sampling within the genome for measuring within-population diversity: trade-offs between markers. Mol Ecol 11:1145–1156
Marshall TJ, Slate J, Kruuk LEB, Pemberton JM (1998) Statistical confidence for likelihood-based paternity inference in natural populations. Mol Ecol 7:639–655
Meinders HC, Jones MD (1950) Pollen shedding and dispersal in the castor plant Ricinus communis L. J Agron 4:206–209
Mignouna HD, Abang MM, Fagbemi SA (2003) A comparative assessment of molecular marker assays (AFLP, RAPD and SSR) for white yam (Dioscorea rotundata) germplasm characterisation. Ann Appl Biol 142:269–276
Mwase WF, Bjørnstad Å, Stedje B, Bokosi JM, Kwapata MB (2006) Genetic diversity of Uapaca kirkiana Muel. Årg. populations as revealed by amplified fragment length polymorphisms (AFLPs). Afr J Biotech 5:1205–1213
Nybom H (2004) Comparison of different nuclear DNA markers for estimating intraspecific genetic diversity in plants. Mol Ecol 13:1143–1155
Oetting WS, Lee HK, Flanders DJ, Wiesner GL, Sellers TA, King RA (1995) Linkage analysis with multiplexed short tandem repeat polymorphisms using infrared fluorescence and M13 tailed primers. Genomics 30:450–458
Park K (2004) Comparisons of allozyme variation of narrow endemic and widespread species of Far East Euphorbia (Euphorbiaceae). Bot Bull Acad Sin 45:221–228
Parzies HK, Spoor W, Ennos RA (2000) Genetic diversity of barley landrace accessions (Hordeum vulgare ssp. vulgare) conserved for different lengths of time in ex situ gene banks Heredity 84:476–486
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Portis E, Barchi L, Acquadro A, Macua JI, Lanteri S (2005) Genetic diversity assessment in cultivated cardoon by AFLP (amplified fragment length polymorphism) and microsatellite markers. Plant Breed 124:299–304
Powell W, Morgante M, Andre C, Hanafey M, Vogel J, Tingey S, Rafalski A (1996) The comparison of RFLP, RAPD, AFLP and SSR (microsatellite) markers for germplasm analysis. Mol Breed 2:225–238
Richard M, Thorpe RS (2001) Can microsatellites be used to infer phylogenies? Evidence from population affinities of the Western Canary Island lizard (Gallotia galloti). Mol Phylogenet Evol 3:351–360
Russell JR, Fuller JD, Macaulay M, Hatz BG, Jahoor A, Powell W, Waugh R (1997) Direct comparison of levels of genetic variation among barley accessions detected by RFLPs, AFLPs, SSRs and RAPDs. Theor Appl Genet 95:714–722
Saitou N, Nei M (1987) The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sethy NK, Shokeen B, Edwards KJ, Bhatia S (2006) Development of microsatellite markers and analysis of intraspecific genetic variability in chickpea (Cicer L.). Theor Appl Genet 112:1416–1428
Tautz D (1989) Hypervariability of simple sequences as a general source for polymorphic DNA markers. Nucleic Acids Res 17:6463–6471
Vekemans XT, Beauwens T, Lemaire M, Roldan-Ruiz I (2002) Data from amplified fragment length polymorphism (AFLP) markers show indication of size homoplasy and of a relationship between degree of homoplasy and fragment size. Mol Ecol 11:139–151
Vos PR, Hogers R, Bleeker M, Reijans M, Van De Lee T, Hornes M, Frijters A, Pot J, Peleman J, Kuiper M, Zabeau M (1995) AFLP: a new technique for DNA fingerprinting. Nucleic Acids Res 23:4407–4414
Weber E (2003) The most complete global overview of invasive species in natural areas. Divers Distrib 10:505
Wolfe AD, Liston A (1998) Contributions of the polymerase chain reaction to plant systematics and evolutionary biology. In: Soltis DE, Soltis PS, Doyle JJ (eds) Molecular systematics of plants II. Kluwer, Dordrecht, pp 43–86
Wurdack KP, Hoffman P, Chase MW (2005) Molecular phylogenetic analysis of uniovulate Euphorbiaceae (Euphorbiaceae sensu stricto) using plastid RbcL and TrnL-F DNA sequences. Am J Bot 92:1397–1420
Yeh FC, Yang R-C, Boyle TBJ, Ye Z-H, Mao JX (1997) POPGENE, the user-friendly shareware for population genetic analysis. Molecular Biology and Biotechnology Centre, University of Alberta. AB, Canada
Zhou Y, Bui T, Auckland LD, Williams CG (2002) Direct fluorescent primers are superior to M13–tailed primers for Pinus taeda microsatellites. Biotechniques 32:46–52
Zhu Q, Zheng X, Luo J, Gaut BS, Ge S (2007) Multilocus Analysis of Nucleotide Variation of Oryza sativa and Its Wild Relatives: severe Bottleneck during Domestication of Rice. Mol Biol Evol 24:875–888
Acknowledgements
We extend our thanks to Dr. Brad Morris at the USDA-ARS Plant Genetic Resource Conservation Unit in Griffin, GA, USA for providing seeds of castor bean accessions discussion in choosing germplasm material. Bradford Blake (NAU) provided expert guidance and care of castor bean seedlings. We also thank Drs. Jim Robertson and Mark Wilson (Federal Bureau of Investigation, Quantico Laboratories, Quantico, Virginia) for providing funding for this project and assistance with sampling of castor bean accessions. This research was also supported in part by Federal funds from the National Institute of Allergy and Infectious Diseases, NIH, DHHS, under contract No. NO1-AI-30071. Dr. Ken Wurdack (Smithsonian Institution) provided helpful information about the systematic and phylogenetic affinities of castor bean. The views expressed in this academic research paper are those of the author(s) and do not reflect the official policy or position of the US Government or the Department of Justice.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Allan, G., Williams, A., Rabinowicz, P.D. et al. Worldwide genotyping of castor bean germplasm (Ricinus communis L.) using AFLPs and SSRs. Genet Resour Crop Evol 55, 365–378 (2008). https://doi.org/10.1007/s10722-007-9244-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-007-9244-3