Abstract
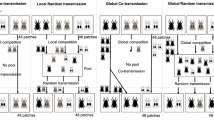
The high prevalence of meiotic recombination—an important element of sexual reproduction—represents one of the greatest puzzles in biology. The influence of either selection by a co-evolving parasite alone or in combination with genetic drift on recombination rates was tested in the host-parasite system Tribolium castaneum and Nosema whitei. After eight generations, populations with smaller genetic drift had a lower recombination rate than those with high drift whereas parasites had no effect. Interestingly, changes in recombination rate at one site of the chromosome negatively correlated with changes at the adjacent site on the same chromosome indicating an occurrence of crossover interference. The occurrence of spontaneous or plastic changes in recombination rates could be excluded with a separate experiment.



Similar content being viewed by others
References
Abdullah MFF, Borts RH (2001) Meiotic recombination frequencies are affected by nutritional states in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A 98:14524–14529
Armstrong E, Bass LK (1986) Effects of infection by Nosema whitei on the mating frequency and fecundity of Tribolium castaneum. J Invertebr Pathol 47:310–316
Barton N (1995) A general model for the evolution of recombination. Genet Res 65:123–144
Barton NH, Otto SP (2005) Evolution of recombination due to random drift. Genetics 169:2353–2370
Brooks LD (1988) The evolution of recombination rates. In: Michod RE, Levin BR (eds) The evolution of sex. Sinauer Associates, Sunderland, Mass, pp 87–105
Burt A (2000) Perspective: sex, recombination, and the efficacy of selection—was Weismann right? Evolution 54:337–351
Dewees AA (1975) Genetic modification of recombination rate in Tribolium castaneum. Genetics 81:537–552
Fischer O, Schmid-Hempel P (2005) Selection by parasites may increase host recombination frequency. Biology Lett 1:193–195
Grell RF (1978) A comparison of heat and interchromosomal effects on recombination and interference in Drosophila melanogaster. Genetics 89:65–77
Hasu T, Valtonen ET, Jokela J (2006) Costs of parasite resistance for female survival and parental care in a freshwater isopod. Oikos 114:322–328
Hillers KJ (2004) Crossover interference. Curr Biol 14:R1036–R1037
Korol AB (1999) Selection for adaptive traits as a factor of recombination evolution: evidence from natural and experimental populations (a review). In: Wasser SP (ed) Evolutionary theory and processes: modern perspectives. Kluwer Academic Publishers, Amsterdam, pp 31–53
Kovalchuk I, Kovalchuk O, Kalck V, Boyko V, Filkowski J, Heinlein M, Hohn B (2003) Pathogen-induced systemic plant signal triggers DNA rearrangements. Nature 423:760–762
Lebel EG, Masson J, Bogucki A, Paszkowski J (1993) Stress-induced intrachromosomal recombination in plant somatic-cells. Proc Natl Acad Sci U S A 90:422–426
Lucht JM, Mauch-Mani B, Steiner H-Y, Metraux J-P, Ryals J, Hohn B (2002) Pathogen stress increases somatic recombination frequency in Arabidposis. Nat Genet 30:311–313
Milner RJ (1972) Nosema whitei, a microsporidian pathogen of some species of Tribolium. I. Morphology, life cycle, and generation time. J Invertebr Pathol 19:231–238
Moret Y, Schmid-Hempel P (2000) Survival for immunity: the price of immune system activation for bumblebee workers. Science 290:1166–1168
Neel JV (1941) A relation between larval nutrition and the frequency of crossing over in the third chromosome of Drosophila melanogaster. Genetics 26:506–516
Otto SP, Barton NH (2001) Selection for recombination in small populations. Evolution 55:1921–1931
Peters AD, Lively CM (1999) The Red Queen and fluctuating epistasis: a population genetic analysis of antagonistic coevolution. Am Nat 154:393–405
Plough HH (1917) The effect of temperature on crossingover in Drosophila. J Exp Zool 24:187–202
Plough HH (1921) Further studies on the effect of temperature on crossing over. J Exp Zool 32:187–202
Puchta H, Swoboda P, Hohn B (1995) Induction of intrachromosomal homologous recombination in whole plants. Plant J 7:203–210
Rice WR (2002) Experimental tests of the adaptive significance of sexual reproduction. Nat Rev Genet 3:241–251
Salathé M, Kouyos RD, Bonhoeffer S (2008) The state of affairs in the kingdom of the Red Queen. Trends Ecol Evol 23:439–445
Schmid-Hempel P (2003) Variation in immune defence as a question of evolutionary ecology. Proc R Soc Lond B 270:357–366
Van Veen JE, Hawley RS (2003) Meiosis; when even two is a crowd. Curr Biol 13:R831–R833
Weinstein A (1936) The theory of multiple-strand crossover. Genetics 21:155–199
Zhao HY, Speed TP, Mcpeek MS (1995) Statistical analysis of crossover interference using the Chi-square model. Genetics 139:1045–1056
Acknowledgments
Beetle stocks were kindly provided by R. W. Beeman (USDA), R. Schröder (University of Tübingen), J. Trauner (University of Erlangen), and G. Bucher (University of Göttingen). We thank D. Trujillo-Villegas for his help on counting the beetles and R. Schmid-Hempel for general help. The Swiss National Science Foundation (grant nr. 3100-066733 to PSH), and the Genetic Diversity Centre of ETH supported this study.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Greeff, M., Schmid-Hempel, P. Influence of co-evolution with a parasite, Nosema whitei, and population size on recombination rates and fitness in the red flour beetle, Tribolium castaneum . Genetica 138, 737–744 (2010). https://doi.org/10.1007/s10709-010-9454-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10709-010-9454-z