Abstract
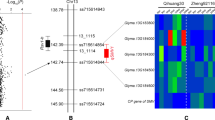
Yellow Mosaic Virus (YMV) is a serious disease of soybean. Resistance to YMV was mapped in 180 soybean genotypes through association mapping approach using 121 simple sequence repeats (SSR) and four resistance gene analogue (RGA)-based markers. The association mapping population (AMP) (96 genotypes) and confirmation population (CP) (84 genotypes) was tested for resistance to YMV at hot-spot consecutively for 3 years (2007–2009). The genotypes exhibited significant variability for YMV resistance (P < 0.01). Molecular genotyping and population structure analysis with ‘admixture’ co-ancestry model detected seven optimal sub-populations in the AMP. Linkage disequilibrium (LD) between the markers extended up to 35 and 10 cM with r2 > 0.15, and >0.25, respectively. The 4 RGA-based markers showed no association with YMV resistance. Two SSR markers, Satt301 and GMHSP179 on chromosome 17 were found to be in significant LD with YMV resistance. Contingency Chi-square test confirmed the association (P < 0.01) and the utility of the markers was validated in the CP. It would pave the way for marker assisted selection for YMV resistance in soybean. This is the first report of its kind in soybean.



Similar content being viewed by others
References
Abdurakhmonov IY, Abdukarimov A (2008) Application of Association mapping to Understanding the Genetic Diversity of Plant Germplasm Resources. Int J Plant Genomics Article ID 574927, 18 pages doi:10.1155/2008/574927
Agrama HA, Eizenga GC (2008) Molecular diversity and genome-wide linkage disequilibrium patterns in a worldwide collection of Oryza sativa and its wild relatives. Euphytica 160:339–355
Basak J, Kundagrami S, Ghose TK, Pal A (2004) Development of yellow mosaic virus (YMV) resistance linked DNA marker in Vigna mungo from populations segregating for YMV-reaction. Mol Breed 14:375–383
Bhattacharyya PK, Ram HH, Kole PC (1999) Inheritance of resistance to yellow mosaic virus in inter specific crosses of soybean. Euphytica 108:157–159
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635
Charlson DV, Cianzio SR, Shoemaker RC (2003) Associating SSR markers with soybean resistance to iron deficiency chlorosis. J Plant Nutr 26:2267–2276
Charlson DV, Bailey TB, Cianzio SR, Shoemaker RC (2005) Molecular marker Satt481 is associated with iron-deficiency chlorosis resistance in a soybean breeding population. Crop Sci 45:2394–2399
Collard BCY, Jahufer MZZ, Brouwer JB, Pang ECK (2005) An introduction to markers, quantitative trait loci (QTL) mapping and marker-assisted selection for crop improvement: the basic concepts. Euphytica 142:169–196
Cregan PB, Jarvik T, Bush AL, Shoemaker RC, Lark KG, Kahler AL, Kaya N, VanToai TT, Lohnes DG, Chung J, Specht JE (1999) An integrated genetic linkage map of the soybean genome. Crop Sci 39:1464–1490
Doerge RW (2002) Mapping and analysis of quantitative trait loci in experimental populations. Nat Rev Genet 3:43–52
Evanno GS, Regnaut Goudet J (2005) Detecting the number of clusters of individuals using the software structure: a simulation study. Mol Ecol 14:2611–2620
Farnir F, Coppieters W, Arranz JJ, Berzi P, Cambisano N, Grisart B, Karim L, Marcq F, Moreau L, Mni M, Nezer C, Simon P, Vanmanshoven P, Wagenaar D, Michel G (2000) Extensive genome wide linkage disequilibrium in cattle. Genome Res 10:220–227
Flint-Garcia SA, Thornsberry JM, Buckler ESIV (2003) Structure of linkage disequilibrium in plants. Annu Rev Plant Biol 54:357–374
Garris AJ, McCouch SR, Kresovich S (2003) Population structure and its effects on haplotype diversity and linkage disequilibrium surrounding the xa5 locus of rice Oryza sativa L. Genetics 165:759–769
Gouis JL, Bordes J, Ravel C, Heumez E, Faure S, Praud S, Galic N, Remoue C, Balfourier F, Allard V, Rousset M (2012) Genome-wide association analysis to identify chromosomal regions determining components of earliness in wheat. Theor Appl Genet 124:597–611
Hardy OJ, Vekemans X (2002) SPAGeDi: a versatile computer program to analyse spatial genetic structure at the individual or population level. Mol Ecol Notes 2:618–620
Hill WG, Robertson A (1968) Linkage disequilibrium in finite populations. Theor Appl Genet 38:226–231
Holland JB (2007) Genetic architecture of complex traits in plants. Curr Opin Plant Biol 10:156–161
Jun TH, Van K, Kim MY, Lee SH, Walker DR (2008) Association analysis using SSR markers to find QTL for seed protein content in soybean. Euphytica 162:179–191
Kraakman ATW, Niks RE, Van den Berg PMMM, Stam P, Van Eeuwijk FA (2004) Linkage disequilibrium mapping of yield and yield stability in modern spring barley cultivars. Genetics 168:435–446
Lal SK, Rana VKS, Sapra RL, Singh KP (2005) Screening and utilization of soybean germplasm for breeding resistance against Mungbean yellow mosaic virus. Soyb Genet Newsl 32. http://soybase.org:8083/articleFiles/45
Lewontin RC (1964) The interaction of selection and linkage. I. General considerations; heterotic models. Genetics 49:49–67
Malysheva-Otto LV, Ganal MW, Roder MS (2006) Analysis of molecular diversity, population structure and linkage disequilibrium in a worldwide survey of cultivated barley germplasm (Hordeum vulgare L.). BMC Genet 7:6
Maskri YA, Sajjad M, Khan SH (2012) Association mapping: a step forward to discovering new alleles for crop improvement. Int J Agr Biol 14:153–160
Meuwissen THE, Goddard ME (2000) Fine mapping of quantitative trait loci using linkage disequilibria with closely linked marker loci. Genetics 155:421–430
Nene YL (1972) A surveys of the viral diseases of pulse crops in India, G B Pant Univ Agric Tech, Pantnagar. India J Res Bull 4:191
Nordborg M, Tavare S (2002) Linkage disequilibrium: what history has to tell us? Trends Genet 18:83–90
Palaisa K, Morgante M, Williams M, Rafalski A (2004) Longrange patterns of diversity and linkage disequilibrium surrounding the maize Y1 gene are indicative of an asymmetric selective sweep. Proc Natl Acad Sci USA 101:9885–9890
Price AH (2006) Believe it or not, QTLs are accurate! Trends Plant Sci 11:213–216
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Saghai-Maroof MA, Soliman KM, Jorgensen RA, Allard RW (1984) Ribosomal DNA spacer-length polymorphisms in barley: mendelian inheritance, chromosomal location and population dynamics. Proc Natl Acad Sci USA 81:8014–8018
Singh BB, Mallick AS (1978) Inheritance of resistance to yellow mosaic in soybean. Indian J Genet 38:258–261
Sun G, Zhu C, Kramer MH, Yang SS, Song W, Piepho HP, Yu J (2010) Variation explained in mixed–model association mapping. Heredity 105:333–340
Szalma SJ, Buckler IVES, Snook ME, McMullen MD (2005) Association analysis of candidate genes for maysin and chlorogenic acid accumulation in maize silks. Theor Appl Genet 110:1324–1333
Talukdar A, Harish GD, Shivakumar M, Kumar B, Verma K, Lal SK, Sapra RL, Singh KP (2013) Inheritance of yellow mosaic virus resistance in cultivated soybean (Glycine max L. Merr.). Legume Res 36(3):263–267
Thavamanikumar S, Tibbits J, McManus L, Ades P, Stackpole D, Hadjigol S, Vaillancourt S, Zhu P, Bossinger G (2011) Candidate gene-based association mapping of growth and wood quality traits in Eucalyptus globulus Labill. BMC Proc 5(Suppl 7):O15
Thornsberry JM, Goodman MM, Doebley J, Kresovich S, Nielsen D, Buckler ESIV (2001) Dwarf8 polymorphisms associate with variation in flowering time. Nat Genet 28:286–289
Wang J, Mc Clean PE, Lee R, Goos RJ, Helms T (2008) Association mapping of iron deficiency chlorosis loci in soybean (Glycine max L. Merr.) advanced breeding lines. Theor Appl Genet 116:777–787
Yu J, Pressoir G, Briggs WH, Bi IV, Yamasaki M, Doebley JF, McMullen MD, Gaut BS, Nielsen DM, Holland JB, Kresovich S, Buckler ES (2006) A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat Genet 38:203–208
Zhao K, Tung CW, Eizenga GC, Wright MH, Ali ML, Price AH, Mezey J, McClung AM, Bustamante CD, McCouch SR (2011) Genome wide association mapping reveals a rich genetic architecture of complex traits in Oryza sativa. Nat Commun 2(467):1467. doi:10.1038/ncomms
Zhu C, Gore M, Buckler ES, Yu J (2008) Status and prospects of association mapping in plants. Plant Genome 1:5–20
Acknowledgments
First author sincerely acknowledges Post Graduate School, Indian Agricultural Research Institute, for providing the fellowship during the Post-Graduate study.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kumar, B., Talukdar, A., Verma, K. et al. Mapping of yellow mosaic virus (YMV) resistance in soybean (Glycine max L. Merr.) through association mapping approach. Genetica 143, 1–10 (2015). https://doi.org/10.1007/s10709-014-9801-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10709-014-9801-6



