Abstract
Phosphoglucan phosphatases (Like-SEX4 1 and 2; LSF1 and LSF2) were reported to play roles in starch metabolism in leaves of Arabidopsis. In this study, we identified and mapped the LSF1 and LSF2 genes in barley (HvLSF1 and HvLSF2), characterized their gene and protein structures, predicted the cis-elements of their promoters, and analysed their expression patterns. HvLSF1 and HvLSF2 were mapped on the long arm of chromosome 1H (1HL) and 5H (5HL), respectively. Our results revealed varied exon–intron structures and conserved exon–intron junctions in both LSF1 and LSF2 from a range of analysed species. Alignment of protein sequences indicated that cTP and CT domains are much less varied than the functional domains (PDZ, DPS and CBM48). LSF2 was mainly expressed in anthers of barley and rice, and in leaf of Arabidopsis. LSF1 was mainly expressed in endosperm of barley and leaf of Arabidopsis and rice. The expression of LSF1 exhibited a diurnal pattern in rice only and that of LSF2 in both rice and Arabidopsis. Of the investigated stresses, only cold stress significantly reduced expression level of LSF1 and LSF2 in barley and LSF2 in Arabidopsis at late stages of the treatments. While heat treatment significantly decreased expression levels of LSF1 at middle stage (4 h) of a treatment in Arabidopsis only. The strong relationships detected between LSF2 and starch excess4 (SEX4), glucan, water dikinases or phosphoglucan, water dikinases were identified and discussed. Taken together, these results provide information of genetic manipulation of LSF1 and LSF2, especially in monocotyledon and further elucidate their regulatory mechanism in plant development.






Similar content being viewed by others
Abbreviations
- aa:
-
Acid residues
- ABA:
-
Abscisic acid
- CBM48:
-
Carbohydrate-binding module 48
- CDS:
-
Coding sequences
- CT:
-
C-terminal
- cTP:
-
Chloroplast transit peptide
- DAP:
-
Days after pollination
- DSP:
-
Dual specificity phosphatase
- GWD:
-
Glucan, water phosphotransferase
- NCBI:
-
National Center for Biotechnology Information
- ORF:
-
Open reading frame
- PWD:
-
Phosphoglucan, water dikinase
- LSF1:
-
Like-SEX4 1
- LSF2:
-
Like-SEX4 2
- SEX4:
-
Phosphoglucan phosphatase starch excess4
References
Bahaji A et al (2013) Starch biosynthesis, its regulation and biotechnological approaches to improve crop yields. Biotechnol Adv 32:87–106
Bauerlein M, Frohberg C (2003) Method for generating maize plants with an increased leaf starch content, and their use for making maize silage. US Patent application no.US 2006/0150278 A1
Comparot-Moss S et al (2010) A putative phosphatase, LSF1, is required for normal starch turnover in Arabidopsis leaves. Plant Physiol 152:685–697. doi:10.1104/pp.109.148981
Emanuelsson O, Brunak S, von Heijne G, Nielsen H (2007) Locating proteins in the cell using TargetP, SignalP and related tools. Nat Protoc 2:953–971
Fordham-Skelton AP et al (2002) A novel higher plant protein tyrosine phosphatase interacts with SNF1-related protein kinases via a KIS (kinase interaction sequence) domain. Plant J: Cell Mol Biol 29:705–715
Frohberg C, Leube M, Rumpler I (2003) Transgenic fodder plants with an increased leaf starch content. US Patent application no. US 2003/0217386 A1
Hejazi M, Fettke J, Paris O, Steup M (2009) The two plastidial starch-related dikinases sequentially phosphorylate glucosyl residues at the surface of both the A-and B-type allomorphs of crystallized maltodextrins but the mode of action differs. Plant Physiol 150:962–976
Hejazi M, Fettke J, Kötting O, Zeeman SC, Steup M (2010) The laforin-like dual-specificity phosphatase SEX4 from Arabidopsis hydrolyzes both C6-and C3-phosphate esters introduced by starch-related dikinases and thereby affects phase transition of α-glucans. Plant Physiol 152:711–722
Higo K, Ugawa Y, Iwamoto M, Korenaga T (1999) Plant cis-acting regulatory DNA elements (PLACE) database. Nucleic Acids Res 27:297–300
Hirose T et al (2013) Disruption of a rice gene for α-glucan water dikinase, OsGWD1, leads to hyperaccumulation of starch in leaves but exhibits limited effects on growth. Front Plant Sci 4
International-Barley-Genome-Sequencing-Consortium (2012) A physical, genetic and functional sequence assembly of the barley genome. Nature 491:711–716
International-Wheat-Genome-Sequencing-Consortium (2014) A chromosome-based draft sequence of the hexaploid bread wheat (Triticum aestivum) genome. Science 345:1251788
Keren H, Lev-Maor G, Ast G (2010) Alternative splicing and evolution: diversification, exon definition and function. Nat Rev Genet 11:345–355
Kötting O et al (2009) STARCH-EXCESS4 is a laforin-like phosphoglucan phosphatase required for starch degradation in Arabidopsis thaliana. Plant Cell 21:334–346
Ma J et al (2013) Structure and expression of barley starch phosphorylase genes. Planta 238:1081–1093
Ma J et al (2014) Conserved structure and varied expression reveal key roles of phosphoglucan phosphatase gene starch excess 4 in barley. Planta 240:1179–1190. doi:10.1007/s00425-014-2140-0
Mahlow S, Hejazi M, Kuhnert F, Garz A, Brust H, Baumann O, Fettke J (2014) Phosphorylation of transitory starch by α-glucan, water dikinase during starch turnover affects the surface properties and morphology of starch granules. New Phytol 203:495–507
Meekins DA et al (2013) Structure of the Arabidopsis glucan phosphatase like sex four2 reveals a unique mechanism for starch dephosphorylation. Plant Cell 25:2302–2314. doi:10.1105/tpc.113.112706
Meekins DA et al (2014) Phosphoglucan-bound structure of starch phosphatase starch Excess4 reveals the mechanism for C6 specificity. Proc Natl Acad Sci USA. 10.1073/pnas.1400757111
Nashilevitz S, Melamed-Bessudo C, Aharoni A, Kossmann J, Wolf S, Levy AA (2009) The legwd mutant uncovers the role of starch phosphorylation in pollen development and germination in tomato. Plant J 57:1–13
Ral JP et al (2012) Down-regulation of glucan, water-dikinase activity in wheat endosperm increases vegetative biomass and yield. Plant Biotechnol J 10:871–882
Reddy AS (2007) Alternative splicing of pre-messenger RNAs in plants in the genomic era. Annu Rev Plant Biol 58:267–294
Regina A et al (2012) Differential effects of genetically distinct mechanisms of elevating amylose on barley starch characteristics. Carbohydr Polym 89:979–991. doi:10.1016/j.carbpol.2012.04.054
Santelia D et al (2011) The phosphoglucan phosphatase like sex Four2 dephosphorylates starch at the C3-position in Arabidopsis. Plant Cell 23:4096–4111. doi:10.1105/tpc.111.092155
Silver DM, Kötting O, Moorhead GB (2014) Phosphoglucan phosphatase function sheds light on starch degradation. Trends Plant Sci. doi:10.1016/j.tplants.2014.01.008
Skeffington AW, Graf A, Duxbury Z, Gruissem W, Smith AM (2014) Glucan, water dikinase exerts little control over starch degradation in Arabidopsis leaves at night. Plant Physiol 165:866–879
Somers D, Bilgic H, Cho S, Garvin DF, Muehlbauer GJ (2007) Mapping barley genes to chromosome arms by transcript profiling of wheat–barley ditelosomic chromosome addition lines. Genome 50:898–906
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Van Hung P, Maeda T, Morita N (2006) Waxy and high-amylose wheat starches and flours—characteristics, functionality and application. Trends Food Sci Technol 17:448–456
Vriet C, Smith AM, Wang TL (2014) Root starch reserves are necessary for vigorous re-growth following cutting back in Lotus japonicus. PloS One 9:e87333
Zeeman SC, Kossmann J, Smith AM (2010) Starch: its metabolism, evolution, and biotechnological modification in plants. Annu Rev Plant Biol 61:209–234
Acknowledgments
This work was supported by the International Science and Cooperation Program of China (No. 2015DFA30600), the Applied Basic Research Programs of Science and Technology Department of Sichuan Province (2016JY0010) and the Key Projects of Education Department of Sichuan Province (16ZA0038). We appreciate the anonymous referees for critical reading of the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Jian Ma and Shang Gao have contributed equally to this paper.
Electronic supplementary material
Below is the link to the electronic supplementary material.
10709_2016_9900_MOESM1_ESM.tif
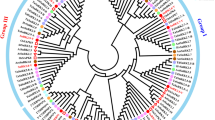
Fig. S1. Phylogenetic relationship from a range of species based on LSF1 and LSF2 protein sequences using maximum likelihood method (see “Materials and methods” for accession numbers) (TIFF 778 kb)
Rights and permissions
About this article
Cite this article
Ma, J., Gao, S., Jiang, QT. et al. Structure and expression of phosphoglucan phosphatase genes of Like Sex Four1 and Like Sex Four2 in barley. Genetica 144, 313–323 (2016). https://doi.org/10.1007/s10709-016-9900-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10709-016-9900-7