Abstract
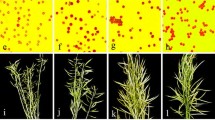
Two wide hybrids, Diplotaxis erucoides (2n = 14) × Brassica rapa (2n = 20) and B. maurorum (2n = 20) × B. rapa, were developed using the sequential ovary–ovule culture. Reciprocal crosses failed, possibly as a consequence of strong unilateral incompatibility. The F 1 hybrids in each combination were completely male sterile and morphologically intermediate to the respective parents. DNA marker polymorphism and chromosome counts confirmed their hybrid nature. High frequency of bivalents in the F 1 and the presence of trivalents/quadrivalents in the derived amphiploids suggested genomic duplications and homoeology of the parental genomes. Up to three homoeologous pairs between the D. erucoides (DeDe) and B. rapa (AA) genomes, and one between B. maurorum (BmBm) and B. rapa genomes were observed. Successful synthesis of the F 1 hybrids and amphiploids of B. rapa with D. erucoides and B. maurorum, and allosyndetic chromosome pairing are expected to permit introgressions of desirable loci into the cultivated Brassica germplasm, especially for resistance to Alternaria brassicae and Albugo candida.



Similar content being viewed by others
References
Armstrong KC, Keller WA (1981) Chromosome pairing in haploids of Brassica campestris. Theor Appl Genet 59:49–52
Banga SS, Deol JS, Banga SK (2003) Alloplasmic male sterile Brassica juncea with Enarthrocarpus lyratus cytoplasm and the introgression of gene(s) for fertility restoration from cytoplasm donor species. Theor Appl Genet 106:1390–1395
Bhaskar PB, Ahuja I, Janeja HS, Banga SS (2002) Intergeneric hybridization between Erucastrum canarience and Brassica rapa : Genetic relatedness between EC and A genomes. Theor Appl Genet 105:754–758
Chandra A, Gupta ML, Ahuja I, Kaur G, Banga SS (2004) Intergeneric hybridization between Erucastrum cardaminoides and two diploid crop brassica species. Theor Appl Genet 108(8):1620–1626
Chrungu B, Verma N, Mohanty A, Pradhan A, Shivanna KR (1999) Production and characterization of interspecific hybrids between B. maurorum and crop Brassicas. Theor Appl Genet 98:608–613
Delourme R, Eber F, Cheavre AM (1989) Intergeneric hybridization of Diplotaxis erucoides with Brassica napus. I. Cytogenetic analysis of F 1 and BC1 progeny. Euphytica 41:123–128
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Fahleson J, Eriksson I, Landgrin M, Stymne S, Glimelius K (1994) Intertribal somatic hybrids between Brassica napus and Thlaspi perfoliatum with high content of T. perfoliatum specific nervonic acid. Theor Appl Genet 87:795–804
Hagimori M, Nagaoka M, Kato N, Yoshikawa H (1992) Production and characterization of somatic hybrids between Japanese radish and cauliflower. Theor Appl Genet 84:819–824
Jauhar PP, Crane CF (1989) An evaluation of Baum et al’s assessment of the genomic system of classification in the Triticeae. Am J Bot 76:571–576
Karpechenko GD (1924) Hybrids of Raphanus sativus x Brassica oleracea L. J Genet 14:375–396
Lysak MA, Koch MA, Pecinka A, Schubert I (2005) Chromosome triplication found across the tribe Brassicaceae. Genome Res 15:516–525
Quiros CF (1999) Genome structure and mapping. In: Gomez-Campo C (ed) Biology of Brassica coenospecies. Elsevier science, Amsterdam
Sageret M (1826) Considerations Sur la production des variants et des varieties esn general, et sur celles de la famille de cucurbitacaes on particular. Ann Sci Nat 8:294–314
Shivanna KR (1996) Incompatibility and wide hybridization. In: Chopra VL and Prakash S (eds) Oilseed and Vegetable Brassicas: Indian Perspective. Oxford and IBH, New Delhi, pp 77–102
Song KM, Osborn TC, Williams PH (1990) Brassica taxonomy based on nuclear restriction fragment length polymorphisms (RFLP’s). 3. Genome relationships in Brassica and related genera and the origin of B. oleracea and B. rapa (syn. campestris). Theor Appl Genet 79:497–506
Takahata Y, Hinata K (1983) Studies on cytodemes in Subtribe Brassicas (Cruciferae) Tohokee. J Agri Res 33:111–124
Truco MJ, Hu J, Sadowski J, Quiros CF (1996) Inter and Intragenomic homology of the Brassica genomes: implications for their origin and evolution. Theor Appl Genet 93:1225–1233
Tsunoda S (1980) Ecophysiology of wild and cultivated forms in Brassica and allied genera. In: Tsunoda S, Hinata K, Gomez-Campo C (eds) Brassica crops and wild allies. Biology and Breeding. Japan Scientific Press, Tokyo, pp. 109–120
Vyas P, Prakash S, Shivanna KR (1995) Production of wide hybrids and backcross progenies between Diplotaxis erucoides and crop brassicas. Theor Appl Genet 90:549–553
Warwick SI, Black LD (1991) Molecular systematics of Brassica and allied genera (Subtribe Brassianal, Brassiceae) chloroplast genome and cytodeme congruence. Theor Appl Genet 82:81–92
Acknowledgements
This study was partly supported by the funds provided under the Indian Council of Agricultural Research aided research project “National network for management of Alternaria blight in Brassica juncea and vegetable crops”. Sincere thanks to Prof. Adam Lukaszewski for helpful suggestions. Gratitude is also expressed to Prof. Shyam Prakash for the supply of seed samples of the wild crucifers.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Garg, H., Banga, S., Bansal, P. et al. Hybridizing Brassica rapa with wild crucifers Diplotaxis erucoides and Brassica maurorum . Euphytica 156, 417–424 (2007). https://doi.org/10.1007/s10681-007-9391-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-007-9391-9