Abstract
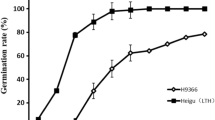
A previously constructed introgression lines (ILs) population including 239 lines was used to evaluate the cold-tolerant ability. The ILs with strong cold tolerance which were identified in the present study were used for further cytological and microsatellite (SSR) marker analyses. The results showed that the IL5243 and IL5335 had strong cold-tolerant ability. Cytological analysis showed that the rate of pollen mother cells (PMCs) with normal meiotic behavior in the IL5243 and IL5335 was to 89.93 and 90.22 %, respectively, and finally formed normal mature pollen. And meanwhile, the low frequency of abnormal chromosome behavior was observed in the IL5243 and IL5335, such as univalent, 8-shape bivalent, multivalent at diakinesis. At anaphase I, one or two lagging chromosomes were observed in some PMCs (3.95–5.15 %). The results of SSR marker analysis further confirmed that partial alien DNA of common wild rice has been transferred into the IL5243 and IL5335. These results implied that the IL5243 and IL5335 might be excellent bridging germplasm for exploring and utilizing the cold-tolerant gene of common wild rice. In addition, IL5243 and IL5335 would provide a better experimental system for understanding some epigenetic phenomenon induced by alien gene introgression.





Similar content being viewed by others
References
Andaya VC, Mackill DJ (2003) Mapping of QTL associated with cold tolerance during the vegetative stage in rice. J Exp Bot 54:2579–2585
Andaya VC, Tai TH (2007) Fine mapping of the qCTS4 locus associated with seedling cold tolerance in rice (Oryza sativa L.). Mol Breeding 20:349–358
Ansari HA, Ellison NW, Williams WM (2008) Molecular and cytogenetic evidence for an allotetraploid origin of Trifolium dubium (Leguminosae). Chromosoma 117:159–167
Bertin P, Bouharmont J (1997) Use of somaclonal variation and in vitro selection for chilling tolerance improvement in rice. Euphytica 96:135–142
Chen JF, Luo XD, Qian CT, Molly MJ, Staub JE, Zhuang FY, Lou QF, Ren G (2004) Cucumis monosomic alien addition lines: morphological, cytological, and genotypic analyses. Theor Appl Genet 108:1343–1348
Chen LZ, Lou QF, Zhuang Y, Chen JF, Zhang XQ, Wolukau JN (2007) Cytological diploidization and rapid genome changes of the newly synthesized allotetraploids Cucumis × hytivus. Planta 225:603–614
Chen XR, Yang KS, Fu JR, Zhu CL, Peng XS, He P, He HH (2008) Identification and genetic analysis of fertility restoration ability in Dongxiang wild rice (Oryza rufipogon). Rice Sci 15:21–28
Fujino K (2004) A major gene for low temperature germinability in rice (Oryza sativa L.). Euphytica 136:63–68
Groszmann M, Greaves IK, Albertyn ZI, Scofield GN, Peacock WJ, Dennis ES (2011) Changes in 24-nt siRNA levels in Arabidopsis hybrids suggest an epigenetic contribution to hybrid vigor. Proc Natl Acad Sci USA 108:2617–2622
Guo JY, Chen JF, Cao QH, Luo XD, Chen LZ (2004) Cytological studies on microsporogenesis and male gametophyte development of a Cucumis allotriploid derived from C. hytivus × C. sativus. Cytologia 69:335–340
Han LZ, Qiao YL, Cao GL, Zhang YY, An YP, Ye JD, Koh HJ (2005) QTL analysis on cold tolerance during early growth period in rice. Chin J Rice Sci 19:122–126
He GM, Zhu XP, Elling AA, Chen LB, Wang XF, Guo L, Liang MZ, He H, Zhang HY, Chen FF, Qi YJ, Chen RS, Deng XW (2010) Global epigenetic and transcriptional trends among two rice subspecies and their reciprocal hybrids. Plant Cell 22:17–33
Liu FX, Sun CQ, Tan LB, Fu YC, Li DJ, Wang XK (2003) Identification of QTL for cold tolerance at booting and flowering stage in Jiangxi Dongxiang wild rice. Chin Sci Bull 48:1864–1867
Lou QJ, Chen L, Sun ZX, Xing YZ, Li J, Xu XY, Mei HW, Luo LJ (2007) A major QTL associated with cold tolerance at seedling stage in rice (Oryza sativa L.). Euphytica 158:87–94
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15:473–497
Murray HG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucl Acids Res 8:4321–4326
Osborn TC, Pires JC, Birchler JA, Auger DL, Chen ZJ, Lee HS, Comai L, Madlung A, Doerge RW, Colot V, Martienssen RA (2003) Understanding mechanisms of novel gene expression in polyploids. Trends Genet 19:141–147
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Qi B, Zhong X, Zhu B, Zhao N, Xu L, Zhang H, Yu X, Liu B (2010) Generality and characteristics of genetic and epigenetic changes in newly synthesized allotetraploid wheat lines. J Genet Genomics 37:737–748
Qu TT, Chen LY, Zhang ZH, Hu ZL, Li P, Zhu LH, Zhu YG (2003) Molecular mapping of genes conferring cold tolerance at seedling stage using doubled haploid lines from an indica × japonica cross in rice. J Wuhan Bot Res 21:385–389
Rahman L, Khanam MS, Koh HJ (2008) QTL analysis for yield related traits using populations derived from an indica-japonica hybrid in rice (Oryza sativa L.). Czech J Genet Plant Breed 44:93–104
Sanguinetti CJ, Dias NE, Simpson AJ (1994) Rapid silver staining and recovery of PCR products separated on polyacrylamide gels. Biotechniques 17:914–921
Sthapit BR, Witcombe JR (1998) Inheritance of tolerance to chilling stressing rice during germination and plumule greening. Crop Sci 38:660–665
Takesawa T, Ito M, Kanzaki H, Kameya N, Nakamura I (2002) Over-expression of ζ glutathione S-transferase in transgenic rice enhances germination and growth at low temperature. Mol Breed 9:93–101
Tan LB, Liu FX, Xue W, Wang GJ, Ye S, Zhu ZF, Fu YC, Wang XK, Sun CQ (2007) Development of Oryza rufipogon and O. sativa introgression lines and assessment for yield-related quantitative trait loci. J Integr Plant Biol 49:871–884
Tanksley SD, Nelson JC (1996) Advanced backcross QTL analysis: a method for the simultaneous discovery and transfer of valuable QTLs from unadapted germplasm into elite breeding lines. Theor Appl Genet 92:91–203
Thomson MJ, Edwards JD, Septiningsih EM, Harrington SE, McCouch SR (2006) Substitution mapping of dth1.1, a flowering-time quantitative trait locus (QTL) associated with transgressive variation in rice, reveals multiple sub-QTL. Genetics 172:2501–2514
Tian F, Li DJ, Fu Q, Zhu ZF, Fu YC, Wang XK, Sun CQ (2006) Construction of introgression lines carrying wild rice (Oryza rufipogon Griff.) segments in cultivated rice (Oryza sativa L.) background and characterization of introgressed segments associated with yield-related traits. Theor Appl Genet 112:570–580
Wang WS, Pan YJ, Zhao XQ, Dwivedi D, Zhu LH, Ali J, Fu BY, Li ZK (2011) Drought-induced site-specific DNA methylation and its association with drought tolerance in rice (Oryza sativa L.). J Exp Bot 62:1951–1960
Xie JK, Hu BL, Wan Y, Zhang T, Li X, Liu RN, Huang YH, Dai LF, Luo XD (2010a) Comparison of the drought resistance characters at seedling stage between Dongxiang common wild rice (Oryza rufipogon Griff.) and cultivars (Oryza sativa L.). Acta Ecol Sin 30:1665–1674
Xie JK, Agrama HA, Kong D, Zhuang J, Hu B, Wan Y, Yan W (2010b) Genetic diversity associated with conservation of endangered Dongxiang wild rice (Oryza rufipogon). Genet Resour Crop Evol 57:597–609
Xu L, He Y, Zhang DF, Dai JR, Wang SC (2009) Identification and fine-mapping of a bacterial brown spot disease resistance gene in maize. Mol Breed 23:709–718
Zeng YW, Pu XY (2006) Genetic analysis for cold tolerance at booting stage for japonica rice (Oryza sativa L.). Indian J Genet Plant Breed 66:100–102
Zeng YW, Yang SM, Cui H, Yang XJ, Xu LM, Du J, Pu XY, Li ZC, Cheng ZQ, Huang XQ (2009) QTLs of cold tolerance-related traits at the booting stage for NIL-RILs in rice revealed by SSR. Genes Genomics 31(2):143–154
Zhan QC, Zhu KY, Chen ZW, Zeng SZ (2005) Studies on the QTL for cold tolerance related characters of rice seedling by molecular markers. Hybrid rice 20:50–55
Zhang ZH, Li S, Wei L, Wei C, Zhu YG (2005) A major QTL conferring cold tolerance at the early seedling stage using recombinant inbred lines of rice (Oryza sativa L.). Plant Sci 168:527–534
Zhang XS, Zhou X, Fu YC, Zhen S, Wang XK, Sun CQ (2006) Identification of a drought tolerant introgression line derived from Dongxiang common wild rice (O. rufipogon Griff.). Plant Mol Biol 62:247–259
Zhang YS, Luo L, Liu T, Xu C, Xing Y (2009) Four rice QTL controlling number of spikelets per panicle expressed the characteristics of single Mendelian gene in near isogenic backgrounds. Theor Appl Genet 118:1035–1044
Acknowledgments
This research was partially supported by the National Natural Science Foundation of China (30860120 and 30900781), the Natural Science Foundation of Jiangxi Province, China (2008GQN0059), the Development Program for Young Scientists of Jiangxi Province, China (20112BCB23007), and the Scientific Planning Project of Jiangxi Provincial Education Department (GJJ12184).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Luo, XD., Dai, LF., Cao, JF. et al. Identification and molecular cytology analysis of cold tolerance introgression lines derived from Oryza sativa L. mating with O. rufipogon Griff.. Euphytica 187, 461–469 (2012). https://doi.org/10.1007/s10681-012-0769-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-012-0769-y