Abstract
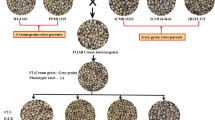
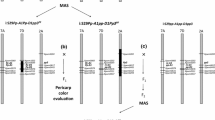
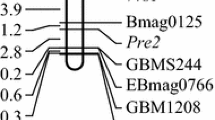
Complementary genes for purple grain colour Pp1, Pp2, Pp3 (now designated Pp1, Pp3b, Pp3a, respectively) were mapped using crosses between purple-grained hexaploid wheats ‘Purple Feed’ – Pp1Pp1/Pp2Pp2 (Pp1Pp1/Pp3bPp3b), ‘Purple’ – Pp1Pp1/Pp3Pp3 (Pp1Pp1/Pp3aPp3a) with non-purple-grained cultivars ‘Novosibirskaya 67’ (‘N67’) and ‘Saratovskaya 29’ (‘S29’). The genes Pp2 (Pp3b) and Pp3 (Pp3a) were inherited as monofactorial dominant when purple-grained wheats were crossed to ‘N67’. Both were mapped in the centromeric region of the chromosome 2A. Therefore, they were suggested being different alleles at the same locus and designated Pp3a and Pp3b. In the crosses between purple-grained wheats and ‘S29’ a segregation ratio of 9 (purple) to 7 (non purple) was obtained suggesting a complementary interaction of two dominant genes, Pp1 and Pp3. To map Pp1 as a single gene, the influence of the other Pp gene was taken into consideration by determining the Pp3 genotype of the F2 plants. The gene was mapped on chromosome 7BL, about 24 cM distal to the centromere. The Pp1gene was shown to be non allelic to the Rc-1 (red coleoptile) and Pc (purple culm) genes, contrary to what was previously suggested. The colouration caused by the Pp genes has no effect on pre-harvest sprouting.
Similar content being viewed by others
References
Abdel-Aal, el-S.M.&P. Hucl, 2003. Composition and stability of anthocyanins in blue-grained wheat. Agric Food Chem 51: 2174–2180.
Arbuzova, V.S.&O.I. Maystrenko, 2000. Chromosomal location of genes for purple grain colour introgressed in common wheat. Cereal Res Comm 28: 235–237.
Arbuzova, V.S., O.I. Maystrenko&O.M. Popova, 1998. Development of near-isogenic lines of the common wheat cultivar ‘Saratovskaya 29’. Cereal Res Comm 26: 39–46.
Blanco, A., C. De Giovanni, B. Laddomada, A. Sciancalepore, R. Simeone, K.M. Devos&M.D. Gale, 1996. Quantitative trait loci influencing grain protein content in trtraploid Wheats. Plant Breed 115: 310–316.
Bolton, F.E., 1968. Inheritance of blue aleurone and purple pericarp in hexaploid wheat. Diss Abstracts 1968:19 order No 68-13089: p. 844B Abstract Plant Breeding Abstracts 40 (1970) No 2684.
Börner, A., M.S. Röder, O. Unger&A. Meinel, 2000. The detection and molecular mapping of a major gene for non-specific adult-plant disease resistance against stripe rust (Puccinia striiformis) in wheat. Theor Appl Genet 100: 1095–1099.
Castro, A.M., A. Vasicek, M. Manifiesto, D.O. Gimenez, M.S. Tacaliti, O. Dobrovolskaya, M.S. Röder, J.W. Snape&A. Börner, 2005. Mapping antixenosis genes on chromosome 6A of wheat to green bug and to a new biotype of Russian wheat aphid. Plant Breed 124: 229–233.
Clark, J.A., 1936. Improvement of wheat. US Dept. Agric. Yearbook 1936: 207–302.
Copp, L.G.L., 1965. Purple graines in hexaploid wheat. Wheat Inform Service No19–20: 18.
Finch, R.A.&E. Simpson, 1978. New colours and complementary colour genes in barley. Z. Pflanzenzücht. 81: 40–53.
Flintham, J.E., R. Adlam&M. Gale, 1999. Seed coat and embryo dormancy in wheat. In: D. Weipert (Ed.), Proc. 8th Int. Symp. Preharvest Sprouting Cereals, pp. 67–76.
Franckowiak, J.D., 1997. Revised linkage maps for morphological markers in barley, Hordeum vulgare. Barley Genet. Newslett. 26: 9–21.
Groos, C., G. Gay, M.-R. Perretant, L. Gervais, M. Bernard, F. Dedryver&G. Charmet, 2002. Study of the relationship between pre-harvest sprouting and grain color by quantitative trait loci analysis in a white×red grain bread-wheat cross. Theor Appl Genet 104: 39–47.
Haley, S.D., P.M. Niklas, J.R. Stanly, J. Birum&J.D. Kelly, 1993. Identification of RAPD markers closely linked to a major rust resistance gene block in common bean. Theor Appl Genet 86: 505–512.
Himi, E.&K. Noda, 2004. Isolation and location of the three homoeologous dihydroflavanol-4-reductase (DFR) genes of wheat and their tissue-dependent expression. Journal Exp Botany 55: 365–375.
Hossian, K.G., V. Kalavacharla, G.R. Lazo, J. Hegstad, M.J. Wentz etal., 2004. A chromosome bin map of 2148 expressed sequence tag loci of wheat homologous group 7. Genetics 168: 687–699.
Huang, X.Q., H. Cöster, M.W. Ganal&M.S. Röder, 2003. Advance backcross QTL analysis for the identification of quantitative trait loci alleles from wild relatives of wheat (Triticum aestivum L.). Theor Appl Genet 106: 1379–1389.
Huang, X.Q., H. Kempf, M.W. Ganal&M.S. Röder, 2004. Advance backcross QTL analysis in progenies derived from a cross between a German elite winter wheat variety and a synthetic wheat (Triticum aestivum L.). Theor Appl Genet 109: 933–943.
Jende-Strid, B., 1993. Genetic control of flavonoid biosynthesis in barley. Hereditas 119: 187–204.
Khlestkina, E.K., E.G. Pestsova, M.S. Röder&A. Börner, 2002. Molecular mapping, phenotypic expression and geographical distribution of genes determining anthocyanin pigmentation of coleoptiles in wheat (Triticum aestivum L.). Theor Appl Genet 104: 632–637.
Kosambi, D.D., 1944. The estimation of map distances from recombination values. Ann Eugen 12: 172–175.
Lander, E.S., P. Green, J. Abrahamson, A. Barlow, M.J. Daly, S.E. Lincoln&L. Newburg, 1987. MAPMAKER: An interactive computer package for construction primary genetic linkage maps of experimental and natural populations. Genomics 1: 174–181.
Lila, A.M., 2004. Anthocyanins and human health: An in vitro investigative approach. J Biomed Biotechnol. 2004: 306–313.
Lundqvist, U.J., D. Franckowiak&T. Konishi, 1997. New and revised descriptions of barley genes. Barley Genet Newslett 26: 22–516.
McIntosh, R.A., 1988. Catalogue of gene symbols for wheat. Proc. Linnean Soc. 92: 204–208.
McIntosh, R.A.&E.P. Backer, 1967. Inheritance of purple pericarp in wheat. Proc. Linnean Soc. 92: 204–208.
McIntosh, R.A., Y. Yamazaki, K.M. Devos, J. Dubcovsky, J. Rogers&R. Appels, 2003. Catalogue of Gene Symbols for Wheat http://www.grs.nig.ac.jp/wheat/komugi/genes/.
Nelson, J.C., M.E. Sorrells, A.E. Van Deynze, Y.H. Lu, M. Atkinson, M. Bernard, P. Leroy, J.D. Faris&J.A. Anderson, 1995. Molecular mapping of wheat. Major genes and rearrangements in homoeologous groups 4, 5, and 7. Genetics 141: 721– 731.
Piech, J.&L.E. Evans, 1979. Monosomic analysis of purple grain colour in hexaploid wheat. Z Pflanzenzuchtg 82: 212–217.
Plaschke, J., M.W. Ganal&M.S. Röder, 1995. Detection of genetic diversity in closely related bread wheat using microsatellite markers. Theor Appl Genet 91: 1001–1007.
Röder, M.S., V. Korzun, K. Wendehake, J. Plaschke, M.-H. Tixier, P. Leroy&M.W. Ganal, 1998. A microsatellite map of wheat. Genetics 149: 2007–2023.
Salina, E., A. Börner, I. Leonova, V. Korzun, L. Laikova, O. Maystrenko&M.S. Röder, 2000. Microsatellite mapping of the induced schaerococcoid mutation genes in Triticum aestivum. Theor Appl Genet 100: 686–689.
Sharman, B.C., 1958. ‘Purple pericarp’: A monofactorial dominant in tetraploid wheats. Nature 181: 929.
Sourdille, P., S. Singh, T. Cadalen, G.L. Brown-Guedira, G. Gay, L. Qi, B.S. Gill, P. Dufour, A. Murigneux&M. Bernard, 2004. Microsatellite –based deletion bin system for the establishment of genetic-physical map relationships in wheat (Triticum aestivum L.). Theor Appl Funct Integr Genomics 4: 12–25.
Vavilov, N.I., 1962. Selected works, USSR Academical Science Publisher, Moscow, Leningrad, Vol. 111, pp. 228–235 (in Russ).
Watanabe, N., S.F. Koval&V.S. Koval, 2003. Wheat near-isogenic lines, Sankeisha, Nagoya, p. 34.
Zang, X.Q., K. Ross&J.P. Gustafson, 2000. Physical location of homoeologous groups 5 and 6 molecular markers mapped in Triticum aestivum L. Jour Heredity 91: 441–445.
Zeven, A.C., 1991. Wheat with purple and blue grains; a review. Euphytica 56: 43–258.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Dobrovolskaya, O., Arbuzova, V.S., Lohwasser, U. et al. Microsatellite mapping of complementary genes for purple grain colour in bread wheat (Triticum aestivum) L.. Euphytica 150, 355–364 (2006). https://doi.org/10.1007/s10681-006-9122-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-006-9122-7