Abstract
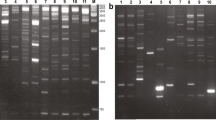
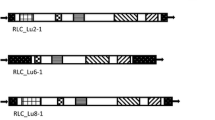
Heterosis is a main force leading the development of the hybrid seed industry in sunflower. The purpose of this study is to evaluate if heterosis effects for morphological traits among sunflower hybrids can be related to differences in the repetitive component of the genome of parental lines. The assumption is that, at least for certain traits, heterosis results from mutations in the cis-regulatory elements of genes, largely related to retrotransposon insertions and/or removals. Our experimental approach consists of a correlation study between hybrid performance and retrotransposon-related genetic distances between inbreds. Six sunflower inbred lines of different origin were crossed in a half diallel fashion; comparing parental lines and hybrids, mid parent heterosis of F1 hybrids was evaluated for six traits. We estimated the parental genetic distances between the six inbreds on data gathered by the inter-retrotransposon-amplified-polymorphism (IRAP) protocol. Different retrotransposons previously isolated in sunflower were targeted by 11 primer pairs designed on conserved LTR domains. As a control, genetic distances were also calculated using 86 genic SNPs. We analysed the correlation between the mid-parent heterosis for each of the six traits analysed and the genetic distance (calculated on data obtained by SNP or IRAP analyses) between the parental lines. Differences between parents showed to be largely related to variations in the retrotransposon component of the genome. Retrotransposon-related genetic distance between parents resulted to be larger than that related to genic SNPs, and significantly correlated to seed yield and, at a lesser extent, to plant height and stem diameter in hybrids. The hypothesis that variations in the repetitive component of the genome, especially LTR-retrotransposons, affect the displaying of heterosis is discussed.




Similar content being viewed by others
References
Ahmad S, Khan MS, Swati MS, Shah GS, Khalil IH (2005) A study on heterosis and inbreeding depression in sunflower (Helianthus annuus L.). Songklanakarin J Sci Technol 27:1–8
Boppenmaier J, Melchinger AE, Brunklaus-Jung E, Geiger HH, Herrmann RG (1992) Genetic distance for RFLPs in European maize inbreds: I. Relation to performance of flint dent crosses for forage traits. Crop Sci 32:895–902
Brunner S, Fengler K, Morgante M, Tingey S, Rafalski A (2005) Evolution of DNA sequence nonhomologies among maize inbreds. Plant Cell 17:343–360
Buckler ES, Gaut BS, McMullen MD (2006) Molecular and functional diversity of maize. Curr Opin Plant Biol 9:172–176
Buti M, Giordani T, Vukich M, Gentzbittel L, Pistelli L, Cattonaro F, Morgante M, Cavallini A, Natali L (2009) HACRE1, a recently inserted copia-like retrotransposon of sunflower (Helianthus annuus L.). Genome 52:904–911
Buti M, Giordani T, Cattonaro F, Cossu RM, Pistelli L, Vukich M, Morgante M, Cavallini A, Natali L (2011) Temporal dynamics in the evolution of the sunflower genome as revealed by sequencing and annotation of three large genomic regions. Theor Appl Genet 123:779–791
Cavallini A, Natali L, Zuccolo A, Giordani T, Jurman I, Ferrillo V, Vitacolonna N, Sarri V, Cattonaro F, Ceccarelli M, Cionini PG, Morgante M (2010) Analysis of genome composition and organization in sunflower (Helianthus annuus L.) and related species. Theor Appl Genet 120:491–508
Charcosset A, Gallais A (2003) Application of markers in selection. In: de Vienne D (ed) Molecular markers in plants genetics and biotechnology. Science Publishers, Enfield, pp 53–176
Charcosset AM, Lefort-Buson M, Gallais A (1991) Relationship between heterosis and heterozygosity at marker loci: a theoretical computation. Theor Appl Genet 81:571–575
Cheres MT, Knapp SJ (1998) Ancestral origins and genetic diversity of cultivated sunflower: analysis of the pedigrees of public germplasm. Crop Sci 38:1476–1482
Cheres MT, Miller JF, Crane JM, Knapp SJ (2000) Genetic distance as a predictor of heterosis and hybrid performance within and between heterotic groups in sunflower. Theor Appl Genet 100:889–894
Ching A, Caldwell KS, Jung M, Dolan M, Smith OS, Tingey S, Morgante M, Rafalski AJ (2002) SNP frequency, haplotype structure and linkage disequilibrium in elite maize inbred lines. BMC Genet 3:19
Clark RM, Wagler TN, Quijada P, Doebley J (2006) A distant upstream enhancer at the maize domestication gene tb1 has pleiotropic effects on plant and inflorescent architecture. Nat Genet 38:594–597
Darvishzadeh R (2012) Phenotypic and molecular marker distance as a tool for prediction of heterosis and F1 performance in sunflower (Helianthus annuus L.) under well-watered and water-stressed conditions. Aust J Crop Sci 6:732–738
Davenport CB (1908) Degeneration, albinism and inbreeding. Science 28:454–455
Diers BW, Mac Vetty PBE, Osborn TC (1996) Relationship between heterosis and genetic distance based on restriction fragment length polymorphism markers in oilseed rape (Brassica napus L.). Crop Sci 36:79–83
Dubreuil P, Dufour P, Krejci E, Causse M, de Vienne D, Gallais A, Charcosset A (1996) Organization of RFLP diversity among inbred lines of maize representing the most significant heterotic groups. Crop Sci 36:790–799
Duvick DN (1999) Heterosis: feeding people and protecting natural resources. In: Coors JG, Pandey S (eds) The genetics and exploitation of heterosis in crops. Crop Science Society of America, Madison, pp 19–30
East EM (1908) Inbreeding in corn. Rep Conn Agric Exp Stn 1907:419–429
Felsenstein J (1989) PHYLIP—Phylogeny Inference Package (Version 3.2). Cladistics 5:164–166
Frascaroli E, Canè MA, Landi P, Pea G, Gianfranceschi L, Villa M, Morgante M, Pè ME (2007) Classical genetic and quantitative trait loci analyses of heterosis in a maize hybrid between two elite inbred lines. Genetics 176:625–644
Giordani T, Natali L, D’Ercole A, Pugliesi C, Fambrini M, Vernieri P, Vitagliano C, Cavallini A (1999) Expression of a dehydrin gene during embryo development and drought stress in ABA-deficient mutants of sunflower (Helianthus annuus L.). Plant Mol Biol 39:739–748
Giordani T, Buti M, Natali L, Pugliesi C, Cattonaro F, Morgante M, Cavallini A (2010) An analysis of sequence variability in eight genes putatively involved in drought response in sunflower (Helianthus annuus L.). Theor Appl Genet 122:1039–1049
Godshalk EB, Lee M, Lamkey KR (1990) Relationship of restriction fragment length polymorphisms to single-cross hybrid performance in maize. Theor Appl Genet 80:273–280
Guo M, Rupe MA, Zinselmeier C, Habben J, Bowen BA, Smith OS (2004) Allelic variation of gene expression in maize hybrids. Plant Cell 16:1707–1716
Guo M, Rupe MA, Yang X, Crasta O, Zinselmeier C, Smith OS, Bowen B (2006) Genome-wide transcript analysis of maize hybrids: allelic additive gene expression and yield heterosis. Theor Appl Genet 113:831–845
Hongtrakul V, Huestis GM, Knapp SJ (1997) Amplified fragment length polymorphisms as a tool for DNA fingerprinting sunflower germplasm: genetic diversity among oilseed inbred lines. Theor Appl Genet 95:400–407
Jaccard P (1908) Nouvelles recherches sur la distribution florale. Bull Soc Vaud Sci Nat 44:223–270
Janick J (1998) Hybrids in horticultural crops. In: Lamkey KR, Staub JE (eds) Concepts and breeding of heterosis in crop plants. Crop Science Society of America, Madison, pp 45–56
Jones DF (1917) Dominance of linked factors as a means of accounting for heterosis. Genetics 2:466–479
Kalendar R, Grob T, Regina M, Suoniemi A, Schulman A (1999) IRAP and REMAP: two new retrotransposon-based DNA fingerprinting techniques. Theor Appl Genet 98:704–711
Kaya Y (2005) Hybrid vigor in sunflower (Helianthus annuus L.). Helia 28:77–86
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Kobayashi S, Goto-Yamamoto N, Hirochika H (2004) Retrotransposon-induced mutations in grape skin color. Science 304:982
Krystkowiak K, Adamski T, Surma M, Kaczmarek Z (2009) Relationship between phenotypic and genetic diversity of parental genotypes and the specific combining ability and heterosis effects in wheat (Triticum aestivum L.). Euphytica 165:419–434
Lai J, Li R, Xu X, Jin W, Xu M, Zhao H et al (2010) Genome-wide patterns of genetic variation among elite maize inbred lines. Nature Genet 42:1027–1030
Martin JM, Talbort LE, Lanning SP, Blake NK (1995) Hybrid performance in wheat as related to parental diversity. Crop Sci 35:104–108
Melchinger AE (1999) Genetic diversity and heterosis. In: Coors JG, Pandey S (eds) The genetics and exploitation of heterosis in crops. Crop Science Society of America, Madison, pp 99–118
Melchinger AE, Gumber RK (1998) Overview of heterosis and heterotic groups in agronomic crops. In: Lamkey KR, Staub JE (eds) Concepts and breeding of heterosis in crop plants. Crop Science Society of America, Madison, pp 29–44
Melchinger AE, Lee M, Lamkey KR, Woodman WW (1990) Genetic diversity for restriction fragment length polymorphisms: relation to genetic effects in maize inbreds. Crop Sci 30:1033–1040
Melchinger AE, Utz HF, Piepho HP, Zeng ZB, Schön CC (2007) The role of epistasis in the manifestation of heterosis: a systems-oriented approach. Genetics 177:1815–1825
Messing J, Dooner HK (2006) Organization and variability of the maize genome. Curr Opin Plant Biol 9:157–163
Meyers BC, Scalabrin S, Morgante M (2004) Mapping and sequencing complex genomes: let’s get physical. Nat Rev Genet 5:578–588
Miller JF (1992) Registration of five oilseed sunflower germplasm restorer lines (RHA 373 to 377) and two nuclear male sterile populations (NMS 274 and 801). Crop Sci 32:1298
Moll RH, Salhuana WS, Robinson HF (1962) Heterosis and genetic diversity in variety crosses of maize. Crop Sci 2:197–198
Morgante M, De Paoli E, Radovic S (2007) Transposable elements and the plant pan-genomes. Curr Opin Plant Biol 10:149–155
Moser H, Lee M (1994) RFLP variation and genealogical distance, multivariate distance, heterosis, and genetic variance in oats. Theor Appl Genet 87:947–956
Natali L, Giordani T, Cavallini A (2003) Sequence variability of a dehydrin gene within Helianthus annuus. Theor Appl Genet 106:811–818
Natali L, Santini S, Giordani T, Minelli S, Maestrini P, Cionini PG, Cavallini A (2006) Distribution of Ty3-gypsy- and Ty1-copia-like DNA sequences in the genus Helianthus and other Asteraceae. Genome 49:64–72
Ouvrard O, Cellier F, Ferrare K, Tousch D, Lamaze T, Dupuis J-M, Casse-Delbart F (1996) Identification and expression of water stress- and abscisic acid-regulated genes in a drought-tolerant sunflower genotype. Plant Mol Biol 31:819–829
Paschold A, Jia Y, Marcon C, Lund S, Larson NB, Yeh CT, Ossowski S, Lanz C, Nettleton D, Schnable PS, Hochholdinger F (2012) Complementation contributes to transcriptome complexity in maize (Zea mays L.) hybrids relative to their inbred parents. Genome Res 22(12):2445–2454. doi:10.1101/gr.138461.112
Rohlf FJ (2008) NTSYSpc: numerical taxonomy system, ver. 2.00. Exeter Publishing Ltd., Setauket
Rychlik W, Rhoads RE (1989) A computer program for choosing optimal oligonucleotides for filter hybridization, sequencing and in vitro amplification of DNA. Nucl Acids Res 17:8543–8551
Sheng JX, Lu GY, Fu TD, Yang GS (2002) Relationships between genetic diversity and hybrid performance in oilseed rape (Brassica napus). Acta Agron Sin 28:622–627
Shull GH (1908) The composition of a field of maize. Am Breed Assoc Rep 4:296–301
Sokal RR, Michener CD (1958) A statistical method for evaluating systematic relationships. Univ Kans Sci Bull 38:1409–1438
Springer NM, Stupar RM (2007) Allelic variation and heterosis in maize: how do two halves make more than a whole? Genome Res 17:264–275
Springer NM, Ying K, Fu Y, Ji T, Yeh CT, Jia Y, Wu W, Richmond T et al (2009) Maize inbreds exhibit high levels of copy number variation (CNV) and presence/absence variation (PAV) in genome content. PLoS Genet 5:e1000734
Stam M, Belele C, Dorweiler JE, Chandler VL (2002) Differential chromatin structure within a tandem array 100 kb upstream of the maize b1 locus is associated with paramutation. Genes Dev 16:1906–1918
Staton SE, Bakken BH, Blackman BK, Chapman MA, Kane NC, Tang S, Ungerer MC, Knapp SJ, Rieseberg LH, Burke JM (2012) The sunflower (Helianthus annuus L.) genome reflects a recent history of biased accumulation of transposable elements. Plant J 72:142–153
Stupar RM, Springer NM (2006) Cis-transcriptional variation in maize inbred lines B73 and Mo17 leads to additive expression patterns in the F1 hybrid. Genetics 173:2199–2210
Tersac M, Blanchard P, Brunel D, Vancourt P (1994) Relations between heterosis and enzymatic polymorphisms in populations of cultivated sunflower (Helianthus annuus L.). Theor Appl Genet 88:49–55
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTALW: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucl Acids Res 22:4673–4680
Vukich M, Schulman AH, Giordani T, Natali L, Kalendar R, Cavallini A (2009a) Genetic variability in sunflower (Helianthus annuus L.) and in the Helianthus genus as assessed by retrotransposon-based molecular markers. Theor Appl Genet 119:1027–1038
Vukich M, Giordani T, Natali L, Cavallini A (2009b) Copia and Gypsy retrotransposons activity in sunflower (Helianthus annuus L.). BMC Plant Biol 9:150
Zanoni U, Dudley JW (1989) Comparison of different methods of identifying inbreds useful for improving elite maize hybrids. Crop Sci 29:577–582
Zhang Q, Gao YJ, Saghai Maroof MA, Yang SH, Li JX (1995) Molecular divergence and hybrid performance in rice. Mol Breed 1:133–142
Zhang G, Angeles ER, Abenes MLP, Khush GS, Huang N (1996) RAPD and RFLP mapping for the bacterial blight resistance gene xa-13 in rice. Theor Appl Genet 93:65–70
Acknowledgments
Thanks are due to Dr. Elisabetta Frascaroli (University of Bologna) for her useful suggestions during the investigation and to Dr. Andrea Cini (Kayser Italia) for revising the manuscript. This work was supported by PRIN-MIUR, Projects “Variabilità di sequenza ed eterosi in piante coltivate” and “SUNREP: caratterizzazione molecolare della componente ripetitiva del genoma di girasole”.
Author information
Authors and Affiliations
Corresponding author
Additional information
M. Buti and T. Giordani contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Buti, M., Giordani, T., Vukich, M. et al. Retrotransposon-related genetic distance and hybrid performance in sunflower (Helianthus annuus L.). Euphytica 192, 289–303 (2013). https://doi.org/10.1007/s10681-013-0883-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-013-0883-5