Abstract
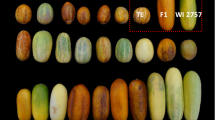
Two cucumber recombinant inbred lines (RILs) differing in plant habit were crossed and progeny self-pollinated to produce F3 individuals upon which phenotypic selection was practiced to identify a base population which in turn underwent either two cycles of MAS or random mating without selection (RAN). MAS and RAN were practiced to produce F4 and F5 progeny sets. RIL, crossing parents, and F3–F5 progeny sets were then evaluated under replicated field conditions for fruit yield and quality (L:D and E:T) to evaluate gain from selection (ΔG). The broad-sense heritability (h 2 B) over cycles (C) of selection ranged 0.22–0.45, 0.09–0.20, and 0.11–0.15 for yield, L:D, and E:T, respectively. Although one cycle of PHE selection followed by MAS was effective in conserving the performance of the traits examined during inbreeding, progeny performance during RAN fluctuated (F4–F5 generation; C2). Lack of ΔG during advanced generations (F4–F5) of MAS was likely due to allelic fixation and/or optimized epistatic complementation.
Similar content being viewed by others
References
Behera TK, Staub JE, Behera S, Rao AR, Mason S (2008) One cycle of phenotypic selection combined with marker assisted selection for improving yield and quality in cucumber. In: Pitrat M (ed) Proceedings of the IXth EUCARPIA meeting on genetics and breeding of Cucurbitaceae. Avignon, France, pp 115–121
Dudley JW (1993) Molecular markers in plant improvement: manipulation of genes affecting quantitative traits. Crop Sci 33:660–668
Edwards MD, Stuber CW, Wendel JF (1987) Molecular marker-facilitated investigations of quantitative trait loci in maize. I. Numbers, distribution, and types of gene action. Genetics 116:113–125
Fan Z, Robbins MD, Staub JE (2006) Population development by phenotypic selection with subsequent marker-assisted selection for line extraction in cucumber (Cucumis sativus L.). Theor Appl Genet 112:843–855
Fazio G (2001) Comparative study of marker assisted and phenotypic selection and genetic analysis of yield components in cucumber. PhD dissertation, University of Wisconsin, Madison
Fazio G, Staub JE, Stevens MR (2003a) Genetic mapping and QTL analysis of horticultural traits in cucumber (Cucumis sativus L.) using recombinant inbred lines. Theor Appl Genet 107:864–874
Fazio G, Chung SM, Staub JE (2003b) Comparative analysis of response to phenotypic and marker-assisted selection for multiple lateral branching in cucumber (Cucumis sativus L.). Theor Appl Genet 107:875–883
Flint-Garcia SA, Darrah LL, McMullen MD, Hibbard BE (2003) Phenotypic versus marker-assisted selection for stalk strength and second-generation European corn borer resistance in maize. Theor Appl Genet 107:1331–1336
Gimelfarb A, Lande R (1994) Simulation of marker assisted selection in hybrid populations. Genet Res 63:39–47
Hospital F, Moreau L, Lacoudre F, Charcosset A, Gallais A (1997) More on the efficiency of marker-assisted selection. Theor Appl Genet 95:1181–1189
Kasha KJ (1999) Biotechnology and world food supply. Genome 42:642–645
Kennard WC, Havey MJ (1995) Quantitative trait analysis of fruit quality in cucumber: QTL detection, confirmation, and comparison with mating design. Theor Appl Genet 91:53–61
Knapp SJ (1998) Marker-assisted selection as a strategy for increasing the probability of selecting superior genotypes. Crop Sci 38:1164–1174
Lande R, Thompson R (1990) Efficiency of marker-assisted selection in the improvement of quantitative traits. Genetics 124:743–756
Lecomte L, Duffe P, Buret M, Servin B, Hospital F, Causse M (2004) Marker-assisted introgression of five QTLs controlling fruit quality traits into three tomato lines revealed interactions between QTLs and genetic backgrounds. Theor Appl Genet 109:658–668
Littell RC, Milliken GA, Stroup WW, Wolfinger RD (1996) SAS system for mixed models. SAS Institute Inc, Cary
Lower RL, Edwards MD (1986) Cucumber breeding. In: Basset MJ (ed) Breeding vegetable crops. AVI Publishing Co., Westport, pp 173–207
Miller MP (1997) Tools for population genetic analysis (TEPGA), Version 3. Department of Biological Sciences. Northern Arizona University, Arizona, USA
Moreau L, Charcosset A, Hospital F, Gallais A (1998) Marker assisted selection efficiency in populations of finite size. Genetics 148:1353–1365
Moreau L, Lamarie S, Charcosset A, Gallais A (2000) Economic efficiency of one cycle of marker assisted selection. Crop Sci 40:329–337
Moreau L, Charcosset A, Gallais A (2004) Experimental evaluation of several cycles of marker-assisted selection in maize. Euphytica 137:111–118
Nei M (1972) Genetic distance between populations. Am Nat 106:283–292
Ortiz R (1998) Critical role of plant biotechnology for the genetic improvement of food crops: Perspectives for the next millennium. Electron J Biotechnol 1:152–159
Paterson AH, Tanksley SD, Sorrells ME (1991) DNA markers in plant improvement. Adv Agron 46:39–90
Robbins MD (2006) Molecular marker development, QTL pyramiding, and comparative analysis of phenotypic and marker assisted selection in cucumber. PhD dissertation University of Wisconsin, Madison
Robbins MD, Staub JE (2009) Comparative analysis of marker-assisted and phenotypic selection for yield components in cucumber. Theor Appl Genet 119:621–634
Robbins MD, Casler M, Staub JE (2008) Pyramiding QTL for multiple lateral branching in cucumber using nearly isogenic lines. Mol Breed 22:131–139
SAS software, Version 8 of SAS System for Unix (1999) SAS Institute Inc., Cary
Serquen FC, Bacher J, Staub JE (1997a) Genetic analysis of yield components in cucumber (Cucumis sativus L.) at low plant density. J Am Soc Hortic Sci 122:522–528
Serquen FC, Bacher J, Staub JE (1997b) Mapping and QTL analysis of a narrow cross in cucumber (Cucumis sativus L.) using random amplified polymorphic DNA markers. Mol Breed 3:257–268
Staub JE, Bacher J (1997) Cucumber as a processed vegetable. In: Smith DS, Cash JN, Nip W, Hui YH (eds) Processing vegetables: science and technology IV. Technomic Publishing Co, Inc Lancaster, PA, pp 129–193
Staub JE, Grumet R (1993) Selection for multiple disease resistance affects cucumber yield potential. Euphytica 67:205–213
Staub JE, Balgooyen B, Tolla GE (1986) Quality and yield of cucumber hybrids using gynoecious and bisexual pollen parents. HortScience 21:510–512
Staub JE, Serquen FC, Gupta M (1996) Genetic markers, map construction and their application in plant breeding. HortScience 31:729–741
Steele R, Torrie JH (1980) Principles and procedures of statistics, 2nd edn. McGraw-Hill, New York
Steele KA, Edwards G, Zhu J, Witcombe RC (2004) Marker evaluated selection in rice: shifts in allele frequency among bulks selected in contrasting agricultural environments identify genomic regions of importance to rice adaptation and breeding. Theor Appl Genet 109:1247–1260
Tanksley SD, Medino-filho H, Rick CM (1981) The effect of isozyme selection on metric characters in an inter-specific backcross of tomato: Basis of an early screen procedure. Theor Appl Genet 60:291–296
Tatlioglu T (1993) Cucumber (Cucumis sativus L.). In: Kalloo GB, Bergh O (eds) Genetic improvement of vegetable crops. Pergamon Press Ltd, Tarrytown, pp 197–234
Thabuis A, Palloix A, Servin B, Daubeze AM, Signoret P, Hospital F, Lefebvre V (2004) Marker-assisted introgression of 4 Phytophthora capsici resistance QTL alleles into a bell pepper line: validation of additive and epistatic effects. Mol Breed 14:9–20
Wehner TC (1989) Breeding for improved yield in cucumber. Plant Breed Rev 6:323–359
Willcox MC, Khairallah MM, Bergvinson D, Crossa J, Deutsch JA, Edmeades GO, Gonzalez de Leon D, Jiang C, Jewell DC, Mihm JA, Williams WP, Hoisington D (2002) Selection for resistance to southwestern corn borer using marker-assisted and conventional backcrossing. Crop Sci 42:1516–1528
Xie C, Xu S (1998) Efficiency of multistage marker-assisted selection in the improvement of multiple quantitative traits. Heredity 80:489–498
Yeh FC, Boyle TJB (1997) Population genetic analysis of co-dominant and dominant markers and quantitative traits. Belg J Bot 129:157
Acknowledgments
The fund provided by the Department of Biotechnology, Ministry of Science and Technology, Govt. of India for sponsoring T.K. Behera to carry out research work at Department of Horticulture, UW Madison, USA is highly acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Behera, T.K., Staub, J.E., Behera, S. et al. Response to phenotypic and marker-assisted selection for yield and quality component traits in cucumber (Cucumis sativus L.). Euphytica 171, 417–425 (2010). https://doi.org/10.1007/s10681-009-0072-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-009-0072-8