Abstract
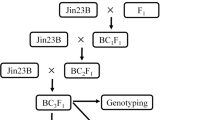
Blast (caused by Magnaporthe oryzae B. Couch) is an economically important and mutable disease of rice (Oryza sativa L.). To analyze the genetic mechanism of blast resistance in rice cultivar CR185; we used CR185 as a donor parent to establish a BC3F1 and derived BC3F2 backcross population in the Zhenshan 97B (ZS97B) background. By challenging the population with natural infection in 2008 and 2009, 11 blast resistance quantitative trait loci (QTLs) were identified. Among them, four major QTLs, qBR-1a, qBR-4d, qBR-6a, and qBR-12a, had large effects during four observation times. QTL effect analyses suggested that resistance increased with the number of QTLs. The major QTL, qBR-6a, which had the largest effect, was fine-mapped to a 100-kb DNA fragment that encompassed the Pi2/Pi9 locus. Sequence comparisons revealed that the Pi allele in CR185 was different from the known alleles of Pi2/Pi9. Our results revealed that accumulation of several QTLs is an effective way to develop highly resistant cultivars.




Similar content being viewed by others
References
Babujee L, Gnanamanickam S (2000) Molecular tools for characterization of rice blast pathogen, Magnaporthe grisea, population and molecular marker-assisted breeding for disease resistance. Curr Sci 78:24–257
Bonman JM (1992) Durable resistance to rice blast disease-environmental influences. Euphytica 63:115–123
Bonman JM, Vergel D, Khin MM (1986) Physiologic specialization of Pyricularia oryzae in the Philippines. Plant Dis 70:767–769
Bryan GT, Wu KS, Farrall L, Jia YL, Hershey HP, McAdams SA, Faulk KN, Donaldson GK, Tarchini R, Valent B (2000) A single amino acid difference distinguishes resistant and susceptible alleles of the rice blast resistance gene Pi-ta. Plant Cell 12:2033–2045
Chen HL, Wang SP, Xing YZ, Xu CG, Hayes PM, Zhang QF (2003) Comparative analyses of genomic locations and race specificities of loci for quantitative resistance to Pyricularia grisea in rice and barley. Proc Natl Acad Sci USA 100:2544–2549
Chen XW, Shang JJ, Chen DX, Lei CL, Zou Y, Zhai WX, Liu GZ, Xu JC, Ling ZZ, Cao G, Ma BT, Wang YP, Zhao XF, Li SG, Zhu LH (2006) A B-lectin receptor kinase gene conferring rice blast resistance. Plant J 46:794–804
Das A, Soubam D, Singh PK, Thakur S, Singh NK, Sharma TR (2012) A novel blast resistance gene, Pi54rh cloned from wild species of rice, Oryza rhizomatis confers broad spectrum resistance to Magnaporthe oryzae. Funct Integr Genomics 12:215–228
Deng YW, Zhu XD, Xu J, Chen HQ, He ZH (2009) Map-based cloning and breeding application of a broad-spectrum resistance gene Pigm to rice. In: Wang GL, Valent B (eds) Advances in genetics, genomics and control of rice blast disease. Springer, Dordrecht, pp 161–171
Fukuoka S, Saka N, Koga H, Ono K, Shimizu T, Ebana K, Hayashi N, Takahashi A, Hirochika H, Okuno K, Yano M (2009) Loss of function of a proline-containing protein confers durable disease resistance in rice. Science 325:998–1001
Fukuoka S, Mizobuchi R, Saka N, Ivan S, Matsumoto T, Okuno K, Yano M (2012) A multiple gene complex on rice chromosome 4 is involved in durable resistance to rice blast. Theor Appl Genet 125:551–559
Greenberg JT, Yao N (2004) The role and regulation of programmed cell death in plant–pathogen interactions. Cell Microbiol 6:201–211
Hayashi K, Yoshida H (2009) Refunctionalization of the ancient rice blast disease resistance gene Pit by the recruitment of a retrotransposon as a promoter. Plant J 57:413–425
Hayashi N, Inoue H, Kato T, Funao T, Shirota M, Shimizu T, Kanamori H, Yamane H, Hayano-Saito Y, Matsumoto T, Yano M, Takatsuji H (2010) Durable panicle blast-resistance gene Pb1 encodes an atypical CC-NBS-LRR protein and was generated by acquiring a promoter through local genome duplication. Plant J 64:498–510
He YY, Li J, Li C, Kiyosawa S, Higashi T, Horisue N (1989) Gene analysis of blast resistance of Yunnan upland rice variety, Haonaihuan. Oryza 26:173–182
Hittalmani S, Parco A, Mew TV, Zeigler RS, Huang N (2000) Fine mapping and DNA marker-assisted pyramiding of the three major genes for blast resistance in rice. Theor Appl Genet 100:1121–1128
Huang HM, Huang L, Feng GP, Wang SH, Wang Y, Liu JL, Jiang N, Yan WT, Xu LC, Sun PY, Li ZQ, Pan SJ, Liu XL, Xiao YH, Liu EM, Dai LY, Wang GL (2010) Molecular mapping of the new blast resistance genes Pi47 and Pi48 in the durably resistant local rice cultivar Xiangzi 3150. Phytopathology 101:620–626
Ishihara T, Hayano-Saito Y, Oide S, Ebana K, La NT, Hayashi K, Ashizawa T, Suzuki F, Koizumi S (2014) Quantitative trait locus analysis of resistance to panicle blast in the rice cultivar Miyazakimochi. Rice 7(1):2. doi:10.1186/s12284-014-0002-9 eCollection 2014
Jeung JU, Kim BR, Cho YC, Han SS, Moon HP, Lee YT, Jena KK (2007) A novel gene, Pi40(t), linked to DNA markers derived from NBS-LRR motifs confers broad spectrum of blast resistance in rice. Theor Appl Genet 115:1163–1177
Jiang HC, Feng YT, Bao L, Li X, Gao GJ, Zhang QL, Xiao JH, Xu CG, He YQ (2012) Improving blast resistance of Jin 23B and its hybrid rice by marker-assisted gene pyramiding. Mol Breed 30:1679–1688
Kiyosawa S (1982) Genetics and epidemiological modelling of breakdown of plant disease resistance. Annu Rev Phytopathol 20:93–117
Li YB, Wu CJ, Jiang GH, Wang LQ, He YQ (2007) Dynamic analyses of rice blast resistance for the assessment of genetic and environmental effects. Plant Breed 126:541–547
Li W, Lei CL, Cheng ZJ, Jia YL, Huang DY, Wang JL, Wang JK, Zhang X, Su N, Guo XP, Zhai HQ, Wan JM (2008a) Identification of SSR markers for a broad-spectrum blast resistance gene Pi20(t) for marker-assisted breeding. Mol Breed 22:141–149
Li YB, Wu CJ, Xing YZ, Chen HL, He YQ (2008b) Dynamic QTL analysis for rice blast resistance under natural infection conditions. Aust J Crop Sci 2:73–82
Liu RH, Meng JL (2003) MapDraw: a Microsoft Excel macro for drawing genetic linkage maps based on given genetic linkage data. Hereditas (Beijing) 25:317–321
Liu SP, Li X, Wang CY, Li XL, He YQ (2003) Improvement of resistance to rice blast in Zhenshan 97 by molecular marker-aided selection. Acta Bot Sin 45:1346–1350
Liu B, Zhang SH, Zhu XY, Yang QY, Wu SZ, Mei MT, Mauleon R, Leach J, Mew T, Leung H (2004) Candidate defense genes as predictors of quantitative blast resistance in rice. Mol Plant-Microbe Interact 17:1146–1152
Liu XQ, Yang QZ, Lin F, Hua LX, Wang CT, Wang L, Pan QH (2007) Identification and fine mapping of Pi39(t), a major gene conferring the broad-spectrum resistance to Magnaporthe oryzae. Mol Genet Genomics 278:403–410
Liu JL, Wang XJ, Mitchell T, Hu YJ, Liu XL, Dai LY, Wang GL (2010) Recent progress and understanding of the molecular mechanisms of the rice and Magnaporthe oryzae interaction. Mol Plant Pathol 11:419–427
McClung AM, Marchetti MA, Webb BD, Bollich CN (1997) Registration of “Jefferson” rice. Crop Sci 37:629–630
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucl Acids Res 8:4321–4326
Nguyen TTT, Koizumi S, La TN, Zenbayashi KS, Ashizawa T, Yasuda N, Imazaki I, Miyasaka A (2006) Pi35(t), a new gene conferring partial resistance to leaf blast in the rice cultivar Hokkai 188. Theor Appl Genet 113:697–704
Qu SH, Liu GF, Zhou B, Bellizzi M, Zeng LR, Dai LY, Han B, Wang GL (2006) The broad-spectrum blast resistance gene Pi9 encodes a nucleotide-binding site-leucine-rich repeat protein and is a member of a multigene family in rice. Genetics 172:1901–1914
Stam P (1993) Construction of integrated genetic linkage maps by means of a new computer package: JOINMAP. Plant J 3:739–744
Tabien E, Li Z, Patterson AH, Marchetti A, Stansel W, Pinson M (2002) Mapping QTLs for field resistance to the rice blast pathogen and evaluating their individual and combined utility in improved varieties. Theor Appl Genet 105:313–324
Tanksley SD, Nelson JC (1996) Advanced backcross QTL analysis: a method for the simultaneous discovery and transfer of valuable QTLs from unadapted germplasm into elite breeding lines. Theor Appl Genet 92:191–203
Thomson MJ, Tai TH, McClung AM, Lai XH, Hinga ME, Lobos KB, Xu Y, Martinez CP, McCouch SR (2003) Mapping quantitative trait loci for yield, yield components and morphological traits in an advanced backcross population between Oryza rufipogon and the Oryza sativa cultivar Jefferson. Theor Appl Genet 107:479–493
Wang GL, Mackill DJ, Bonman JM, McCouch SR, Champoux MC, Nelson RJ (1994) RFLP mapping of genes conferring complete and partial resistance to blast in a durably resistant rice cultivar. Genetics 136:1421–1434
Wang S, Basten CJ, Zeng ZB (2007) Windows QTL cartographer 2.5. Department of Statistics, North Carolina State University, Raleigh
Wang Y, Wang D, Deng XJ, Liu JL, Sun PY, Liu Y, Huang HM, Jiang N, Kang HX, Ning Y, Wang ZL, Xiao YH, Liu XL, Liu EM, Dai LY, Wang GL (2012) Molecular mapping of the blast resistance genes Pi2-1 and Pi51(t) in the durably resistant rice ‘Tianjingyeshengdao’. Phytopathology 102:779–786
Wu JL, Fan YY, Li DB, Zheng KL, Leung H, Zhuang JY (2005) Genetic control of rice blast resistance in the durably resistant cultivar Gumei 2 against multiple isolates. Theor Appl Genet 111:50–56
Zenbayashi K, Ashizawa T, Tani T, Koizumi S (2002) Mapping of the QTL (quantitative trait locus) conferring partial resistance to leaf blast in rice cultivar Chubu 32. Theor Appl Genet 104:547–552
Zeng ZB (1994) Precision mapping of quantitative trait loci. Genetics 136:1457–1468
Zhou B, Qu SH, Liu GF, Dolan M, Sakai H, Lu GD, Bellizzi M, Wang GL (2006) The eight amino-acid differences within three leucine-rice repeats between Pi2 and Piz-t resistance proteins determine the resistance specificity to Magnaporthe grisea. Mol Plant-Microbe Interact 11:1216–1228
Zhu ML, Wang L, Pan QH (2004) Identification and characterization of a new blast resistance gene located on rice chromosome 1 through linkage and differential analyses. Phytopathology 94:515–519
Zhu XY, Chen S, Yang JY, Zhou SC, Zeng LX, Han JL, Su J, Wang L, Pan QH (2012) The identification of Pi50(t), a new member of the rice blast resistance Pi2/Pi9 multigene family. Theor Appl Genet 124:1295–1304
Acknowledgments
This work was partially supported by Grants from the National High Technology Research (2012AA101102, 2012AA10A303), the National Program on Research and Development of Transgenic Plants of China (2011ZX08001-002), the Foundation of the Ministry of Agriculture (CARS-01-03, 2011-Z66), and the National Natural Science Foundation of China (31171617) and the Bill & Melinda Gates Foundation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Haichao Jiang and Bin Yan contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Jiang, H., Yan, B., Duan, T. et al. Mapping and evaluating quantitative trait loci for blast resistance under natural infection conditions using an advanced backcross population in rice. Euphytica 204, 121–133 (2015). https://doi.org/10.1007/s10681-014-1347-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-014-1347-2